Abstract
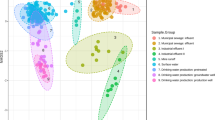
Pyrosequencing targeting the V1–V3 hypervariable of the 16S rDNA was used to investigate the bacterial diversity in river and roof-harvested rainwater (RHRW) used for potable purposes by rural households in Luthengele village in the Eastern Cape Province of South Africa. The phylum Proteobacteria dominated the data set (80.5 % of all reads), while 4.2 % of the reads could not be classified to any of the known phyla at a probability of 0.8 or higher (unclassified bacteria). At class level, the classes; Betaproteobacteria (50.4 % of all reads), Alphaproteobacteria (16.2 %), Verrucomicrobiae (6.6 %), Planctomycetacia (5.7 %), and Sphingobacteria (3 %) dominated the data set in all the samples. Although the class Verrucomicrobiae constituted 6.6 % of all sequences, 88.6 % of the sequences were from the river sample where the class represented 43.7 % of the observed sequences in the sample. The bacteria community structure clearly showed significant similarities between RHRW and differences with the river water control sample, suggesting different levels of contamination and environmental factors affecting the various water sources. Moreover, signatures of potential pathogens including Legionella, Acinetobacter, Pseudomonas, Clostridia, Chromobacterium, Yersinia, and Serratia were detected, and the proportions of Legionella were relatively higher suggesting a potential health risk to households using RHRW. This work provides guidance for prioritizing subsequent culturable and quantitative analysis to ensure that potentially significant pathogens are not left out of risk estimations.









Similar content being viewed by others
References
Ahmed, W., Huygens, F., Goonetilleke, A., & Gardner, T. (2008). Real-Time PCR detection of pathogenic microorganisms in roof-harvested rainwater in Southeast Queensland, Australia. Applied and Environmental Microbiology, 74(17), 5490–5496.
Ahmed, W., Goonetilleke, A., & Gardner, T. (2010a). Implications of faecal indicator bacteria for the microbiological assessment of roof-harvested rainwater quality in southeast Queensland, Australia. Canadian Journal of Microbiology, 56(6), 471–479.
Ahmed, W., Vieritz, A., Goonetilleke, A., & Gardner, T. (2010b). Health risk from the use of roof-harvested rainwater in Southeast Queensland, Australia, as potable or nonpotable water, determined using quantitative microbial risk assessment. Applied and Environmental Microbiology, 76(22), 7382–7391.
Ahmed, W., Gardner, T., & Toze, S. (2011). Microbiological quality of roof-harvested rainwater and health risks: a review. Journal of Environmental Quality, 40(1), 13–21.
Ahmed, W., Sidhu, J. P. S., Toze, S., & Hodgers, L. (2012a). Fecal indicators and zoonotic pathogens in household drinking water taps fed from rainwater tanks in Southeast Queensland. Applied and Environmental Microbiology, 78(1), 219–226.
Ahmed, W., Sidhu, J. P. S., & Toze, S. (2012b). An attempt to identify the likely sources of Escherichia coli harboring toxin genes in rainwater tanks. Environmental Science & Technology, 46(9), 5193–5197.
Ashbolt, N. J. (2004). Microbial contamination of drinking water and disease outcomes in developing regions. Toxicology, 198(1), 229–238.
Bibby, K., Viau, E., & Peccia, J. (2010). Pyrosequencing of the 16S rRNA gene to reveal bacterial pathogen diversity in biosolids. Water Research, 44(14), 4252–4260.
Bricheux, G., Morin, L., Moal, G., Coffe, G., Balestrino, D., Charbonnel, N., et al. (2013). Pyrosequencing assessment of prokaryotic and eukaryotic diversity in biofilm communities from a French river. Microbiology Open, 2(3), 402–414.
Caporaso, J. G., Kuczynski, J., Stombaugh, J., Bittinger, K., Bushman, F. D., Costello, E. K., et al. (2010). QIIME allows analysis of high-throughput community sequencing data. Nature Methods, 7(5), 335–336.
Chen, C. L., Liu, W. T., Chong, M. L., Wong, M. T., Ong, S. L., Seah, H., et al. (2004). Community structure of microbial biofilms associated with membrane-based water purification processes as revealed using a polyphasic approach. Applied Microbiology and Biotechnology, 63(4), 466–473.
Cleary, D. F. R., Smalla, K., Mendonc, L. C. S., & Gomes, N. C. M. (2012). Assessment of variation in bacterial composition among microhabitats in a mangrove environment using DGGE fingerprints and barcoded pyrosequencing. PloS One, 7(1), e29380.
Costa, J., Tiago, I., da Costa, M. S., & Veríssimo, A. (2005). Presence and persistence of Legionella spp. in ground-water. Applied and Environmental Microbiology, 71(2), 663–671.
Costello, E. K., Lauber, C. L., Hamady, M., Fierer, N., Gordon, J. I., & Knight, R. (2009). Bacterial community variation in human body habitats across space and time. Science, 326(5960), 1694–1697.
Cottrell, M. T., Waidner, L. A., Yu, L., & Kirchman, D. L. (2005). Bacterial diversity of metagenomic and PCR libraries from the Delaware River. Environmental Microbiology, 7(12), 1883–1895.
DeSantis, T. Z., Brodie, E. L., Moberg, J. P., Zubieta, I. X., Piceno, Y. M., & Andersen, G. L. (2007). High-density universal 16S rRNA microarray analysis reveals broader diversity than typical clone library when sampling the environment. Microbial Ecology, 53(3), 371–383.
Dolan, J. R. (2005). Biogeography of aquatic microbes. Aquatic Microbial Ecology, 41, 39–48.
Eichler, S., Christen, R., Höltje, C., Westphal, P., Bötel, J., Brettar, I., et al. (2006). Composition and dynamics of bacterial communities of a drinking water supply system as assessed by RNA-and DNA-based 16S rRNA gene fingerprinting. Applied and Environmental Microbiology, 72(3), 1858–1872.
Fields, B. S., Benson, R. F., & Besser, R. E. (2002). Legionella and Legionnaires’ disease: 25 years of investigation. Clinical Microbiology Reviews, 15(3), 506–526.
Gich, F., Schubert, K., Bruns, A., Hoffelner, H., & Overmann, J. (2005). Specific detection, isolation, and characterization of selected, previously uncultured members of the freshwater bacterio-plankton community. Applied and Environmental Microbiology, 71(10), 5908–5919.
Hong, P. Y., Hwang, C., Ling, F., Andersen, G. L., Le Chevallier, M. W., Liu, W., et al. (2010). Pyrosequencing analysis of bacterial biofilm communities in water meters of a drinking water distribution system. Applied and Environmental Microbiology, 76(16), 5631–5635.
Hur, I., & Chun, J. (2004). A method for comparing multiple bacterial community structures from 16S rDNA clone library sequences. Journal of Microbiology, 42(1), 9–13.
Janssen, P. H. (2006). Identifying the dominant soil bacterial taxa in libraries of 16S rRNA and 16S rRNA genes. Applied and Environmental Microbiology, 72(3), 1719–1728.
Jukes, T. H., & Cantor, C. R. (1969). Evolution of protein molecules, Munro HN, Mammalian protein metabolism (pp. 21–132). New York: Academic Press.
Kahinda, J. M., Taigbenu, A. E., & Boroto, R. J. (2010). Domestic rainwater harvesting as an adaptation measure to climate change in South Africa. Physics and Chemistry of the Earth Parts ABC, 35, 742–751.
Kaushik, R., Balasubramanian, R., & de la Cruz, A. A. (2012). Influence of Air Quality on the Composition of Microbial Pathogens in Fresh Rainwater. Applied and Environmental Microbiology, 78(8), 2813–2818.
Kaushik, R., Balasubramanian, R., & Dunstan, H. (2014). Microbial Quality and Phylogenetic Diversity of Fresh Rainwater and Tropical Freshwater Reservoir. PLoS One, 9(6), e100737.
Kirchman, D. L. (2002). The ecology of Cytophaga-Flavobacteria in aquatic environments. FEMS Microbiology Ecology, 39(2), 91–100.
Lautenschlager, K., Hwang, C., Liu, W., Boon, N., Ko, O., Vrouwenvelder, H., et al. (2013). A microbiology-based multi-parametric approach towards assessing biological stability in drinking water distribution networks. Water Research, 47, 3015–3025.
Lindström, E. S., Kamst-Van Agterveld, M. P., & Zwart, G. (2005). Distribution of typical freshwater bacterial groups is associated with pH, temperature, and lake water retention time. Applied and Environmental Microbiology, 71(12), 8201–8206.
Lozupone, C. A., Hamady, M., Kelley, S. T., & Knight, R. (2007). Quantitative and qualitative β diversity measures lead to different insights into factors that structure microbial communities. Applied and Environmental Microbiology, 73(5), 1576–1585.
Lozupone, C., Lladser, M. E., Knights, D., Stombaugh, J., & Knight, R. (2010). UniFrac: An effective distance metric for microbial community comparison. The ISME Journal, 5(2), 169–172.
Luxmy, B., Nakajima, F., & Yamamoto, K. (2000). Analysis of bacterial community in membrane-separation bioreactors by fluorescent in situ hybridization (FISH) and denaturing gradient gel-electrophoresis (DGGE) techniques. Water Science and Technology, 41(10–11), 259–268.
Lye, D. J. (2009). Rooftop runoff as a source of contamination : A review. Science of the Total Environment, 407(21), 5429–5434.
MacLean, D., Jones, J. D. G., & Studholme, D. J. (2009). Application of next-generation’ sequencing technologies to microbial genetics. Nature Reviews Microbiology, 7(4), 287–296.
Margulies, M., Egholm, M., Altman, W. E., Attiya, S., Bader, J. S., Bemben, L. A., et al. (2005). Genome sequencing in micro-fabricated high-density picolitre reactors. Nature, 437(7057), 376–380.
Martiny, A. C., Albrechtsen, H.-J., Arvin, E., & Molin, S. (2005). Identification of bacteria in biofilm and bulk water samples from a non-chlorinated model drinking water distribution system: detection of a large nitrite-oxidizing population associated with Nitrospira spp. Applied and Environmental Microbiology, 71(12), 8611–8617.
Matcher, G. F., Dorrington, R. A., Henninger, T. O., & Froneman, P. W. (2011). Insights into the bacterial diversity in a freshwater-deprived permanently open Eastern Cape estuary, using 16S rRNA pyrosequencing analysis. Water SA, 37(3), 381–390.
Meays, C. L., Broersma, K., Nordin, R., & Mazumder, A. (2004). Source tracking faecal bacteria in water: A critical review of current methods. Journal of Environmental Management, 73(1), 71–79.
Metzker, M. L. (2009). Sequencing technologies: the next generation. Nature Reviews Genetics, 11(1), 31–46.
Muder, R. R., & Yu, V. L. (2002). Infection due to Legionella species other than L. pneumophila. Clinical Infectious Diseases, 35, 990–998.
Murga, R., Forster, T. S., Brown, E., Pruckler, J. M., Fields, B. S., & Donlan, R. M. (2001). Role of biofilms in the survival of Legionella pneumophila in a model potable-water system. Microbiology, 147(11), 3121–3126.
Muyzer, G., De Waal, E. C., & Uitterlinden, A. G. (1993). Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Applied and Environmental Microbiology, 59(3), 695–700.
Navarro-Noya, Y. E., Suárez-Arriaga, M. C., Rojas-Valdes, A., Montoya-Ciriaco, N. M., Gómez-Acata, S., Fernández-Luqueño, F., & Dendooven, L. (2013). Pyrosequencing Analysis of the Bacterial Community in Drinking Water Wells. Microbial Ecology, 66(1), 19–29.
Nocker, A., Burr, M., & Camper, A. K. (2007). Genotypic microbial community profiling: A critical technical review. Microbial Ecology, 54(2), 276–289.
Palmer, C. J., Tsai, Y. L., Paszko-Kolva, C., Mayer, C., & Sangermano, L. R. (1993). Detection of Legionella species in sewage and ocean water by polymerase chain reaction, direct fluorescent-antibody, and plate culture methods. Applied and Environmental Microbiology, 59(11), 3618–3624.
Pinto, A. J., Xi, C., & Raskin, L. (2012). Bacterial community structure in the drinking water microbiome is governed by filtration processes. Environmental Science & Technology, 46(16), 8851–8859.
Poitelon, J.-B., Joyeux, M., Welté, B., Duguet, J.-P., Prestel, E., Lespinet, O., et al. (2009). Assessment of phylogenetic diversity of bacterial microflora in drinking water using serial analysis of ribosomal sequence tags. Water Research, 43(17), 4197–4206.
Rasid, R. A. (2009). Biofilm and multimedia filtration for rainwater treatment. Journal of Sustainable Development, 2(1), 196–199.
Roesch, L. F. W., Fulthorpe, R. R., Riva, A., Casella, G., Hadwin, A. K. M., Kent, A. D., & Triplett, E. W. (2007). Pyrosequencing enumerates and contrasts soil microbial diversity. The ISME Journal, 1(4), 283–290.
Rowbotham, T. J. (1986). Current views on the relationships between amoebae, legionellae and man. Israel Journal of Medical Sciences, 22(9), 678.
Rusin, P. A., Rose, J. B., Haas, C. N., & Gerba, C. P. (1997). Risk assessment of opportunistic bacterial pathogens in drinking water. In Reviews of Environmental Contamination and Toxicology (pp. 57–83). New York: Springer.
Schlesner, H., Jenkins, C., & Staley, J. T. (2006). The phylum Verrucomicrobia: A phylogenetically heterogeneous bacterial group. In The Prokaryotes (pp. 881–896). New York: Springer.
Shanks, O. C., White, K., Kelty, C. A., Sivaganesan, M., Blannon, J., Meckes, M., et al. (2010). Performance of PCR-based assays targeting Bacteroidales genetic markers of human faecal pollution in sewage and faecal samples. Environmental Science and Technology, 44, 6281.
Shi, P., Jia, S., Zhang, X. X., Zhang, T., Cheng, S., & Li, A. (2013). Metagenomic insights into chlorination effects on microbial antibiotic resistance in drinking water. Water Research, 47(1), 111–120.
Sinclair, R., Boone, S. A., Greenberg, D., Keim, P., & Gerba, C. P. (2008). Persistence of category A select agents in the environment. Applied and Environmental Microbiology, 74(3), 555–563.
Sogin, M. L., Morrison, H. G., Huber, J. A., Welch, D. M., Huse, S. M., Neal, P. R., & Herndl, G. J. (2006). Microbial diversity in the deep sea and the underexplored “rare biosphere”. Proceedings of the National Academy of Sciences, 103(32), 12115–12120.
Steelman, S. M., Chowdhary, B. P., Dowd, S., Suchodolski, J., & Jane, J. E. (2012). Pyrosequencing of 16S rRNA genes in faecal samples reveals high diversity of hindgut micro-flora in horses and potential links to chronic laminitis. BMC Veterinary Research, 8(231), 1–11.
Stocker, S., Snajder, R., Rainer, J., Trajanoski, S., Gorkiewicz, G., Trajanoski, Z., & Thallinger, G. G. (2011). SnoWMAn: high-throughput phylotyping, analysis and comparison of microbial communities. In Proceedings of the 110th ASM General Meeting 23-5-2010.
Stout, J. E., Yu, V. L., & Best, M. G. (1985). Ecology of Legionella pneumophila within water distribution systems. Applied and Environmental Microbiology, 49(1), 221–228.
Tamura, K., Peterson, D., Peterson, N., Stecher, G., Nei, M., & Kumar, S. (2011). MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Molecular Biology and Evolution, 28(10), 2731–2739.
US EPA. (2009). Drinking water contaminant candidate List 3. Federal Register, 74(194), 51850–51862.
Vaz-Moreira, I., Egas, C., Nunes, O. C., & Manaia, C. M. (2011). Culture-dependent and culture-independent diversity surveys target different bacteria: a case study in a freshwater sample. Antonie Van Leeuwenhoek, 100(2), 245–257.
Ward, N., Staley, J. T., Fuerst, J. A., Giovannoni, S., Schlesner, H., & Stackebrandt, E. (2006). The order Planctomycetales, including the genera Planctomyces, Pirellula, Gemmata and Isosphaera and the Candidatus genera Brocadia, Kuenenia and Scalindua. In M. Dworkin, S. Falkow, E. Rosenberg, K. H. Schleifer & E. Stackebrandt (Eds.). The Prokaryotes: A Handbook on the Biology of Bacteria (757–793). New York: Springer.
Weisburg, W. G., Barns, S. M., Pelletier, D. A., & Lane, D. J. (1991). 16S ribosomal DNA amplification for phylogenetic study. Journal of Bacteriology, 173(2), 697–703.
Zwart, G., Crump, B. C., Kamst-van Agterveld, M. P., Hagen, F., & Han, S. K. (2002). Typical freshwater bacteria: an analysis of available 16S rRNA gene sequences from plankton of lakes and rivers. Aquatic Microbial Ecology, 28(2), 141–155.
Acknowledgments
This study was undertaken as part of a Water Research Commission (WRC) unsolicited project: “Evaluation of the risks associated with the use of rainwater harvested from roofs, for domestic use and, homestead food gardens; and groundwater for domestic use and livestock watering” (WRC Project No K5/2175, Water Research Commission, 2013).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chidamba, L., Korsten, L. Pyrosequencing analysis of roof-harvested rainwater and river water used for domestic purposes in Luthengele village in the Eastern Cape Province of South Africa. Environ Monit Assess 187, 41 (2015). https://doi.org/10.1007/s10661-014-4237-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10661-014-4237-0