Abstract
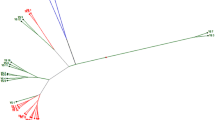
The genetic diversity and the relationships among a collection of Brassica napus L. European populations were evaluated using random amplified polymorphic DNA markers. The study included 33 accessions of B. napus collected from Galicia (northwestern Spain) and 18 British cultivars, 16 accessions of B. napus and two accessions of Brassica oleracea L. used as controls. DNA from 25 individuals per population was analyzed using 18 decamer primers. One hundred thirty-eight amplification products were scored of which 105 were polymorphic. These bands ranged in size from 350 to 2500 base pairs. Similarity coefficients and cluster analysis were computed and six groups were obtained. Cluster I was the largest and included all the landraces from northwestern Spain, except two accessions that grouped separately into Clusters III and IV, respectively. A low level of genetic variability was detected among the B. napus Spanish genotypes, while considerable diversity was present among the British ones, which grouped into three groups, two main clusters and one group formed by one accession. Cluster II included all commercial varieties grown in Great Britain whereas Cluster V grouped local varieties maintained by the growers for many years. Cluster VI was a singularity formed by one entry. British accessions of B. oleracea had the greatest dissimilarity with all the other populations and grouped separately in Clusters VII and VIII. As conclusion, B. napus landraces used in northwestern Spain as leafy-green vegetable probably have an independent origin from B. napus crops grown in other European regions. Besides, separate domestication in northwestern Spain and Great Britain for a different end use might have led to two distinct gene pools.
Similar content being viewed by others
References
P. Arús J.J. Baladrón A. Ordás (1987) ArticleTitleSpecies identification of cultivated brassicas with isozyme electrophoresis Crucifer. Newsl. 12 26–27
R.L. Cassian S. Echeverrigaray (2000) ArticleTitleDiscrimination among cultivars of cabbage using random amplified polymorphic DNA markers HortScience 35 1155–1158
P.A. Crockett P.L. Bhalla C.K. Lee M.B. Singh (2000) ArticleTitleRAPD analysis of seed purity in a commercial hybrid cabbage (Brassica oleracea var. capitata) cultivar Genome 43 317–321 Occurrence Handle10791820 Occurrence Handle1:CAS:528:DC%2BD3cXjtVyjsr4%3D Occurrence Handle10.1139/gen-43-2-317
T. Demeke R.P. Adams R. Chibbar (1992) ArticleTitlePotential taxonomic use of random amplified polymorphic DNA (RAPD): a case study in Brassica Theor. Appl. Genet. 84 990–994 Occurrence Handle1:CAS:528:DyaK3sXjs1Sktw%3D%3D Occurrence Handle10.1007/BF00227415
I. Divaret E. Margalé G. Thomas (1999) ArticleTitleRAPD markers on seed bulks efficiently assess the genetic diversity of a Brassica oleracea L. collection Theor. Appl. Genet. 98 1029–1035 Occurrence Handle1:CAS:528:DyaK1MXktFSms7Y%3D Occurrence Handle10.1007/s001220051164
M.W. Farnham (1996) ArticleTitleGenetic variation among and within United States collard cultivars and landraces as determined by randomly amplified polymorphic DNA markers J. Amer. Soc. Hort. Sci. 121 374–379
C. Gallástegui (1926) ArticleTitleNúmero de cromosomas en algunas especies del género Brassica Bol. Real Soc. Esp. Hist. Nat. 26 185–191
T. Gladis K. Hammer (1992) ArticleTitleDie Gaterslebener Brassica-Kollektion-Brassica juncea B. napus B. nigra B. rapa Feddes Rep. 103 469–507
C. Gómez-Campo S. Prakash (1999) Origin and domestication C. Gómez-Campo (Eds) Biology of Brassica Coenospecies Elsevier Science B.V. Amsterdam The Netherlands 33–58
J. Hu C.F. Quirós (1991) ArticleTitleIdentification of broccoli and cauliflower cultivars with RAPD markers Plant Cell Rep. 10 505–511 Occurrence Handle10.1007/BF00234583
A. Jain S. Bathia S.S. Banga S. Prakash M. Laksmikumaran (1994) ArticleTitlePotential use of random amplified polymorphic DNA (RAPD) technique to study the genetic diversity in Indian mustard (Brassica juncea) and its relationship to heterosis Theor. Appl. Genet. 88 116–122 Occurrence Handle1:CAS:528:DyaK2MXjsVWrsg%3D%3D Occurrence Handle10.1007/BF00222403
S. Kresovich J.G.K. Williams J.R. McFerson E.J. Routman B.A. Schaal (1992) ArticleTitleCharacterization of genetic identities and relationships of Brassica oleracea L. via random amplified polymorphic DNA assay Theor. Appl. Genet. 85 190–196 Occurrence Handle1:CAS:528:DyaK3sXlslOltbs%3D Occurrence Handle10.1007/BF00222859
Y. Liu R.F. Whittier (1994) ArticleTitleRapid preparation of megabase plant DNA from nuclei in agarose plugs and microbeads Nucl. Acid Res. 22 2168–2169 Occurrence Handle1:CAS:528:DyaK2cXltVSktbo%3D
R.J. Mailer R. Searth B. Fristensky (1994) ArticleTitleDiscrimination among cultivars of rapeseed (Brassica napus L.) using DNA polymorphisms amplified from arbitrary primers Theor. Appl. Genet. 87 697–704 Occurrence Handle1:CAS:528:DyaK2MXkt1Cqtw%3D%3D Occurrence Handle10.1007/BF00222895
M. Nei W.H. Li (1979) ArticleTitleMathematical model for studying genetic variation in terms of restriction endonucleases Proc. Natl. Acad. Sci. USA 76 5269–5273 Occurrence Handle291943 Occurrence Handle1:CAS:528:DyaL3cXitVWn
A. Ordás J.J. Baladrón (1985) ArticleTitleCollecting of brassicas in northwestern Spain Crucifer. Newsl. 10 14
W.B. Phippen S. Kresovich F.G. Candelas J.R. McFerson (1997) ArticleTitleMolecular characterization can quantify and partition variation among genebank holdings: a case study with phenotypically similar accessions of Brassica oleracea var. capitata L. (cabbage) ‘8Golden Acre’9 Theor. Appl. Genet. 94 227–234 Occurrence Handle1:CAS:528:DyaK2sXhvV2gsbw%3D Occurrence Handle10.1007/s001220050404
S. Prakash Y. Takahata P.B. Kirti V.L. Chopra (1999) Cytogenetics C. Gómez-Campo (Eds) Biology of Brassica Coenospecies Elsevier Science B.V. Amsterdam The Netherlands 59–106
M.A. Rabbani A. Iwabuchi Y. Murakami T. Suzuki K. Takayanagi (1998) ArticleTitleGenetic diversity in mustard (Brassica juncea L.) germplasm from Pakistan as determined by RAPDs Euphytica 103 235–242 Occurrence Handle1:CAS:528:DyaK1MXlt1agsw%3D%3D Occurrence Handle10.1023/A:1018304921526
J. Ren J.R. McFerson S. Kresovich W.F. Lamboy (1995) ArticleTitleIdentities and relationships among Chinese vegetable brassicas as determined by random amplified polymorphic DNA markers J. Amer. Soc. Hort. Sci. 120 548–555
F.J. Rohlf (1998) NTSYS-pc. Numerical Taxonomy and Multivariate Analysis System. Version 2.02e EXETER software Setauket, New York
K. Song K. Tang T.C. Osborn (1993) ArticleTitleDevelopment of synthetic Brassica amphiploids and comparison to natural amphiploids Theor. Appl. Genet. 86 811–821 Occurrence Handle10.1007/BF00212606
J.G.K. Williams A.R. Kubelik K.J. Livak J.A. Rafalski S.V. Tingey (1990) ArticleTitleDNA polymorphisms amplified by arbitrary primers are useful as genetic markers Nucl. Acids Res. 18 6531–6535 Occurrence Handle1979162 Occurrence Handle1:CAS:528:DyaK3MXjslWmsA%3D%3D
A.C. Zeven K.J. Dehmer T. Gladis K. Hammer H. Lux (1990) ArticleTitleAre the duplicates of perennial kales (Brassica oleracea L. var. ramosa DC.) true duplicates as determined by RAPD analysis? Genet. Resour. Crop Evol. 45 105–111
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Cartea, M.E., Soengas, P., Picoaga, A. et al. Relationships Among Brassica napus (L.) Germplasm from Spain and Great Britain as Determined by RAPD Markers. Genet Resour Crop Evol 52, 655–662 (2005). https://doi.org/10.1007/s10722-003-6014-8
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s10722-003-6014-8