Abstract
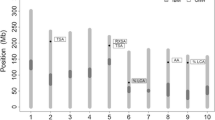
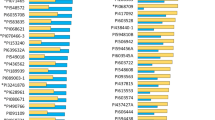
Soybean research has found that nodule traits, especially nodule biomass, are associated with N2 fixation ability. Two genotypes, differing in nodule number per plant and individual nodule weight, KS4895 and Jackson, were mated to create 17 F3- and 80 F5-derived RILs. The population was mapped with 664 informative markers with an average distance of less than 20 cM between adjacent markers. Nodule traits were evaluated in 3-year field trials. Broad-sense heritability for nodule number (no. plant−1), individual nodule dry weight (mg nodule−1), individual nodule size (mm nodule−1), and total nodule dry weight (g plant−1) was 0.41, 0.42, 0.45, and 0.27, respectively. Nodule number was negatively correlated with individual nodule weight and size. Nodule number, individual nodule weight, and size are major components which likely contributed to increased total nodule weight per plant. Composite interval mapping (CIM) identified eight QTLs for nodule number with R2 values ranging from 0.14 to 0.20. Multiple interval mapping (MIM) identified two QTLs for nodule number, one of which was located close to the QTL identified with CIM. Six QTLs for individual nodule weight were detected with CIM, and one QTL was identified with MIM. For nodule size, CIM identified seven QTLs with R2 values ranging from 0.14 to 0.27. Five QTLs for total nodule weight were detected with CIM, one of which was located close to a QTL identified with MIM. These results document the first QTL information on nodule traits in soybean from field experiments utilizing a dense, complete linkage map.



Similar content being viewed by others
Abbreviations
- C.I.:
-
Confidence interval
- CIM:
-
Composite interval mapping
- LG:
-
Linkage group
- LS:
-
Least square means
- LOD:
-
Logarithm of odds
- LRT:
-
Likelihood ratio test
- MIM:
-
Multiple interval mapping
- QTL:
-
Quantitative trait loci
- RIL:
-
Recombinant inbred line
- SSR:
-
Simple sequence repeat
- SNP:
-
Single nucleotide polymorphism
References
Akkaya MS, Shoemaker RC, Specht JE, Bhagwat AA, Cregan PB (1995) Integration of simple sequence repeat DNA markers into a soybean linkage map. Crop Sci 35:1439–1445
Arahana VS, Graef GL, Specht JE, Steadman JR, Eskridge KM (2001) Identification of QTLs for Resistance to Sclerotinia sclerotiorum in Soybean. Crop Sci 41:180–188
Bourion V, Rizvi SMH, Fournier S, Larambergue HD, Galmiche F, Marget P, Duc G, Burstin J (2010) Genetic dissection of nitrogen nutrition in pea through a QTL approach of root, nodule, and shoot variability. Theor Appl Genet 121:71–86
Broman KW, Wu H, Sen S, Churchill GA (2003) R/qtl: QTL mapping in experimental crosses. Bioinformatics 19:889–890
Burias N, Planchon C (1990) Increasing soybean productivity through selection for nitrogen fixation. Agron J 82:1031–1034
Caetano-Annolès G, Gresshoff PM (1991) Plant genetic control of nodulation. Ann Rev Microbiol 45:345–382
Charlson DV, Bhatnagar S, King CA, Ray JD, Sneller CH, Carter TE Jr, Purcell LC (2009) Polygenic inheritance of canopy wilting in soybean [Glycine max (L.) Merr.]. Theor Appl Genet 119(4):587–594
Chung J, Babka HL, Graef GL, Staswick PE, Lee DJ, Cregan PB, Shoemaker RC, Specht JE (2003) The seed protein, oil, and yield QTL on soybean linkage group I. Crop Sci 43:1053–1067
Churchill GA, Doerge RW (1994) Empirical threshold values for quantitative trait mapping. Genetics 138:967–971
Csanadi G, Vollmann J, Stift G, Lelley T (2001) Seed quality QTLs identified in a molecular map of early maturing soybean. Theor Appl Genet 103:912–919
Du W, Wang M, Fu S, Yu D (2009) Mapping WTLs for seed yield and drought susceptiblity index in soybean (Glycine max L.) across different environments. J Genet Genomics 36:721–731
Fehr WR, Caviness CE (1977) Stages of soybean development. Coop Ext Serv Spec Rep 80. Iowa State University, Ames
Gage DJ (2009) Nodule development in legumes. In: Emerich DW, Krishnan HB (eds) Nitrogen fixation in crop production. ASA, CSSA, SSSA, Madison, pp 1–24
Greder RR, Orf JH, Lambert JW (1986) Heritabilities and associations of nodule mass and recovery of Bradyrhizobium japonicum serogroup USDA 110 in soybean. Crop Sci 26:33–37
Hanson WD, Leffel RC, Howell RW (1961) Genetic analysis of energy production in the soybean. Crop Sci 1:121–126
Harper JE (1987) Nitrogen metabolism. In: Wilcox JR (ed) Soybeans: Improvement, production, and uses. 2nd edn. Agron Monogr 16, ASA, CSSA, SSSA, Madison, WI, p 497–533
Helms TC, Orf JH (1998) Protein, oil and yield in soybean lines selected for increased protein. Crop Sci 38:707–711
Hwang S, King CA, Davies MK, Ray JD, Cregan PB, Purcell LC (2013) QTL analysis of shoot ureide and nitrogen concentrations in soybean. Crop Sci (in press)
Hyten DL, Pantalone VR, Sams CE, Saxton AM, Landau-Ellis D, Stefaniak TR, Schmidt ME (2004) Seed quality QTL in a prominent soybean population. Theor Appl Genet 109:552–561
Hyten DL, Choi IY, Song Q, Specht JE, Carter TE Jr, Shoemaker RC, Hwang EY, Matukumalli LK, Cregan PB (2010) A high density integrated genetic linkage map of soybean and the development of a 1536 universal soy linkage panel for quantitative trait locus mapping. Crop Sci 50:1–9
Indrasumunar A, Kereszt A, Searle I, Miyagi M, Li D, Nguyen CDT, Men A, Carroll BJ, Gresshoff PM (2010) Inactivation of duplicated Nod factor receptor 5 (NFR5) genes in recessive loss-of-function non-nodulation mutants of allotetraploid soybean (Glycine max L. Merr.). Plant Cell Physiol 51:201–214
Indrasumunar A, Searle I, Lin MH, Kereszt A, Men A, Carroll BJ, Gresshoff PM (2011) Nodulation factor receptor kinase 1α controls nodule organ number in soybean (Glycine max L. Merr). Plant J 65:39–50
Jiang C, Zeng Z-B (1995) Multiple trait analysis and genetic mapping for quantitative trait loci. Genetics 136:1447–1455
Johnson HW (1958) Registration of soybean varieties, VII. Agron J 11:659–660
Kabelka EA, Diers BW, Fehr WR, LeRoy AR, Baianu IC, You T, Neece DJ, Nelson RL (2004) Putative alleles for increased yield from soybean plant introductions. Crop Sci 44:784–791
Kao C-H, Zeng Z-B, Teasdale RD (1999) Multiple interval mapping for quantitative trait loci. Genetics 152:1203–1216
Kim H, Kang S, Cho J, Choung M, Suh D (2005) Quantitative trait loci associated with oligosaccharide and sucrose contents in soybean (Glycine max L.). J Plant Biol 48:106–112
King CA, Purcell LC (2001) Soybean nodule size and relationship to nitrogen fixation response to water deficit. Crop Sci 41:1099–1107
Knapp SJ, Stroup WW, Ross WM (1985) Exact confidence intervals for heritability on a progeny mean basis. Crop Sci 25:192–194
Komatsu K, Okuda S, Takahashi M, Matsunaga R, Nakazawa Y (2005) QTL mapping of antibiosis resistance to common cutworm (Spodoptera litura Fabricius) in soybean. Crop Sci 45:2044–2048
Lander ES, Green P, Abrahamson J, Barlow A, Daly MJ, Lincoln SE, Newburg L (1987) Mapmaker an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Manly KF, Cudmore RH Jr, Meer JM (2001) Map Manager QTX, cross-platform program software for genetic mapping. Mamm Genome 12:930–932
Mastrodomenico AT, Purcell LC (2012) Soybean nitrogen fixation and nitrogen remobilization during reproductive development. Crop Sci 52:1281–1289
Meng X-L, Rubin DB (1993) Maximum likelihood estimation via the ECM algorithm: a general framework. Biometrika 80:267–278
Narvel JM, Walker DR, Rector BG, All JN, Parrott WA, Boerma HR (2001) A retrospective DNA marker assessment of the development of insect resistant soybean. Crop Sci 41:1931–1939
Nicolás MF, Hungria M, Arias CAA (2006) Identification of quantitative trait loci controlling nodulation and shoot mass in progenies from two Brazillian soybean cultivars. Field Crops Res 95:355–366
Orf JH, Chase K, Jarvik T, Mansur LM, Cregan PB, Adler FR, Lark KG (1999) Genetics of soybean agronomic traits: I. Comparison of three related recombinant inbred populations. Crop Sci 39:1642–1651
Panthee DR, Kwanyuen P, Sams CE, West DR, Saxton AM, Pantalone VR (2004) Quantitative trait loci for B-conglycinin (7S) and glycinin (11S) fractions of soybean storage protein. J Am Oil Chem Soc 81:1005–1012
Panthee DR, Pantalone VR, Saxton AM, West DR, Sams CE (2006) Genomic regions associated with amino acid composition in soybean. Mol Breed 17:79–89
Pazdernik DL, Graham PH, Vance CP, Orf JH (1996) Host genetic variation in the early nodulation and dinitrogen fixation of soybean. Crop Sci 36:1102–1107
Piepho HP, Gauch HG (2001) Marker pair selection for mapping quantitative trait loci. Genetics 157:433–444
Primomo V, Poysa V, Ablett G, Jackson C, Gijzen M, Rajcan I (2005) Mapping QTL for individual and total isoflavone content in soybean seeds. Crop Sci 45:2464–2545
Purcell LC (2009) Physiological responses of N2 fixation to drought and selecting genotypes for improved N2 fixation. In: Emerich DW, Krishnan HB (eds) Nitrogen fixation in crop production. ASA, CSSA, SSSA, Madison, pp 211–238
Purcell LC, deSilva M, King CA, Kim WH (1997) Biomass accumulation and allocation in soybean associated with genotypic differences in tolerance of nitrogen fixation to water deficits. Plant Soil 196:101–113
Purcell LC, King CA, Ball RA (2000) Soybean cultivar differences in ureides, drought tolerant nitrogen fixation, and manganese nutrition. Crop Sci 40:1062–1070
Purcell LC, Edwards JT, Brye KR (2007) Soybean yield and biomass responses to cumulative transpiration: questioning widely held beliefs. Field Crops Res 101:10–18
Qi Z, Wu Q, Han X, Sun Y, Du X, Liu C, Jiang H, Hu G, Chen Q (2011) Soybean oil content QTL mapping and integrating with meta-analysis method for mining genes. Euphytica 179:499–514
Reinprecht Y, Poysa V, Yu K, Rajcan I, Ablett G, Pauls K (2006) Seed and agronomic QTL in low linolenic acid, lipoxygenase-free soybean (Glycine max (L.) Merrill) germplasm. Genome 49:1510–1527
Sall K, Sinclair TR (1991) Soybean genotypic differences in sensitivity of symbiotic nitrogen fixation to soil dehydration. Plant Soil 133:31–37
Santos MA, Geraldi IO, Garcia AAF, Bortolatti N, Schiavon A, Hungria M (2013) Mapping of QTLs associated with biological nitrogen fixation traits in soybean. Hereditas 150:17–25
Schapaugh WT, Dille RE (1998) Registration of ‘KS4895′ soybean. Crop Sci 38:892
Schnabel E, Journet EP, de Carvalho-Niebel F, Duc G, Frugoli J (2005) The Medicago truncatula SUNN gene encodes a CLV1-like leucine-rich repeat receptor kinase that regulates nodule number and root length. Plant Mol Biol 58:809–822
Searle IR, Men AE, Laniya TS, Buzas DM, Iturbe-Ormaetxe I, Carroll BJ, Gresshoff PM (2003) Long-distance signaling in nodulation directed by a CLAVATA1-like receptor kinase. Science 299:109–112
Sen S, Satagopan JM, Broman KW, Churchill GA (2007) R/qtldesign: inbred line cross experimental design. Mamm Genome 18:87–93
Sinclair TR, deWitt C (1975) Photosynthate and nitrogen requirements for seed production by various crops. Science 189:565–567
Sinclair TR, Soffes AR, Hinson K, Albrecht SL, Pfahler PL (1991) Genotypic variation in soybean nodule number and weight. Crop Sci 31:301–304
Singleton PW, Stockinger KR (1983) Compensation against ineffective nodulation in soybean. Crop Sci 23:69–72
Tanya P, Srinives P, Toojinda T, Vanavichit A, Lee SH (2005) Identification of SSR markers associated with N2-fixation component in soybean [Glycine max (L.) Merr.]. Korean J Genetics 27:351–359
Tasma IM, Lorenzen LL, Green DE, Shoemaker RC (2001) Mapping genetic loci for flowering time, maturity, and photoperiod insensitivity in soybean. Mol Breed 8(1):25–35
Terry LI, Chase K, Jarvik T, Orf JH, Mansur LM, Lark KG (2000) Soybean quantitative trait loci for resistance to insects. Crop Sci 40(2):375–382
Thorne JC, Fehr WR (1970) Incorporation of high-protein, exotic germplasm into soybean populations by 2- and 3-way crosses. Crop Sci 10:652–655
Tischner T, Allphin L, Chase K, Orf JH, Lark KG (2003) Genetics of seed abortion and reproductive traits in soybean. Crop Sci 43:464–473
Tominaga A, Gondo T, Akashi R, Zheng S-H, Arima S, Suzuki A (2012) Quatitative trait locus analysis of symbiotic nitrogen fixation activity in the model legume Lotus japonicus. J Plant Res 125:395–406
Wang CS (1994) Bayesian analysis of mixed linear models via Gibbs sampling with an application to litter size in Iberian pigs. Genet Sel Evol 26:91–115
Weller JI (1986) Maximum likelihood techniques for the mapping and analysis of quantitative trait loci with the aid of genetic markers. Biometrics 42:627–640
Wilcox JR, Cavins JF (1995) Backcrossing high seed protein to a soybean cultivar. Crop Sci 35:1036–1041
Williams LF, Lynch DL (1954) Inheritance of a non-nodulation character in the soybean. Agron J 46:28–29
Yang J, Zhu J, Williams RW (2007) Mapping the genetic architecture of complex traits in experimental populations. Bioinformatics 23:1527–1536
Yang J, Hu C, Hu H, Yu R, Xia Z, Ye X, Zhu J (2008) QTL Network: mapping and visualizing genetic architecture of complex traits in experimental populations. Bioinformatics 24:721–723
Zeng Z-B (1994) Precise mapping of quantitative trait loci. Genetics 136:1457–1468
Acknowledgments
The authors gratefully acknowledge partial financial support for this research from the United Soybean Board and from the United States Department of Agriculture-Agriculture Research Service (USDA-ARS) project number 6402-21220-012-00D. Editorial assistance from Penny McGee is greatly appreciated.
Author information
Authors and Affiliations
Corresponding author
Additional information
Mention of a trademark or proprietary product does not constitute a guarantee or warranty of the product by the US Department of Agriculture and does not imply approval or the exclusion of other products that may also be suitable. USDA is an equal opportunity provider and employer.
Rights and permissions
About this article
Cite this article
Hwang, S., Ray, J.D., Cregan, P.B. et al. Genetics and mapping of quantitative traits for nodule number, weight, and size in soybean (Glycine max L.[Merr.]). Euphytica 195, 419–434 (2014). https://doi.org/10.1007/s10681-013-1005-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-013-1005-0