Abstract
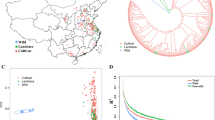
Soybean originated in ancient China has been quickly extended globally as a major protein and oil crop. The QTL–allele constitution of seed protein content (SPC) in the Chinese soybean landrace population (CSLRP) was studied using a representative sample composed of 365 accessions tested under multiple environments and analysed under the novel restricted two-stage multi-locus genome-wide association study (RTM-GWAS) procedure based on 29,121 SNPLDB (single nucleotide polymorphism linkage disequilibrium blocks) markers. The SPC varied from 37.51 to 50.46% among accessions, for which 89 QTLs, each with 2–9 alleles in a total of 255 alleles were identified, accounting for 83.16% of the phenotypic variation covering most of the genetic variation (h2 = 84.31%). The QTL–alleles of the 365 landraces were organized into a 255 × 365 QTL–allele matrix as the compact form of SPC genetic constitution in CSLRP. Of the 89 QTLs, 53 showed significantly differentiated allele frequency distribution patterns among geographic eco-regions (sub-populations). There were 32.09% alleles not common among sub-populations but found only in some sub-populations; new allele(s) emerged on some loci in some respective sub-populations, with Eco-region III showing less but Eco-region VI more emergence. The QTL–allele matrix was also used for prediction of optimal crosses for breeding purpose to reach a 99th percentile potential of up to 54.81%, more than the highest accession (50.46%). From the 89 QTLs, 59 SPC candidate genes involving biological processes, cellular components and molecular functions were annotated. Among them, Glyma18g13574 and Glyma20g21370 were inferred as two of the major SPC genes in the whole genome.



Similar content being viewed by others
References
Bolon YT, Joseph B, Cannon SB et al (2010) Complementary genetic and genomic approaches help characterize the linkage group I seed protein QTL in soybean. BMC Plant Biol 10:41
Brummer EC, Graef GL, Orf J, Wilcox JR, Shoemaker RC (1997) Mapping QTL for seed protein and oil content in eight soybean populations. Crop Sci 37:370–378
Cai SG, Yu G, Chen XH et al (2013) Grain protein content variation and its association analysis in barley. BMC Plant Biol 13:35
Chung J, Babka HL, Graef GL et al (2003) The seed protein, oil, and yield QTL on soybean linkage group I. Crop Sci 43:1053–1067
Diers BW, Keim P, Fehr WR, Shoemaker RC (1992) RFLP analysis of soybean seed protein and oil content. Theor Appl Genetics 83:608–612
Du Z, Zhou X, Ling Y, Zhang Z, Su Z (2010) Agrigo: a GO analysis toolkit for the agricultural community. Nucleic Acids Res 38:W64–W70
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Res 10:564–567
Flint-Garcia SA, Thuillet A, Yu J et al (2005) Maize association population: a high-resolution platform for quantitative trait locus dissection. Plant J 44:1054–1064
Gai J, Wang Y (2001) A study on the varietal eco-regions of soybeans in China. Sci Agric Sin 34:139–145
Gai J, Chen L, Zhang Y, Zhao T, Xing G, Xing H (2012) Genome-wide genetic dissection of germplasm resources and implications for breeding by design in soybean. Breed Sci 61:495–510
Hamblin MT, Jannink J-L (2011) Factors affecting the power of haplotype markers in association studies. Plant Genome 4:145–153
Hanson CH, Robinson HF, Comstock RE (1956) Biometrical studies of yield in segregating populations of Korean. Lespedeza Agron J 48:268–272
He J, Shan M, Zhao T et al (2017) An innovative procedure of genome-wide association analysis fits studies on germplasm population and plant breeding. Theor Appl Genet 130:2327–2343
Hwang E-Y, Song Q, Jia G et al (2014) A genome-wide association study of seed protein and oil content in soybean. BMC Genom 15:1–12
Jun TH, Van K, Kim MY, Lee SH, Walker DR (2008) Association analysis using SSR markers to find QTL for seed protein content in soybean. Euphytica 162:179–191
Korir PC, Qi B, Wang Y et al (2011) A study on relative importance of additive, epistasis and unmapped QTL for Aluminium tolerance at seedling stage in soybean. Plant Breed 130:551–562
Larsson SJ, Lipka AE, Buckler ES (2013) Lessons from Dwarf8 on the strengths and weaknesses of structured association mapping. PLoS Genet 9:e1003246
Li S, Cao Y, He J, Zhao T, Gai J (2017) Detecting the QTL–allele system conferring flowering date in a nested association mapping population of soybean using a novel procedure. Theor Appl Genet 130:2297–2314
Meng S, He J, Zhao T et al (2016) Detecting the QTL–allele system of seed isoflavone content in Chinese soybean landrace population for optimal cross design and gene system exploration. Theor Appl Genet 129:1557–1576
Nichols DM, Glover KD, Carlson SR, Specht JE, Diers BW (2006) Fine mapping of a seed protein QTL on soybean linkage group I and its correlated effects on agronomic traits. Crop Sci 46:834–839
Qiu LJ, Xing LL, Guo Y, Wang J, Jackson SA, Chang RZ (2013) A platform for soybean molecular breeding: the utilization of core collections for food security. Plant Mol Biol 83:41–50
Sebolt AM, Shoemaker RC, Diers BW (2000) Analysis of a quantitative trait locus allele from wild soybean that increases seed protein concentration in soybean. Crop Sci 40:1438–1444
Sonah H, O’Donoughue L, Cober E, Rajcan I, Belzile F (2015) Identification of loci governing eight agronomic traits using a GBS-GWAS approach and validation by QTL mapping in soya bean. Plant Biotechnol J 13:211–221
Tajuddin T, Watanabe S, Yamanaka N, Harada K (2003) Analysis of quantitative trait loci for protein and lipid contents in soybean seeds using recombinant inbred lines. Breed Sci 53:133–140
Thornsberry JM, Goodman MM, Doebley J, Kresovich S, Nielsen D, Buckler ES (2001) Dwarf8 polymorphisms associate with variation in flowering time. Nat Genet 28:286–289
Uauy C, Distelfeld A, Fahima T, Blechl A, Dubcovsky J (2006) A NAC Gene regulating senescence improves grain protein, zinc, and iron content in wheat. Science 314:1298–1301
Valliyodan B, Dan Q, Patil G et al (2016) Landscape of genomic diversity and trait discovery in soybean. Sci Rep 6:23598
Wang HW (2011) Genetic dissection and elite allele identification of seed traits in soybean culivars released from Huang-huai valleys and southern China. Nanjing Agricultural University, Nanjing
Wen ZX, Zhao TJ, Zheng YZ et al (2008) Association analysis of agronomic and quality traits with SSR markers in Glycine max and Glycine soja in China: I. Population structure and associated markers. Acta Agron Sin 34:1169–1178
Wen Z, Boyse JF, Song Q, Cregan PB, Wang D (2015) Genomic consequences of selection and genome-wide association mapping in soybean. BMC Genom 16:1–14
Xiong D, Zhao T, Gai J (2008) Parental analysis of soybean cultivars released in China. Sci Agric Sin 41:2589–2598
Zeng A, Chen P, Korth K et al (2017) Genome-wide association study (GWAS) of salt tolerance in worldwide soybean germplasm lines. Mol Breed 37:30
Zhang D, Kan G, Hu Z et al (2014) Use of single nucleotide polymorphisms and haplotypes to identify genomic regions associated with protein content and water-soluble protein content in soybean. Theor Appl Genetics 127:1905–1915
Zhang YH, He JB, Wang YF et al (2015a) Establishment of a 100-seed weight quantitative trait locus-allele matrix of the germplasm population for optimal recombination design in soybean breeding programmes. J Exp Bot 66:6311–6325
Zhang YH, Liu MF, He JB et al (2015b) Marker-assisted breeding for transgressive seed protein content in soybean [Glycine max (L.) Merr.]. Theor Appl Genet 128:1061–1072
Zhang D, Lü H, Chu S et al (2017) The genetic architecture of water-soluble protein content and its genetic relationship to total protein content in soybean. Sci Reports 7:5053
Zhao K, Tung CW, Eizenga GC et al (2011) Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nat Commun 2:467
Zhu C, Gore M, Buckler ES, Yu J (2008) Status and prospects of association mapping in plants. Plant Genome 1:5–20
Acknowledgements
This work was supported by the National Key R & D Program for Crop Breeding in China (2017YFD0101500, 2016YFD0100304), the Natural Science Foundation of China (31701452, 31701447, 31671718, 31571695), the MOE 111 Project (B08025), the MOE Program for Changjiang Scholars and Innovative Research Team in University (PCSIRT_17R55), the MOA CARS-04 program, the Jiangsu Higher Education PAPD Program, the Fundamental Research Funds for the Central Universities and the Jiangsu JCIC-MCP, the open funds of the State Key Laboratory of Crop Genetics and Germplasm Enhancement (ZW201712). The funders had no role in work design, data collection and analysis, and decision and preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
YHZ, TJZ and JYG designed the study; YHZ, SM, GNX and JYY conducted the field experiment; YHZ, MFL, SPY and YL conducted the lab work; JBH developed the molecular data analysis procedure; YHZ and JYG drafted the manuscript.
Corresponding author
Ethics declarations
Conflict of interests
The authors declare no competing interests or conflicts of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, Y., He, J., Meng, S. et al. Identifying QTL–allele system of seed protein content in Chinese soybean landraces for population differentiation studies and optimal cross predictions. Euphytica 214, 157 (2018). https://doi.org/10.1007/s10681-018-2235-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-018-2235-y