Abstract
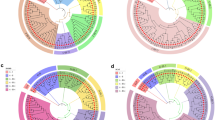
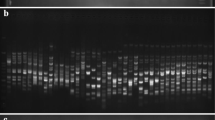
A polymerase chain reaction (PCR) based approach involving the directed amplification of minisatellite DNA region (DAMD-PCR) was used to identify accession specific DNA markers and study genetic relationships between and within 15 accessions corresponding to 11 species in genus Capsicum. A touch down PCR profile and unique chemical concentration of ingredients resulted in reproducible and reliable DNA amplifications. The number of amplified products varied from 1 to 12 fragments depending on the template DNA and the primers. The DAMD-PCR technique provided a total of 38 accession specific DNA markers (diagnostic DAMD-PCR) which can be utilized in accession identification, preservation and genetic studies of Capsicum germplasm. Based on 1,292 polymorphic and monomorphic DNA markers directed with 22 minisatellite specific primers, accessions were divided into four major groups, three of which corresponded to the three distinct Capsicum complexes. Capsicum chacoense was found to be the most distinct species.


Similar content being viewed by others
References
Baral JB, Bosland PW (2004) Unraveling the species dilemma in Capsicum frutescens and C. chinense (Solanaceae): a multiple evidence approach using morphology, molecular analysis, and sexual compatibility. J Am Soc Hortic Sci 129:826–832
Bebeli PJ, Zhou Z, Somers DJ, Gustafson JP (1997) PCR primed with minisatellite core sequences yields DNA fingerprinting probes in wheat. Theor Appl Genet 95:276–283
Bosland PW, Votava EJ (2000) Peppers: vegetable and spice capsicums. Crop production science in horticulture 12. CAB International Publishing, Wallingford, 204 pp
Buse GSC, Amaral ZP, de Bianchetti L, Machado FRB, Ferreira ME (2003) Genetic variability and phylogenetic analysis of Brazilian species of Capsicum. Capsicum Eggplant Newsl 22:13–16
Choong CY (1998) DNA polymorphisms in the study of relationships and evolution in Capsicum. PhD Thesis, The University of Reading, UK
DeWitt D, Bosland P (1996) Peppers of the world. Ten Speed Press, NY
Ellsworth DL, Rittenhouse KD, Honeycutt RL (1993) Artifactual variation in randomly amplified polymorphic DNA banding patterns. Biotechniques 14:214–216
Eshbaugh WH (1993) Peppers: history and exploitation of a serendipitous new crop discovery. In: Janick J, Simon JE (eds) New crops. Wiley, New York, pp 132–139
Guzman FA, Ayala H, Azurdia C, Duque MC, de Vicen MC (2005) AFLP assessment of genetic diversity of Capsicum genetic resources in Guatemala: home gardens as an option for conservation. Crop Sci 45:363–370
Heath DD, Iwama GK, Devlin RH (1993) PCR primed with VNTR core sequence yields species specific patterns and hypervariable probes. Nucleic Acids Res 21:5782–5785
Hunziker AT (2001) Genera Solanacearum: the genera of Solanaceae illustrated, arranged according to a new system. Gantner Verlag, Ruggell, p 516
Jarret RL, Dang P (2004) Revisiting the waxy locus and the Capsicum annuum L. complex. Ga J Sci 62:117–133
Jeffreys AJ, Wilson V, Thein SL (1985) Individual specific ‘fingerprints’ of human DNA. Nature 332:278–281
Jensen RJ, McLeod MJ, Eshbaugh WH, Guttman SI (1979) Numerical taxonomic analysis of allozymic variation in Capsicum (Solanaceae). Taxon 28:315–327
Kang HW, Park DS, Go SJ, Eun MY (2002) Fingerprinting of diverse genomes using PCR with universal rice primers generated from repetitive sequence of Korean weedy rice. Mol Cells 13:281–287
Karaca M, Ince AG (2008) Minisatellites as DNA markers to classify bermudagrasses (Cynodon spp.): confirmation of minisatellite in amplified products. J Genet 87:83–86
Karaca M, Saha S, Zipf A, Jenkins JN, Lang DJ (2002) Genetic diversity among forage bermudagrass (Cynodon spp.): evidence from chloroplast and nuclear DNA fingerprinting. Crop Sci 42:2118–2127
Karaca M, Ince AG, Elmasulu SY, Onus AN, Turgut K (2005) Coisolation of genomic and organelle DNAs from 15 genera and 31 species of plants. Anal Biochem 343:353–355
Knapp S (2002) Tobacco to tomatoes: a phylogenetic perspective on fruit diversity in the Solanaceae. J Exp Bot 53:2001–2022
Lefebvre V, Goffinet B, Chauvet JC, Caromel B, Signoret P, Brand R et al (2001) Evaluation of genetic distances between pepper inbred lines for cultivar protection purposes: comparison of AFLP, RAPD and phenotypic data. Theor Appl Genet 102:741–750
Manly BFJ (1994) Multivariate statistical methods: a primer. Chapman & Hall, London
Moscone EA, Loid J, Ehrendorfer F, Hunziker AT (1995) Analysis of active nucleolus organizing regions in Capsicum (Solanaceae) by silver staining. Am J Bot 82:267–287
Moscone EA, Lambrou M, Ehrendorfer F (1996) Fluorescent chromosome banding in the cultivated species of Capsicum (Solanaceae). Plant Syst Evol 202:37–63
Moscone EA, Baranyi M, Ebert I, Greilhuber J, Ehrendorfer F, Hunziker AT (2003) Analysis of nuclear DNA content in Capsicum (Solanaceae) by flow cytometry and Feulgen densiometry. Ann Bot (Lond) 92:21–29
Murray MJ, Haldeman BA, Grant FJ, O’Hara PJ (1988) Probing the human genome with minisatellite-like sequences from the human coagulation factor VII gene. Nucleic Acids Res 16:4166
Nakamura Y, Leppert M, O’Connell P, Wol R, Holm T, Culver M et al (1987) Variable number of tandem repeat (VNTR) markers for human gene mapping. Science 235:1616–1622
Onus AN, Pickersgill B (2004) Unilateral incompatibility in Capsicum (Solanaceae): occurrence and taxonomic distribution. Ann Bot (Lond) 94:289–295
Pickersgill B (1971) Relationships between weedy and cultivated forms in some species of chili peppers (genus Capsicum). Evolution Int J Org Evolution 25:683–691
Pickersgill B (1991) Cytogenetics and evolution of Capsicum L. In: Tsuchiya T, Gupta PK (eds) Chromosome engineering in plants: genetics, breeding, evolution, part B. Elsevier, Amsterdam, pp 139–160
Pickersgill B, Heiser CB Jr, McNeill J (1979) Numerical taxonomic studies on variation and domestication in some species of Capsicum. In: Hawkes JG, Lester RN, Skelding AD (eds) The biology and taxonomy of the Solanaceae. Linnean society symposium series no. 7. Academic Press, London, pp 679–700
Prince JP, Loaiza-Figueroa F, Tanksley SD (1992) Restriction fragment length polymorphism and genetic distance among Mexican accessions of Capsicum. Genome 35:726–732
Rodriguez JM, Berke T, Engle L, Nienhuis J (1999) Variation among and within Capsicum species revealed by RAPD markers. Theor Appl Genet 99:147–156
Ryzhova NN, Kochieva EZ (2004) Analysis of microsatellite loci of the chloroplast genome in the genus Capsicum (pepper). Russ J Genet 40:892–896
Swofford DL (2002) PAUP* 4 phlyogenetic analysis using parsimony (and other methods), version 4. Sinauer Associates, Sunderland
Tong N, Bosland PW (1999) Capsicum tovarii, a new member of the Capsicum baccatum complex. Euphytica 109:71–77
Vassaet G, Georges M, Monsieur M, Brocas H, Lequarre AS, Christophe D (1987) A sequence of M13 phage detect hypervariable minisatellites in human and animal DNA. Science 235:683–684
Vergnaud G (1989) Polymers of random short oligonucleotides detect polymorphic loci in the human genome. Nucleic Acids Res 17:7623–7630
Walsh BM, Hoot SB (2001) Phylogenetic relationships of Capsicum (Solanaceae) using DNA sequences from two noncoding regions: the chloroplast atpB-rbcl spacer region and nuclear waxy introns. Int J Plant Sci 162:1409–1418
Wasko AP, Galetti PM (2003) PCR primed with minisatellite core sequences yields species-specific patterns and assessment of population variability in fishes of the genus Brycon. J Appl Ichthyol 1:109–113
Williams CE, St Clair DA (1993) Phylogenetic relationships and levels of variability detected by restriction fragment length polymorphism and random amplified polymorphic DNA analysis of cultivated and wild accessions of Lycopersicon esculentum. Genome 36:619–630
Acknowledgement
This research was in part supported by the Scientific Research Projects Administration Unit of Akdeniz University.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ince, A.G., Karaca, M. & Onus, A.N. Development and utilization of diagnostic DAMD-PCR markers for Capsicum accessions. Genet Resour Crop Evol 56, 211–221 (2009). https://doi.org/10.1007/s10722-008-9356-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-008-9356-4