Abstract
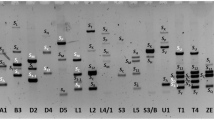
The Latvian and the Swedish sweet cherry (Prunus avium L.) genetic resources collections comprise valuable material for breeding. The collections represent local Latvian and Scandinavian genetic resources: semi-wild samples, landraces, and cultivars developed in local breeding programmes, as well as diverse germplasm from the northern temperate zone. The objective of this investigation was to determine which S 1 –S 6 alleles are most important in the sweet cherry genetic resources collections and to compare the identified allelic and genotypic frequencies in material of different origin. Accessions in the two collections were screened for the presence of the self-incompatibility (S) S 1 to S 6 alleles, using PCR based typing. Significant differences (P < 0.05) between screened collections were found in frequencies of S 4 and S 5 alleles. Analysis of allele combinations identified the high occurrence of selections with the S-genotype S 3 S 6 in both collections. Compared to the S-allele frequencies published for over 250 sweet cherry cultivars from Western and Southern Europe, the Latvian and Swedish germplasm appeared to have a high frequency of the S 6 allele in both collections, and a relatively high frequency of the S 5 allele in Latvian germplasm. This study represents the first comprehensive S-allele screening for the sweet cherry genetic resources collections in Latvia and Sweden. Both sweet cherry collections contain high proportion of accessions adapted to north central European growing conditions, not typical for the majority of the documented sweet cherry genetic resources, which explains differences in certain S-allele occurrence.

Similar content being viewed by others
References
Blukmanis M, Ikase L, Kaufmane E, Ruisa S, Strautina S, Skrivele M, Rashal I (1997) Peteris Upitis (1896–1976), Horticulturist and Breeder. Proc Latvian Acad Sci 51(1/2):88–91
Bošković R, Tobutt KR (2001) Genotyping cherry cultivars assigned to incompatibility groups, by analysing stylar ribonucleases. Theor Appl Genet 103:475–485
Crane MB, Lawrence WJC (1929) Genetical and cytological aspects of incompatibility and sterility in cultivated fruits. J Pomol Hort Sci 7:276–301
Crane MB, Brown AG (1937) Incompatibility and sterility in the sweet cherry Prunus avium L. J Pomol 15:86–116
de Nettancourt D (1977) Incompatibility in angiosperms. Sex Plant Reprod 10:185–199
Hauck NR, Iezzoni AF, Yamane H, Tao R (2001) Revising the S-allele Nomenclature in Sweet Cherry (Prunus avium) using RFLP profiles. J Am Soc Hort Sci 126:654–660
Nybom H, Schaal BA (1990) DNA ‘fingerprints’ reveal genotypic distributions in natural populations of blackberries and raspberries (Rubus, Rosaceae). Am J Bot 44:883–888
Rashal I, Lacis G (1999) Accessions of horticultural plants in the Latvian plant genetic resources data base. In: Ikase L, Skrivele M, Rasals I, Ruisa S, Trajkovski V (eds) Fruit growing today and tomorrow. Proceedings of the International Conference, Dobele, pp 124–130
Ruisa S (1998) Genetic resources of Latvian sweet cherries and their use. Acta Horticulturae 468(1):153–160
Sassa H, Nishio T, Kowyama Y, Hirano H, Koba T, Ikehashi H (1996) Self- incompatibility (S) alleles of the Rosaceae encode members of a distinct class of the T2/S ribonuclease superfamily. Mol Gen Genet 250:547–557
Sonneveld T, Robbins TP, Boškovič R Tobutt KR (2001) Cloning of six cherry self- incompatibility alleles and development of allele specific PCR detection. Theor Appl Genet 102:1046–1055
Sonneveld T, Tobutt KR, Robbins TP (2003) Allele-specific PCR detection of sweet cherry self-incompatibilty (S) alleles S1 to S16 using consensus and allele-specific primers. Theor Appl Genet 107:1059–1070
Tao R, Yamane H, Sugiura A, Murayama H, Sassa H, Mori H (1999) Molecular typing of S-alleles through identification, characterization and cDNA cloning for S-RNases in sweet cherry. J Am Soc Hort Sci 124:224–233
Tehrani G, Brown SK (1992) Pollen-incompatibility and self-fertility in sweet cherry. Plant Breeding Rev 9:368–370
Tobutt KR, Sonneveld T, Bekefi T, Bošković R (2004) Cherry (in)compatibility genotypes – an updated cultivar table. Acta Horticulturae 663:667–671
Trajkovski V (1996) A review of the cherry breeding program in Sweden. Acta Horticulturae 410:387–388
Ushijima K, Sassa H, Dandekar AM, Gradziel TM, Tao R, Hirano H (2003) Structural and transcriptional analysis of the self-incompatibility locus of almond: identification of a pollen-expressed F-Box gene with haplotype-specific polymorphism. The Plant Cell 15:71–781
Wiersma PA, Wu Z, Zhou L, Hampson C, Kappel F (2001) Identification of new self- incompatibility alleles in sweet cherry (Prunus avium L.) and clarification of incompatibility groups by PCR and sequencing analysis. Theor Appl Genet 102:700–708
Acknowledgements
This research was enabled by the support from the Swedish Research Council for Environment, Agricultural Sciences and Spatial Planning, the Royal Swedish Academy of Agriculture and Forestry, the Swedish Institute (SI), the Swedish Council for Forestry and Agricultural Research (SJFR) and the Knut and Alice Wallenberg Foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lacis, G., Kaufmane, E., Rashal, I. et al. Identification of self-incompatibility (S) alleles in Latvian and Swedish sweet cherry genetic resources collections by PCR based typing. Euphytica 160, 155–163 (2008). https://doi.org/10.1007/s10681-007-9496-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-007-9496-1