Abstract
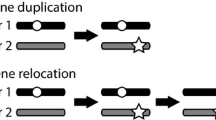
Gene duplication is a major force driving genome evolution, and examples of this mode of evolution and of the functions of duplicated genes are needed to reveal general patterns. Here, our study focuses on a particular retrogene (i.e., CG9573) that originated about 5–13 million years ago that we have named Drcd-1 related. It originated in Drosophila through retroposition of the parental gene Required for cell differentiation 1 of Drosophila (Drcd-1; CG14213), which is a known transcription cofactor. Drcd-1r is only present in D. melanogaster, D. simulans, D. sechellia, and D. mauritiana. Drcd-1r is an X to autosome retroposition event. Many retrogenes are X to autosome copies and it has been shown that positive selection underlies this bias. We sought to understand Drcd-1r mode of evolution and function to contribute to the understanding of the selective pressures acting on X to autosome retrogenes. Drcd-1r overlaps with another gene, it is within the 3′ UTR of the gene CG13102 and is encoded in the opposite orientation. We have studied the characteristics of the transcripts and quantified expression of CG13102 and Drcd-1r in wild-type flies. We found that Drcd-1r is transcribed specifically in testes. We also studied the molecular evolution of Drcd-1r and Drcd-1 and found that the parental gene has evolved under very strong purifying selection but the retrogene has evolved very rapidly (Ka/Ks ~1) under both positive and purifying selection, as revealed using divergence and polymorphism data. These results indicate that Drcd-1r has a novel function in the Drosophila testes. To further explore Drcd-1r function we used a strain containing a P element inserted in the region where CG13102 and Drcd-1r are located that shows recessive male sterility. Analysis of this strain reveals the difficulties that can be encountered in studying the functions of genes with overlapping transcripts. Avenues for studying of the function of this gene are proposed.




Similar content being viewed by others
References
Aravin AA, Naumova NM, Tulin AV, Vagin VV, Rozovsky YM, Gvozdev VA (2001) Double-stranded RNA-mediated silencing of genomic tandem repeats and transposable elements in the D. melanogaster germline. Curr Biol 11:1017–1027
Arguello JR, Chen Y, Yang S, Wang W, Long M (2006) Origination of an X-linked testes chimeric gene by illegitimate recombination in drosophila. PLoS Genet 2:e77
Bai Y, Casola C, Feschotte C, Betran E (2007) Comparative genomics reveals a constant rate of origination and convergent acquisition of functional retrogenes in drosophila. Genome Biol 8:R11
Bayes JJ, Malik HS (2009) Altered heterochromatin binding by a hybrid sterility protein in Drosophila sibling species. Science 326:1538–1541
Betran E, Long M (2003) Dntf-2r: a young Drosophila retroposed gene with specific male expression under positive Darwinian selection. Genetics 164:977–988
Betran E, Thornton K, Long M (2002) Retroposed new genes out of the X in Drosophila. Genome Res 12:1854–1859
Betran E, Emerson JJ, Kaessmann H, Long M (2004) Sex chromosomes and male functions: where do new genes go? Cell Cycle 3:873–875
Blumer N, Schreiter K, Hempel L, Santel A, Hollmann M, Schafer MA, Renkawitz-Pohl R (2002) A new translational repression element and unusual transcriptional control regulate expression of don juan during Drosophila spermatogenesis. Mech Dev 110:97–112
Brosius J (1991) Retroposons—seeds of evolution. Science 251:753
Castrillon DH, Gonczy P, Alexander S, Rawson R, Eberhart CG, Viswanathan S, Dinardo S, Wasserman SA (1993) Toward a molecular genetic analysis of spermatogenesis in Drosophila melanogaster: characterization of male-sterile mutants generated by single P element mutagenesis. Genetics 135:489–505
Chen X, Hiller M, Sancak Y, Fuller M (2005) Tissue-specific TAFs counteract Polycomb to turn on terminal differentiation. Science 310:869–872
Chintapalli VR, Wang J, Dow JA (2007) Using FlyAtlas to identify better Drosophila melanogaster models of human disease. Nat Genet 39:715–720
Chippindale AK, Gibson JR, Rice WR (2001) Negative genetic correlation for adult fitness between sexes reveals ontogenetic conflict in Drosophila. Proc Natl Acad Sci USA 98:1671–1675
Clark AG, Eisen MB, Smith DR, Bergman CM, Oliver B, Markow TA, Kaufman TC, Kellis M, Gelbart W, Iyer VN, Pollard DA, Sackton TB, Larracuente AM, Singh ND, Abad JP, Abt DN, Adryan B, Aguade M, Akashi H, Anderson WW, Aquadro CF, Ardell DH, Arguello R, Artieri CG, Barbash DA, Barker D, Barsanti P, Batterham P, Batzoglou S, Begun D, Bhutkar A, Blanco E, Bosak SA, Bradley RK, Brand AD, Brent MR, Brooks AN, Brown RH, Butlin RK, Caggese C, Calvi BR, Bernardo De Carvalho A, Caspi A, Castrezana S, Celniker SE, Chang JL, Chapple C, Chatterji S, Chinwalla A, Civetta A, Clifton SW, Comeron JM, Costello JC, Coyne JA, Daub J, David RG, Delcher AL, Delehaunty K, Do CB, Ebling H, Edwards K, Eickbush T, Evans JD, Filipski A, Findeiss S, Freyhult E, Fulton L, Fulton R, Garcia AC, Gardiner A, Garfield DA, Garvin BE, Gibson G, Gilbert D, Gnerre S, Godfrey J, Good R, Gotea V, Gravely B, Greenberg AJ, Griffiths-Jones S, Gross S, Guigo R, Gustafson EA, Haerty W, Hahn MW, Halligan DL, Halpern AL, Halter GM, Han MV, Heger A, Hillier L, Hinrichs AS, Holmes I, Hoskins RA, Hubisz MJ, Hultmark D, Huntley MA, Jaffe DB, Jagadeeshan S et al (2007) Evolution of genes and genomes on the Drosophila phylogeny. Nature 450:203–218
Dorus S, Freeman ZN, Parker ER, Heath BD, Karr TL (2008) Recent origins of sperm genes in Drosophila. Mol Biol Evol 25:2157–2166
Emerson JJ, Kaessmann H, Betran E, Long M (2004) Extensive gene traffic on the mammalian X chromosome. Science 303:537–540
Esnault C, Maestre J, Heidmann T (2000) Human LINE retrotransposons generate processed pseudogenes. Nat Genet 24:363–367
Fontanillas P, Hartl DL, Reuter M (2007) Genome organization and gene expression shape the transposable element distribution in the Drosophila melanogaster euchromatin. PLoS Genet 3:2256–2267
Garces RG, Gillon W, Pai EF (2007) Atomic model of human Rcd-1 reveals an armadillo-like-repeat protein with in vitro nucleic acid binding properties. Protein Sci 16:176–188
Haas M, Siegert M, Schurmann A, Sodeik B, Wolfes H (2004) c-Myb protein interacts with Rcd-1, a component of the CCR4 transcription mediator complex. Biochemistry 43:8152–8159
Hense W, Baines JF, Parsch J (2007) X chromosome inactivation during Drosophila spermatogenesis. PLoS Biol 5:e273
Hiller MA, Lin TY, Wood C, Fuller MT (2001) Developmental regulation of transcription by a tissue-specific TAF homolog. Genes Dev 15:1021–1030
Hiller M, Chen X, Pringle MJ, Suchorolski M, Sancak Y, Viswanathan S, Bolival B, Lin TY, Marino S, Fuller MT (2004) Testis-specific TAF homologs collaborate to control a tissue-specific transcription program. Development 131:5297–5308
Hiroi N, Ito T, Yamamoto H, Ochiya T, Jinno S, Okayama H (2002) Mammalian Rcd1 is a novel transcriptional cofactor that mediates retinoic acid-induced cell differentiation. EMBO J 21:5235–5244
Hollocher H, Ting CT, Wu ML, Wu CI (1997) Incipient speciation by sexual isolation in Drosophila melanogaster: variation in mating preference and correlation between sexes. Evolution 51:1175–1181
Hwa JJ, Zhu AJ, Hiller MA, Kon CY, Fuller MT, Santel A (2004) Germ-line specific variants of components of the mitochondrial outer membrane import machinery in Drosophila. FEBS Lett 572:141–146
Kalamegham R, Sturgill D, Siegfried E, Oliver B (2007) Drosophila mojoless, a retroposed GSK-3, has functionally diverged to acquire an essential role in male fertility. Mol Biol Evol 24:732–742
Kempe E, Muhs B, Schafer M (1993) Gene regulation in Drosophila spermatogenesis: analysis of protein binding at the translational control element TCE. Dev Genet 14:449–459
Kosakovsky Pond SL, Frost SD (2005) Not so different after all: a comparison of methods for detecting amino acid sites under selection. Mol Biol Evol 22:1208–1222
Lifschytz E, Lindsley DL (1972) The role of X-chromosome inactivation during spermatogenesis. Proc Natl Acad Sci USA 69:182–186
Long M, Langley CH (1993) Natural selection and the origin of jingwei, a chimeric processed functional gene in Drosophila. Science 260:91–95
Mcdonald JH, Kreitman M (1991) Adaptive protein evolution at the Adh locus in Drosophila. Nature 351:652–654
Nielsen R, Yang Z (1998) Likelihood models for detecting positively selected amino acid sites and applications to the HIV-1 envelope gene. Genetics 148:929–936
Ohno S (1970) Evolution by gene duplication. Berlin, Springer
Okazaki N, Okazaki K, Watanabe Y, Kato-Hayashi M, Yamamoto M, Okayama H (1998) Novel factor highly conserved among eukaryotes controls sexual development in fission yeast. Mol Cell Biol 18:887–895
Pond SL, Frost SD (2005) Datamonkey: rapid detection of selective pressure on individual sites of codon alignments. Bioinformatics 21:2531–2533
Presgraves DC (2007) Does genetic conflict drive rapid molecular evolution of nuclear transport genes in Drosophila? Bioessays 29:386–391
Presgraves DC, Stephan W (2007) Pervasive adaptive evolution among interactors of the Drosophila hybrid inviability gene, Nup96. Mol Biol Evol 24:306–314
Pröschel M, Zhang Z, Parsch J (2006) Widespread adaptive evolution of Drosophila genes with sex-biased expression. Genetics 174:893–900
Puig M, Caceres M, Ruiz A (2004) Silencing of a gene adjacent to the breakpoint of a widespread Drosophila inversion by a transposon-induced antisense RNA. Proc Natl Acad Sci USA 101:9013–9018
Ranz JM, Castillo-Davis CI, Meiklejohn CD, Hartl DL (2003) Sex-dependent gene expression and evolution of the Drosophila transcriptome. Science 300:1742–1745
Rice WR (1992) Sexually antagonistic genes: experimental evidence. Science 256:1436–1439
Rozas J, Sanchez-Delbarrio JC, Messeguer X, Rozas R (2003) DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19:2496–2497
Schäfer M, Kuhn R, Bosse F, Schäfer U (1990) A conserved element in the leader mediates post-meiotic translation as well as cytoplasmic polyadenylation of a Drosophila spermatocyte mRNA. EMBO J 9:4519–4525
Schäfer M, Nayernia K, Engel W, Schäfer U (1995) Translational control in spermatogenesis. Dev Biol 172:344–352
Schlenke TA, Begun DJ (2003) Natural selection drives Drosophila immune system evolution. Genetics 164:1471–1480
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 3:1101–1108
Sturgill D, Zhang Y, Parisi M, Oliver B (2007) Demasculinization of X chromosomes in the Drosophila genus. Nature 450:238–241
Sun S, Ting CT, Wu CI (2004) The normal function of a speciation gene, Odysseus, and its hybrid sterility effect. Science 305:81–83
Swanson WJ, Vacquier VD (2002) The rapid evolution of reproductive proteins. Nat Rev Genet 3:137–144
Swanson WJ, Clark AG, Waldrip-Dail HM, Wolfner MF, Aquadro CF (2001) Evolutionary EST analysis identifies rapidly evolving male reproductive proteins in Drosophila. Proc Natl Acad Sci USA 98:7375–7379
Tamura K, Subramanian S, Kumar S (2004) Temporal patterns of fruit fly (Drosophila) evolution revealed by mutation clocks. Mol Biol Evol 21:36–44
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Timakov B, Zhang P (2001) The hsp60B gene of Drosophila melanogaster is essential for the spermatid individualization process. Cell Stress Chaperones 6:71–77
Ting CT, Tsaur SC, Wu ML, Wu CI (1998) A rapidly evolving homeobox at the site of a hybrid sterility gene. Science 282:1501–1504
Ting CT, Tsaur SC, Wu CI (2000) The phylogeny of closely related species as revealed by the genealogy of a speciation gene, Odysseus. Proc Natl Acad Sci USA 97:5313–5316
Tracy C, Rio J, Motiwale M, Christensen SM, Betran E (2010) Convergently recruited nuclear transport retrogenes are male biased in expression and evolving under positive selection in Drosophila. Genetics 184:1067–1076
Tripoli G, D’elia D, Barsanti P, Caggese C (2005) Comparison of the oxidative phosphorylation (OXPHOS) nuclear genes in the genomes of Drosophila melanogaster, Drosophila pseudoobscura and Anopheles gambiae. Genome Biol 6:R11
Vibranovski MD, Lopes HF, Karr TL, Long M (2009) Stage-specific expression profiling of Drosophila spermatogenesis suggests that meiotic sex chromosome inactivation drives genomic relocation of testis-expressed genes. PLoS Genet 5:e1000731
Vinckenbosch N, Dupanloup I, Kaessmann H (2006) Evolutionary fate of retroposed gene copies in the human genome. Proc Natl Acad Sci USA 103:3220–3225
Wilson RJ, Goodman JL, Strelets VB, Consortium F (2008) FlyBase: integration and improvements to query tools. Nucleic Acids Res 36:D588–D593
Wu C-I, Xu EY (2003) Sexual antagonism and X inactivation—the SAXI hypothesis. TIGS 19:243–247
Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Yang Z, Nielsen R, Goldman N, Pedersen AM (2000) Codon-substitution models for heterogeneous selection pressure at amino acid sites. Genetics 155:431–449
Yang Z, Wong WS, Nielsen R (2005) Bayes empirical Bayes inference of amino acid sites under positive selection. Mol Biol Evol 22:1107–1118
Yuan X, Miller M, Belote JM (1996) Duplicated proteasome subunit genes in Drosophila melanogaster encoding testes-specific isoforms. Genetics 144:147–157
Zhong L, Belote JM (2007) The testis-specific proteasome subunit Prosalpha6T of D. melanogaster is required for individualization and nuclear maturation during spermatogenesis. Development 134:3517–3525
Acknowledgments
We thank Dr. Chitra Chandrasekaran, Sidi Chen and Dr. Manyuan Long for comments on this work. We also thank Mao-Lien Wu, Chung-I Wu, Tessa Bauer DuMont and Chip Aquadro for providing stocks. Research was supported by UTA startup funds and grant R01 GM071813 from the National Institutes of Health, USA to EB.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
10709_2010_9474_MOESM1_ESM.pdf
Supplementary Figure 1. Alignment of genomic regions of Drcd-1 and Drcd-1r. The alignment shows ATG and stop codons for both genes. Note that sequence conservation exists only in the CDS. Sequence conservation does not extend to other gene regions (i.e., UTRs). Neither remnants of poly-A tract nor direct repeats remain in the retrogene insertion site. The non-homologous 3’ region is shown in blue. (PDF 23 kb)
10709_2010_9474_MOESM2_ESM.pdf
Supplementary Figure 2. Drcd-1r genomic region. The genomic organization of the region where Drcd-1r is inserted is given as annotated in FlyBase. CDS regions are in bold. The Drcd-1r (CG9573) sequence has been reversed and complemented. The location of the P element insertion of line 11773 is also shown. (PDF 39 kb)
10709_2010_9474_MOESM3_ESM.pdf
Supplementary Figure 3. Drcd-1r alignment between species. Potentially positively selected codons are highlighted in blue. (PDF 27 kb)
10709_2010_9474_MOESM4_ESM.pdf
Supplementary Figure 4. Alignment of D. melanogaster Drcd-1r and Drcd-1 with human Rcd-1. Amino acids that participate in dimerization in humans are highlighted in green. Amino acids of the positively charged cleft that binds DNA in humans are highlighted in blue. Residues are highlighted in Drosophila genes in either green or blue if conserved. Differences between Rcd-1 and Drcd-1r proteins are highlighted in red. (PDF 30 kb)
Rights and permissions
About this article
Cite this article
Quezada-Díaz, J.E., Muliyil, T., Río, J. et al. Drcd-1 related: a positively selected spermatogenesis retrogene in Drosophila. Genetica 138, 925–937 (2010). https://doi.org/10.1007/s10709-010-9474-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-010-9474-8