Abstract
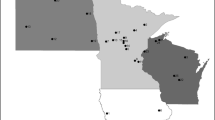
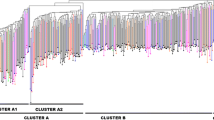
The American hazelnut (Corylus americana Marshall) is native to the eastern United States and Southern Canada. In the early decades of the last century, breeding work by several individuals attempted to combine the cold hardiness and disease resistance of the American hazelnut with the larger nut size of the European hazelnut (C. avellana L.). Many hybrid selections grown today trace back to these early efforts. Over the past three decades, representatives of C. americana were collected and are preserved in Corvallis, Oregon. Seeds were collected along roadsides, in hedgerows, in woodland clearings, and in gardens. The seeds were germinated and the best of the resulting seedlings were preserved for later use. In this study, genetic diversity was studied in 87 American and 67 hybrid hazelnut selections using 21 microsatellite loci. The total number of alleles was 266 in the 162 accessions examined, of which 229 were present in C. americana accessions, but only 168 in the Arbor Day Farm hybrids, and 116 in the hybrids of C. americana ‘Rush’. In the American accessions, a high level of genetic diversity was observed (H e = 0.74, H o = 0.68). A genetic similarity matrix of the American and hybrid accessions, and a few European cultivars, was constructed and the resulting dendrogram revealed seven major groups: European cultivars and ‘Rush’ hybrids, two groups of Arbor Day Farm hybrids, three groups of C. americana accessions, and a mixed group of hybrid and American accessions. Analysis confirmed the reported parentage of 10 hybrids. The genetic diversity among the American hazelnut accessions, untapped to date, will be useful in breeding hybrids that will allow expansion of hazelnut production into eastern North America.






Similar content being viewed by others
References
Bassil NV, Botta R, Mehlenbacher SA (2005a) Microsatellite markers in hazelnut: isolation, characterization and cross-species amplification. J Am Soc Hort Sci 130:543–549
Bassil NV, Botta R, Mehlenbacher SA (2005b) Additional microsatellite markers of the European hazelnut. Acta Hort 686:105–110
Boccacci P, Botta R (2010) Microsatellite variability and genetic structure in hazelnut (Corylus avellana L.) cultivars from different growing regions. Sci Hortic 124:128–133
Boccacci P, Akkak A, Bassil NV, Mehlenbacher SA, Botta R (2005) Characterization and evaluation of microsatellite loci in European hazelnut (Corylus avellana L.) and their transferability to other Corylus species. Mol Ecol Notes 5:934–937
Boccacci P, Akkak A, Botta R (2006) DNA typing and genetic relations among European hazelnut (Corylus avellana L.) cultivars using microsatellite markers. Genome 49:598–611
Coyne CJ, Mehlenbacher SA, Smith DC (1998) Sources of resistance to eastern filbert blight in hazelnut. J Am Soc Hort Sci 123:253–257
Crane HL, Reed CA, Wood MN (1937) Nut breeding. In: Yearbook of agriculture, pp 835–844
Erdoğan V, Mehlenbacher SA (2000) Interspecific hybridization in hazelnut (Corylus). J Am Soc Hort Sci 125:489–497
Fuller AS (1910) Filbert or hazelnut. In: The nut culturist. Orange Judd, New York, pp 118–146
Gökirmak T, Mehlenbacher SA, Bassil NV (2009) Characterization of European hazelnut (Corylus avellana) cultivars using SSR markers. Genet Resour Crop Evol 56:147–172
Gürcan K, Mehlenbacher SA (2010a) Development of microsatellite marker loci for European hazelnut (Corylus avellana L.) from ISSR fragments. Mol Breed 26:551–559
Gürcan K, Mehlenbacher SA (2010b) Transferability of microsatellite markers in the Betulaceae. J Am Soc Hort Sci 135:159–173
Gürcan K, Mehlenbacher SA, Botta R, Boccacci P (2010a) Development, characterization, segregation, and mapping of microsatellite for European hazelnut (Corylus avellana L.) from enriched genomic libraries and usefulness in genetic diversity studies. Tree Genet Genomes 6:513–531
Gürcan K, Mehlenbacher SA, Erdoğan V (2010b) Genetic diversity in hazelnut cultivars from Black Sea countries assessed using SSR markers. Plant Breed 129:422–434
Johnson KB, Mehlenbacher SA, Stone JK, Pscheidt JW, Pinkerton JN (1996) Eastern filbert blight of hazelnut: it’s becoming a manageable disease. Plant Dis 80:1308–1316
Kalinowski ST, Taper ML, Marshall TC (2007) Revising how the computer program CERVUS accommodates genotyping error increases success in paternity assignment. Mol Ecol 16:1099–1106
Liu K, Muse SV (2005) PowerMarker: integrated analysis environment for genetic marker data. Bioinformatics 21:2128–2129
Lunde CF, Mehlenbacher SA, Smith DC (2000) Survey of hazelnut cultivars for response to eastern filbert blight inoculation. HortScience 35:729–731
Molnar T, Goffreda JC, Funk CR (2005) Developing hazelnuts for the eastern United States. Acta Hort 686:609–617
Rehder A (1940) Manual of cultivated trees and shrubs. Macmillan, New York, pp 144–145
Rutter PA (1987) Badgersett Research Farm––plantings, projects and goals. Annu Rep Northern Nut Growers Assn 78:173–186
Rutter M (1991) Variation in resistance to eastern filbert blight in hybrid hazelnuts. Ann. Rept. Northern Nut Growers Assn. 82:159–162
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, and Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, Evolutionary distance, and maximum parsimony methods. Mol Biol Evol. doi:10.1093/molbev/msr121
Weschcke C (1954) Growing nuts in the north. Webb, St. Paul, MN, pp 24–38
Acknowledgments
This research was supported by funds from the Oregon Hazelnut Commission, State of Oregon, Hatch Act, and USDA Specialty Crops Research Initiative Grant 2009-51181-06028.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sathuvalli, V.R., Mehlenbacher, S.A. Characterization of American hazelnut (Corylus americana) accessions and Corylus americana × Corylus avellana hybrids using microsatellite markers. Genet Resour Crop Evol 59, 1055–1075 (2012). https://doi.org/10.1007/s10722-011-9743-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-011-9743-0