Abstract
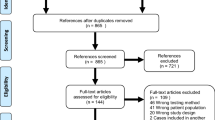
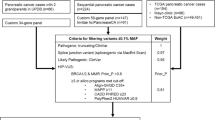
Approximately 5–10% of all pancreatic cancer patients carry a predisposing mutation in a known susceptibility gene. Since >90% of patients present with late stage disease, it is crucial to identify high risk individuals who may be amenable to early detection or other prevention. To explore the spectrum of hereditary pancreatic cancer susceptibility, we evaluated germline DNA from pancreatic cancer participants (n = 53) from a large hereditary cancer registry. For those without a known predisposition mutation gene (n = 49), germline next generation sequencing was completed using targeted capture for 706 candidate genes. We identified 16 of 53 participants (30%) with a pathogenic (P) or likely pathogenic (LP) variant that may be related to their hereditary pancreatic cancer predisposition; seven had mutations in genes associated with well-known cancer syndromes (13%) [ATM (2), BRCA2 (3), MSH2 (1), MSH6 (1)]. Many had mutations in Fanconi anemia complex genes [BRCA2 (3 participants), FANCF, FANCM]. Eight participants had rare protein truncating variants of uncertain significance with no other P or LP variants. Earlier age of pancreatic cancer diagnosis (57.5 vs 64.8 years) was indicative of possessing a P or LP variant, as was cancer family history (p values <0.0001). Our multigene panel approach for identifying known cancer predisposing genetic susceptibility in those at risk for hereditary pancreatic cancer may have direct applicability to clinical practice in cases with mutations in actionable genes. Future pancreatic cancer predisposition studies should include evaluation of the Fanconi anemia genes.

Similar content being viewed by others
References
Surveillance Epidemiology and End, Results Program N (2016) SEER Fact Sheets: Pancreas Cancer. National Cancer Institute. http://seer.cancer.gov/statfacts/html/pancreas.html. Accessed 31 Oct 2016
Klein AP (2012) Genetic susceptibility to pancreatic cancer. Mol Carcinog 51(1):14–24. doi:10.1002/mc.20855
Holter S, Borgida A, Dodd A, Grant R, Semotiuk K, Hedley D, Dhani N, Narod S, Akbari M, Moore M, Gallinger S (2015) Germline BRCA mutations in a large clinic-based cohort of patients with pancreatic adenocarcinoma. J Clin Oncol 33(28):3124–3129. doi:10.1200/jco.2014.59.7401
Connor AA, Gallinger S (2015) Hereditary pancreatic cancer syndromes. Surg Oncol Clin N Am 24(4):733–764. doi:10.1016/j.soc.2015.06.007
Solomon S, Das S, Brand R, Whitcomb DC (2012) Inherited pancreatic cancer syndromes. Cancer J 18(6):485–491. doi:10.1097/PPO.0b013e318278c4a6
NCCN (2017) NCCN genetic/familial high-risk assessment: breast and ovarian version 2.2017.
MacDonald DJ, Blazer KR, Weitzel JN (2010) Extending comprehensive cancer center expertise in clinical cancer genetics and genomics to diverse communities: the power of partnership. J Natl Compr Cancer Netw 8(5):615–624
Weitzel JN, Blazer KR, MacDonald DJ, Culver JO, Offit K (2011) Genetics, genomics and cancer risk assessment: state of the art and future directions in the era of personalized medicine. CA Cancer J Clin 61(5):327–359. doi:10.3322/caac.20128
Amundadottir LT (2016) Pancreatic cancer genetics. Int J Biol Sci 12(3):314–325. doi:10.7150/ijbs.15001
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25(14):1754–1760. doi:10.1093/bioinformatics/btp324
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25(16):2078–2079. doi:10.1093/bioinformatics/btp352
COSMIC (2016) Catalogue of somatic mutations in cancer. http://cancer.sanger.ac.uk/wgs/browse/tissue. Accessed 13 July 2016
National Center for Biotechnology Information (NCBI) (2016) dbSNP. http://www.ncbi.nlm.nih.gov/projects/SNP/. Accessed 15 Sept 2016
1000 Genomes Project (2010) 1000 Genomes: a deep catalog of human genetic variation. http://www.1000genomes.org/. Accessed 01 March 2011
National Heart, Lung, and Blood Institute (NHLBI) (2016) NHLBI Grand Opportunity Exome Sequencing Project (ESP). https://esp.gs.washington.edu/drupal/. Accessed 15 Sept 2016
JASPAR Database (2016) JASPAR. http://jaspar.genereg.net/. Accessed 15 Sept 2016
Siepel A, Bejerano G, Pedersen JS, Hinrichs AS, Hou M, Rosenbloom K, Clawson H, Spieth J, Hillier LW, Richards S, Weinstock GM, Wilson RK, Gibbs RA, Kent WJ, Miller W, Haussler D (2005) Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res 15(8):1034–1050. doi:10.1101/gr.3715005
Pollard KS, Hubisz MJ, Rosenbloom KR, Siepel A (2010) Detection of nonneutral substitution rates on mammalian phylogenies. Genome Res 20(1):110–121. doi:10.1101/gr.097857.109
VISTA Comparative Genomics Analysis (2016) VISTA enhancer browser. http://enhancer.lbl.gov/. Accessed 15 Sept 2016
Complete Genomics (2016) Complete Genomics. http://www.completegenomics.com/. Accessed 20 Sept 2016
Sim NL, Kumar P, Hu J, Henikoff S, Schneider G, Ng PC (2012) SIFT web server: predicting effects of amino acid substitutions on proteins. Nucleic Acids Res 40(Web Server issue):W452–W457. doi:10.1093/nar/gks539
The Cancer Genome Atlas (2011). http://cancergenome.nih.gov. Accessed 26 Feb 2011
Adzhubei I, Jordan DM, Sunyaev SR (2013) Predicting functional effect of human missense mutations using PolyPhen-2. Curr Protoc Hum Genet Chapter 7:Unit7 20. doi:10.1002/0471142905.hg0720s76
Lek M, Karczewski KJ, Minikel EV, Samocha KE, Banks E, Fennell T, O’Donnell-Luria AH, Ware JS, Hill AJ, Cummings BB, Tukiainen T, Birnbaum DP, Kosmicki JA, Duncan LE, Estrada K, Zhao F, Zou J, Pierce-Hoffman E, Berghout J, Cooper DN, Deflaux N, DePristo M, Do R, Flannick J, Fromer M, Gauthier L, Goldstein J, Gupta N, Howrigan D, Kiezun A, Kurki MI, Moonshine AL, Natarajan P, Orozco L, Peloso GM, Poplin R, Rivas MA, Ruano-Rubio V, Rose SA, Ruderfer DM, Shakir K, Stenson PD, Stevens C, Thomas BP, Tiao G, Tusie-Luna MT, Weisburd B, Won H-H, Yu D, Altshuler DM, Ardissino D, Boehnke M, Danesh J, Donnelly S, Elosua R, Florez JC, Gabriel SB, Getz G, Glatt SJ, Hultman CM, Kathiresan S, Laakso M, McCarroll S, McCarthy MI, McGovern D, McPherson R, Neale BM, Palotie A, Purcell SM, Saleheen D, Scharf JM, Sklar P, Sullivan PF, Tuomilehto J, Tsuang MT, Watkins HC, Wilson JG, Daly MJ, MacArthur DG, Exome Aggregation Consortium (2016) Analysis of protein-coding genetic variation in 60,706 humans. Nature 536 (7616):285–291. doi:10.1038/nature19057. http://www.nature.com/nature/journal/v536/n7616/abs/nature19057.html#supplementary-information
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, Grody WW, Hegde M, Lyon E, Spector E, Voelkerding K, Rehm HL (2015) Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. doi:10.1038/gim.2015.30
Landrum MJ, Lee JM, Riley GR, Jang W, Rubinstein WS, Church DM, Maglott DR (2014) ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res 42(Database issue):D980–D985. doi:10.1093/nar/gkt1113
Schwartz MK (1999) Genetic testing and the clinical laboratory improvement amendments of 1988: present and future. Clin Chem 45(5):739–745
Klein AP, Brune KA, Petersen GM, Goggins M, Tersmette AC, Offerhaus GJ, Griffin C, Cameron JL, Yeo CJ, Kern S, Hruban RH (2004) Prospective risk of pancreatic cancer in familial pancreatic cancer kindreds. Cancer Res 64(7):2634–2638
Witkiewicz AK, McMillan EA, Balaji U, Baek G, Lin WC, Mansour J, Mollaee M, Wagner KU, Koduru P, Yopp A, Choti MA, Yeo CJ, McCue P, White MA, Knudsen ES (2015) Whole-exome sequencing of pancreatic cancer defines genetic diversity and therapeutic targets. Nat Commun 6:6744. doi:10.1038/ncomms7744
Bailey P, Chang DK, Nones K, Johns AL, Patch AM, Gingras MC, Miller DK, Christ AN, Bruxner TJ, Quinn MC, Nourse C, Murtaugh LC, Harliwong I, Idrisoglu S, Manning S, Nourbakhsh E, Wani S, Fink L, Holmes O, Chin V, Anderson MJ, Kazakoff S, Leonard C, Newell F, Waddell N, Wood S, Xu Q, Wilson PJ, Cloonan N, Kassahn KS, Taylor D, Quek K, Robertson A, Pantano L, Mincarelli L, Sanchez LN, Evers L, Wu J, Pinese M, Cowley MJ, Jones MD, Colvin EK, Nagrial AM, Humphrey ES, Chantrill LA, Mawson A, Humphris J, Chou A, Pajic M, Scarlett CJ, Pinho AV, Giry-Laterriere M, Rooman I, Samra JS, Kench JG, Lovell JA, Merrett ND, Toon CW, Epari K, Nguyen NQ, Barbour A, Zeps N, Moran-Jones K, Jamieson NB, Graham JS, Duthie F, Oien K, Hair J, Grutzmann R, Maitra A, Iacobuzio-Donahue CA, Wolfgang CL, Morgan RA, Lawlor RT, Corbo V, Bassi C, Rusev B, Capelli P, Salvia R, Tortora G, Mukhopadhyay D, Petersen GM, Munzy DM, Fisher WE, Karim SA, Eshleman JR, Hruban RH, Pilarsky C, Morton JP, Sansom OJ, Scarpa A, Musgrove EA, Bailey UM, Hofmann O, Sutherland RL, Wheeler DA, Gill AJ, Gibbs RA, Pearson JV, Waddell N, Biankin AV, Grimmond SM (2016) Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 531(7592):47–52. doi:10.1038/nature16965
Biankin AV, Waddell N, Kassahn KS, Gingras MC, Muthuswamy LB, Johns AL, Miller DK, Wilson PJ, Patch AM, Wu J, Chang DK, Cowley MJ, Gardiner BB, Song S, Harliwong I, Idrisoglu S, Nourse C, Nourbakhsh E, Manning S, Wani S, Gongora M, Pajic M, Scarlett CJ, Gill AJ, Pinho AV, Rooman I, Anderson M, Holmes O, Leonard C, Taylor D, Wood S, Xu Q, Nones K, Fink JL, Christ A, Bruxner T, Cloonan N, Kolle G, Newell F, Pinese M, Mead RS, Humphris JL, Kaplan W, Jones MD, Colvin EK, Nagrial AM, Humphrey ES, Chou A, Chin VT, Chantrill LA, Mawson A, Samra JS, Kench JG, Lovell JA, Daly RJ, Merrett ND, Toon C, Epari K, Nguyen NQ, Barbour A, Zeps N, Kakkar N, Zhao F, Wu YQ, Wang M, Muzny DM, Fisher WE, Brunicardi FC, Hodges SE, Reid JG, Drummond J, Chang K, Han Y, Lewis LR, Dinh H, Buhay CJ, Beck T, Timms L, Sam M, Begley K, Brown A, Pai D, Panchal A, Buchner N, De Borja R, Denroche RE, Yung CK, Serra S, Onetto N, Mukhopadhyay D, Tsao MS, Shaw PA, Petersen GM, Gallinger S, Hruban RH, Maitra A, Iacobuzio-Donahue CA, Schulick RD, Wolfgang CL, Morgan RA, Lawlor RT, Capelli P, Corbo V, Scardoni M, Tortora G, Tempero MA, Mann KM, Jenkins NA, Perez-Mancera PA, Adams DJ, Largaespada DA, Wessels LF, Rust AG, Stein LD, Tuveson DA, Copeland NG, Musgrove EA, Scarpa A, Eshleman JR, Hudson TJ, Sutherland RL, Wheeler DA, Pearson JV, McPherson JD, Gibbs RA, Grimmond SM (2012) Pancreatic cancer genomes reveal aberrations in axon guidance pathway genes. Nature 491(7424):399–405. doi:10.1038/nature11547
Barber LJ, Rosa Rosa JM, Kozarewa I, Fenwick K, Assiotis I, Mitsopoulos C, Sims D, Hakas J, Zvelebil M, Lord CJ, Ashworth A (2011) Comprehensive genomic analysis of a BRCA2 deficient human pancreatic cancer. PLoS ONE 6(7):e21639. doi:10.1371/journal.pone.0021639
Kohlmann W, Gruber SB (1993–2014) Lynch syndrome. In: Pagon RA, Adam MP, Ardinger HH, et al. (eds) GeneReviews™ [Internet]
Canto MI, Harinck F, Hruban RH, Offerhaus GJ, Poley JW, Kamel I, Nio Y, Schulick RS, Bassi C, Kluijt I, Levy MJ, Chak A, Fockens P, Goggins M, Bruno M (2013) International Cancer of the Pancreas Screening (CAPS) Consortium summit on the management of patients with increased risk for familial pancreatic cancer. Gut 62(3):339–347. doi:10.1136/gutjnl-2012-303108
Javle M, Golan T, Maitra A (2016) Changing the course of pancreatic cancer—focus on recent translational advances. Cancer Treat Rev 44:17–25. doi:10.1016/j.ctrv.2016.01.004
Michl J, Zimmer J, Tarsounas M (2016) Interplay between Fanconi anemia and homologous recombination pathways in genome integrity. EMBO J 35(9):909–923. doi:10.15252/embj.201693860
NCCN (2016) NCCN guidelines genetic/familial high-risk assessment: breast and ovarian V.2.2016.
Sawyer SL, Tian L, Kahkonen M, Schwartzentruber J, Kircher M, Majewski J, Dyment DA, Innes AM, Boycott KM, Moreau LA, Moilanen JS, Greenberg RA (2015) Biallelic mutations in BRCA1 cause a new Fanconi anemia subtype. Cancer Discov 5(2):135–142. doi:10.1158/2159-8290.cd-14-1156
Zhen DB, Rabe KG, Gallinger S, Syngal S, Schwartz AG, Goggins MG, Hruban RH, Cote ML, McWilliams RR, Roberts NJ, Cannon-Albright LA, Li D, Moyes K, Wenstrup RJ, Hartman A-R, Seminara D, Klein AP, Petersen GM (2015) BRCA1, BRCA2, PALB2, and CDKN2A mutations in familial pancreatic cancer: a PACGENE study. Genet Med 17(7):569–577. doi:10.1038/gim.2014.153
Roberts NJ, Norris AL, Petersen GM, Bondy ML, Brand R, Gallinger S, Kurtz RC, Olson SH, Rustgi AK, Schwartz AG, Stoffel E, Syngal S, Zogopoulos G, Ali SZ, Axilbund J, Chaffee KG, Chen YC, Cote ML, Childs EJ, Douville C, Goes FS, Herman JM, Iacobuzio-Donahue C, Kramer M, Makohon-Moore A, McCombie RW, McMahon KW, Niknafs N, Parla J, Pirooznia M, Potash JB, Rhim AD, Smith AL, Wang Y, Wolfgang CL, Wood LD, Zandi PP, Goggins M, Karchin R, Eshleman JR, Papadopoulos N, Kinzler KW, Vogelstein B, Hruban RH, Klein AP (2016) Whole genome sequencing defines the genetic heterogeneity of familial pancreatic cancer. Cancer Discov 6 (2):166–175. doi:10.1158/2159-8290.cd-15-0402
Smith AL, Alirezaie N, Connor A, Chan-Seng-Yue M, Grant R, Selander I, Bascunana C, Borgida A, Hall A, Whelan T, Holter S, McPherson T, Cleary S, Petersen GM, Omeroglu A, Saloustros E, McPherson J, Stein LD, Foulkes WD, Majewski J, Gallinger S, Zogopoulos G (2016) Candidate DNA repair susceptibility genes identified by exome sequencing in high-risk pancreatic cancer. Cancer Lett 370(2):302–312. doi:10.1016/j.canlet.2015.10.030
Mathew CG (2006) Fanconi anaemia genes and susceptibility to cancer. Oncogene 25(43):5875–5884
van der Heijden MS, Brody JR, Gallmeier E, Cunningham SC, Dezentje DA, Shen D, Hruban RH, Kern SE (2004) Functional defects in the fanconi anemia pathway in pancreatic cancer cells. Am J Pathol 165(2):651–657. doi:10.1016/s0002-9440(10)63329-9
Couch FJ, Johnson MR, Rabe K, Boardman L, McWilliams R, de Andrade M, Petersen G (2005) Germ line Fanconi anemia complementation group C mutations and pancreatic cancer. Cancer Res 65(2):383–386
Rogers CD, van der Heijden MS, Brune K, Yeo CJ, Hruban RH, Kern SE, Goggins M (2004) The genetics of FANCC and FANCG in familial pancreatic cancer. Cancer Biol Ther 3(2):167–169
van der Heijden MS, Yeo CJ, Hruban RH, Kern SE (2003) Fanconi anemia gene mutations in young-onset pancreatic cancer. Cancer Res 63(10):2585–2588
Oddoux C, Struewing JP, Clayton CM, Neuhausen S, Brody LC, Kaback M, al e (1996) The carrier frequency of the BRCA2 6174delT mutation among Ashkenazi Jewish individuals is approximately 1%. Nat Genet 14:188–190
Devgan SS, Sanal O, Doil C, Nakamura K, Nahas SA, Pettijohn K, Bartek J, Lukas C, Lukas J, Gatti RA (2011) Homozygous deficiency of ubiquitin-ligase ring-finger protein RNF168 mimics the radiosensitivity syndrome of ataxia-telangiectasia. Cell Death Differ 18(9):1500–1506. doi:10.1038/cdd.2011.18
O’Driscoll M, Cerosaletti KM, Girard PM, Dai Y, Stumm M, Kysela B, Hirsch B, Gennery A, Palmer SE, Seidel J, Gatti RA, Varon R, Oettinger MA, Neitzel H, Jeggo PA, Concannon P (2001) DNA ligase IV mutations identified in patients exhibiting developmental delay and immunodeficiency. Mol Cell 8(6):1175–1185
Li D, Suzuki H, Liu B, Morris J, Liu J, Okazaki T, Li Y, Chang P, Abbruzzese JL (2009) DNA repair gene polymorphisms and risk of pancreatic cancer. Clin Cancer Res 15(2):740–746. doi:10.1158/1078-0432.ccr-08-1607
Chun SG, Yee NS, Holland JM, Shohet RV, Palalay MP, Bryant-Greenwood PK (2011) Pancreatic Adenocarcinoma Associated With Werner’s Syndrome (Adult-Onset Progeria). Gastrointest Cancer Res 4(1):24–28
Alemar B, Herzog J, Netto CBO, Artigalas O, Schwartz IV, Bittar CM, Ashton-Prolla P, Weitzel JN (2016) Prevalence of Hispanic BRCA1 and BRCA2 mutations among hereditary breast and ovarian cancer patients from Brazil reveals differences among Latin American populations. Cancer Genet 209:417–422. doi:10.1016/j.cancergen.2016.06.008
Theis B, Boyd N, Lockwood G, Tritchler D (1994) Accuracy of family cancer history in breast cancer patients. Eur J Cancer Prev 3(4):321–327
Acknowledgements
The research reported in this publication was supported by the National Cancer Institute (NCI) of the National Institutes of Health (NIH) under award number P30CA33572 (Integrative Genomics and Bioinformatics Cores). The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH. The City of Hope Clinical Cancer Genomics Community Research Network and the Hereditary Cancer Research Registry was supported in part by the NCI NIH award number RC4CA153828 (PI: J. Weitzel). Other sources of support include: Breast Cancer Research Foundation (PI: J. Weitzel), Morris and Horowitz Families Professor (S. Neuhausen), 2015 STOP CANCER Research Career Development Award (PI: T. Slavin), and the Oxnard Foundation (PI: T. Slavin). We would like to thank all sites that contributed research effort to the Clinical Cancer Genomics Community Research Network as well as the patients who allow this research to be completed. We would like to thank Drs. Yuan Chun Ding and Yuan Yate-Ching for help accessing informatics resources. We would also like to thank the following research assistants: Tanya Chavez, Lily Van Tongeren, and Rosa Mejia.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
All procedure performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Slavin, T.P., Neuhausen, S.L., Nehoray, B. et al. The spectrum of genetic variants in hereditary pancreatic cancer includes Fanconi anemia genes. Familial Cancer 17, 235–245 (2018). https://doi.org/10.1007/s10689-017-0019-5
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10689-017-0019-5