Abstract
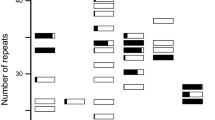
Native species show adaptive traits that are difficult to find in introduced species. The Pampas region in Argentina is a valuable nature reserve of grasses and Paspalum dilatatum Poir. is one of the most important grasses found there. Based on ploidy level and on morphological traits, five biotypes of P. dilatatum have been described. Two of them were included in this study: a tetraploid biotype with sexual reproduction and a pentaploid biotype with apomictic reproduction. We analyzed the genetic diversity in eight native populations from the Salado basin, Argentina, using both quantitative traits and molecular data (RAPD) with these aims: to obtain information of the degree of phenotypic variation in that area, to know which the pattern of distribution of this variation is and to look for association between molecular markers with populational or biotypic differentiation. Cluster analysis based on morphological data grouped the individuals of the different populations by ploidy level. Molecular markers showed the inverse situation because individuals were grouped by geographic origin as opposed to biotype. Moreover, since RAPD did not discriminate between biotypes with sexual or apomictic reproduction, they are probably not associated with mating system. The results let us conclude that polygenic traits such as LP, LBSR, NRT and NSP can discriminate between biotypes and molecular markers such as bands 12, 40, 19 and 46 can be used to discriminate among populations, probably because they detect neutral variation.
Similar content being viewed by others
References
Barreto IL (1974) O género Paspalum (Gramineae) no rio grande do Sul. Tesis (Livre Docência). Departamento de Fitotecnia, Facultade do Rio Grande do Sul, Porto Alegre, 258 pp
Bashaw EC, Holt E (1958) Megasporogenesis, embryo sac development and embryogenesis in dallisgrass, Paspalum dilatatum Poir. Agron J 50:753–756
Bassi P (1999) The effect of environmental stress on repetitive DNA behavior in plants. In: Lerner R (ed) Plant responses to environmental stresses. Marcel Dekker, Inc. New York, 730 pp
Bennet HW, Burson BL, Bashaw S (1969) Intraspeciefic hybridization in dallisgrass, Paspalum dilatatum Poir. Crop Sci 9:807–809
Beer SC, Goffreda J, Phillips TD, Murphy JP, Sorrells ME (1993) Assessment of genetic variation in Avena sterilis using morphological traits, isozymes and RFLP. Crop Sci 33:1386–1393
Burkart A (1969) Flora ilustrada de Entre Ríos (Argentina). Parte II. INTA. 551 pp
Burson BL (1991) Genome relationships between tetraploid and hexaploid biotypes of dallisgrass, Paspalum dilatatum. Bot Gaz 152(2):219–223
Burson BL, Bennett HW (1972) Genome relations between an intraspecific Paspalum dilatatum hybrid and two diploid Paspalum species. Can J Genet Cytol 14:609–613
Burson BL, Lee H, Bennett HW (1973) Genome relations between tetraploid Paspalum dilatatum and four diploid Paspalum species. Crop Sci 13:739–743
Burson BL, Voigt PW, Evers GW (1991) Cytology, reproductive behavior, and forage potential of hexaploid dallisgrass biotypes. Crop Sci 31:636–641
Caetano-Anollés G (1998) DNA analysis of turfgrass genetic diversity. Crop Sci 38:1415–1424
Carámbula M (1982) Paspalum dilatatum, características agronómicas y su rol en las pasturas. Revista Argentina de Producción Animal 2:68–84
Casa AM (1995) Caracterizaçao do germoplasma de especies de Paspalum (grupo Dilatata) pelo uso de marcadores bioquimicos e moleculares. Tesis (Mestre em Ciências Biológicas), Instituto de Biociencias, Universidade Estadual Paulista “Julio de Mesquita Filho”, Botucatu. 102 pp
Cullis CA (1999) The environment as an active generator of adaptive genomic variation. In: Lerner R (ed) Plant responses to environmental stresses. Marcel Dekker, Inc. New York, 730 pp
Espinoza F, Quarín CL (2000) 2n + n hybridization of apomictic Paspalum dilatatum with diploid Paspalum species. Int J Plant Sci 161(1):221–225
Fernández Grecco RC, Viviani Rossi EM (1997) Guía de reconocimiento de especies de campo natural. INTA Centro regional Buenos Aires Sur. Materiales didácticos N° 13. 67 pp
Ferreira ME, Grattaplagia D (1996) Introducao ao uso de marcadores moleculares em analise genética. EMBRAPA – CENARGEN, Documento 20, 220 pp
García MV, Arturi MJ, Ansín OE (2002) Variabilidad fenotípica y genética en poblaciones de Paspalum dilatatum Poir”. Agricultura Técnica (Chile) 62(2):237–244
Garita F, Alonso F, Clausen A (1994) Caracterización de subespecies de Paspalum dilatatum Poir. Resúmenes del Congreso Latinoamericano de Botánica. 499 pp
Holt EC (1956) Dallisgrass. Texas Agricultural Experiment Station, EUA Boletín 829. 14 pp
Jarret RL, Liu ZW, Webster RW (1998) Genetic diversity among Paspalum spp. as determined by RFLPs. Euphytica 104:119–125
Josifovich J, Maddaloni J, Serrano H, Echeverría I (1982) Areas forrajeras y de producción animal en la Argentina. Informe Técnico N°169 EEA INTA Pergamino. 110 pp
Legendre L, Legendre P (1983) Numerical ecology. Esevier, New York, 419 pp
Liu, ZW, Jarret RL, Duncan RR, Kresovich S (1994) Genetic relationship and variation among ecotypes of seashore paspalum (Paspalum vaginatum) determined by random amplified polymorphic DNA markers. Genome 37:1011–1017
M’Ribu HK, Hilu KW (1996) Application of random amplified polymorphic DNA to study genetic diversity in Paspalum scrobiculatum L. (Kodo millet, Poaceae). Genet Resour Crop Evol 43:203–210
Ortiz JPA, Pessino SC, Leblanc O, Hayward MD, Quarín CL (1997) Genetic fingerprinting for determining the mode of reproduction in Paspalum notatum, a subtropical apomictic forage grass. Theor Appl Genet 95:850–856
Pahlen A (1986) Evaluation of genetic variability of some native forage plants. Boletín Genético 14:1–6, CICA Castelar
Percival NS, Couchman JN (1979) Evaluation of paspalum (Paspalum dilatatum Poir.) selections. 1. Variation among populations. NZ J Exp Agric 7:59–64
Pereira J, Fajardo A, Sabbía V (1999) Caracterización genética y filogenética del genero Paspalum a partir de marcadores moleculares. Agrociencia 3(1):11–19
Pereira J, Sabbía V, Fajardo A, Speranza PR (2000) Análisis genético en gramíneas nativas del género Paspalum a partir de datos isoenzimáticos y RAPD. Agrociencia 4(1):1–11
Quarín CL, Norrmann GA (1986) Anales del IV Congreso Latinoamericano de Botánica, Medellín. pp 25–34
Skroch P, Tivang J, Nienhuis J (1992) Analysis of genetic relationships using RAPD marker data. En: Proceedings of the Symposium Applications of RAPD technology to plant breeding, Minnesota. pp 26–30
Smouse PE, Long JC, Sokal RR (1986) Multiple regression and correlation extensions of the Mantel test of matrix correspondence. Syst Zool 35(4):627–632
Sneath PHA, Sokal RR (1983) Numerical taxonomy. WH Freeman and Co
Steel RGD, Torrie JH (1980) Principles and procedures of statistics. A biometrical approach, 2nd ed. McGraw Hill Book Company
Steiner JJ, Poklemba CJ, Fjellstrom RG, Elliot LF (1995) A rapid one tube genomic DNA extraction process for PCR and RAPD analyses. Nucl Acids Res 23(13):2569–2570
Stewart CN, Excoffier L (1996) Assessing population genetic structure and variability with RAPD data: application to Vaccinium macrocarpa (American cranberry). J Evol Biol 9:153–171
Acknowledgements
This study has been funded by grants from the Consejo Nacional de Investigaciones Científicas y Técnicas (1074/98) and from Universidad Nacional de La Plata. We would like to thanks to P.K.Ingvarsson for helpful comments to the manuscript and N. Di Lorenzo for the data on resistance to Claviceps paspali.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
García, M.V., Balatti, P.A. & Arturi, M.J. Genetic variability in natural populations of Paspalum dilatatum Poir. analyzed by means of morphological traits and molecular markers. Genet Resour Crop Evol 54, 935–946 (2007). https://doi.org/10.1007/s10722-006-9147-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-006-9147-8