Abstract
Background
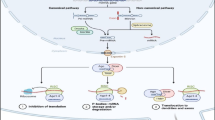
Competitive endogenous RNA (ceRNA) regulation suggested complex network of all transcript RNAs including long noncoding RNAs (lncRNAs), which can act as natural miRNA sponges to inhibit miRNA functions and modulate mRNA expression. Until now, the specific ceRNA regulatory mechanism of lncRNA–miRNA–mRNA in colorectal cancer (CRC) still remains unclear.
Materials and Methods
RNA sequencing data of 478 colon adenocarcinoma cases and 41 controls as well as 166 rectum adenocarcinoma cases and 10 controls were obtained from The Cancer Genome Atlas (TCGA) to investigate the significant changes of lncRNAs, miRNAs and mRNAs in colorectal carcinogenesis. The target lncRNAs and mRNAs of miRNAs were predicted by miRWalk. Functional and enrichment analyses were conducted by DAVID database. The lncRNA–miRNA–mRNA interaction network was constructed using Cytoscape.
Results
We constructed ceRNA regulatory networks including 22 up-regulated lncRNAs, 12 down-regulated miRNAs and 122 up-regulated mRNAs, as well as 8 down-regulated lncRNAs, 43 up-regulated miRNAs and 139 down-regulated mRNAs. The GO enrichment showed that up-regulated genes mainly enriched in biological process including organic anion transport, collagen catabolic process, wound healing, Wnt receptor signalling and in pathways of tyrosine metabolism, taurine and hypotaurine metabolism, melanogenesis and phenylalanine metabolism. For down-regulated genes, significant enrichment was found in biological process of metal ion homeostasis, transmission of nerve impulse, cell–cell signalling, transmembrane transport and in pathways of ABC transporters, neuroactive ligand–receptor interaction, retinol metabolism, nitrogen metabolism and steroid hormone biosynthesis.
Conclusion
We identified significantly altered lncRNAs, miRNAs and mRNAs in colorectal carcinogenesis, which might serve as potential biomarkers for tumorigenesis of CRC. In addition, the ceRNA regulatory network of lncRNA–miRNA–mRNA was constructed, which would elucidate novel molecular mechanisms involved in initiation and progression of CRC, thus providing promising clues for clinical diagnosis and therapy.




Similar content being viewed by others
References
Anastasiadou E, Jacob LS, Slack FJ. Non-coding RNA networks in cancer. Nat Rev Cancer. 2018;18:5–18.
Esteller M, Pandolfi PP. The epitranscriptome of noncoding RNAs in Cancer. Cancer Discov. 2017;7:359–368.
Bhan A, Soleimani M, Mandal SS. Long noncoding RNA and cancer: a new paradigm. Cancer Res. 2017;77:3965–3981.
Bolha L, Ravnik-Glavac M, Glavac D. Long noncoding RNAs as biomarkers in cancer. Dis Mark. 2017;2017:7243968.
Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell. 2011;146:353–358.
Giroud M, Scheideler M. Long non-coding RNAs in metabolic organs and energy homeostasis. Int J Mol Sci. 2017;18:2578.
Hu X, Sood AK, Dang CV, Zhang L. The role of long noncoding RNAs in cancer: the dark matter matters. Curr Opin Genet Dev. 2017;48:8–15.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2015. CA Cancer J Clin. 2015;65:5–29.
Donadon M, Ribero D, Morris-Stiff G, Abdalla EK, Vauthey JN. New paradigm in the management of liver-only metastases from colorectal cancer. Gastrointest Cancer Res GCR. 2007;1:20–27.
Li B, Shi C, Zhao J, Li B. Long noncoding RNA CCAT1 functions as a ceRNA to antagonize the effect of miR-410 on the down-regulation of ITPKB in human HCT-116 and HCT-8 cells. Oncotarget. 2017;8:92855–92863.
Chen DL, Lu YX, Zhang JX, et al. Long non-coding RNA UICLM promotes colorectal cancer liver metastasis by acting as a ceRNA for microRNA-215 to regulate ZEB2 expression. Theranostics. 2017;7:4836–4849.
Xu J, Zhang R, Zhao J. The novel long noncoding RNA TUSC7 inhibits proliferation by sponging MiR-211 in colorectal cancer. Cell Physiol Biochem Int J Exp Cell Physiol Biochem Pharmacol. 2017;41:635–644.
Zhou XG, Huang XL, Liang SY, et al. Identifying miRNA and gene modules of colon cancer associated with pathological stage by weighted gene co-expression network analysis. OncoTargets Ther. 2018;11:2815–2830.
Wei HT, Guo EN, Liao XW, et al. Genomescale analysis to identify potential prognostic microRNA biomarkers for predicting overall survival in patients with colon adenocarcinoma. Oncol Rep. 2018;40:1947–1958.
Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature. 2012;490:61–70.
Dweep H, Sticht C, Pandey P, Gretz N. miRWalk–database: prediction of possible miRNA binding sites by “walking” the genes of three genomes. J Biomed Inform. 2011;44:839–847.
Dweep H, Gretz N, Sticht C. miRWalk database for miRNA–target interactions. Methods Mol Biol (Clifton NJ). 2014;1182:289–305.
Dennis G Jr, Sherman BT, Hosack DA, et al. DAVID: database for annotation, visualization, and integrated discovery. Genome Biol. 2003;4:P3.
The Gene Ontology (GO) project in 2006. Nucleic Acids Research. 2006;34:D322–326.
Kanehisa M, Goto S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000;28:27–30.
Lin C, Yang L. Long noncoding RNA in cancer: wiring signaling circuitry. Trends Cell Biol. 2017;28:287–301.
Renganathan A, Felley-Bosco E. Long noncoding RNAs in cancer and therapeutic potential. Adv Exp Med Biol. 2017;1008:199–222.
Slaby O, Laga R, Sedlacek O. Therapeutic targeting of non-coding RNAs in cancer. Biochem J. 2017;474:4219–4251.
Sun W, Yang Y, Xu C, Guo J. Regulatory mechanisms of long noncoding RNAs on gene expression in cancers. Cancer Genet. 2017;216–217:105–110.
Zhang J, Jiang Y, Zhu J, et al. Overexpression of long non-coding RNA colon cancer-associated transcript 2 is associated with advanced tumor progression and poor prognosis in patients with colorectal cancer. Oncol Lett. 2017;14:6907–6914.
Ling H, Spizzo R, Atlasi Y, et al. CCAT2, a novel noncoding RNA mapping to 8q24, underlies metastatic progression and chromosomal instability in colon cancer. Genome Res. 2013;23:1446–1461.
Han Y, Yang YN, Yuan HH, et al. UCA1, a long non-coding RNA up-regulated in colorectal cancer influences cell proliferation, apoptosis and cell cycle distribution. Pathology. 2014;46:396–401.
Ni B, Yu X, Guo X, et al. Increased urothelial cancer associated 1 is associated with tumor proliferation and metastasis and predicts poor prognosis in colorectal cancer. Int J Oncol. 2015;47:1329–1338.
Rupaimoole R, Slack FJ. MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat Rev Drug Discov. 2017;16:203–222.
Mima K, Nishihara R, Yang J, et al. MicroRNA MIR21 (miR-21) and PTGS2 expression in colorectal cancer and patient survival. Clin Cancer Res. 2016;22:3841–3848.
Ding M, Zhang T, Li S, Zhang Y, Qiu Y, Zhang B. Correlation analysis between liver metastasis and serum levels of miR200 and miR141 in patients with colorectal cancer. Mol Med Rep. 2017;16:7791–7795.
Amado NG, Predes D, Moreno MM, Carvalho IO, Mendes FA, Abreu JG. Flavonoids and Wnt/beta-catenin signaling: potential role in colorectal cancer therapies. Int J Mol Sci. 2014;15:12094–12106.
Masuda M, Sawa M, Yamada T. Therapeutic targets in the Wnt signaling pathway: feasibility of targeting TNIK in colorectal cancer. Pharmacol Ther. 2015;156:1–9.
Xue X, Taylor M, Anderson E, et al. Hypoxia-inducible factor-2alpha activation promotes colorectal cancer progression by dysregulating iron homeostasis. Cancer Res. 2012;72:2285–2293.
Matsumura T, Sugimachi K, Iinuma H, et al. Exosomal microRNA in serum is a novel biomarker of recurrence in human colorectal cancer. Br J Cancer. 2015;113:275–281.
Wang X, Ding X, Nan L, et al. Investigation of the roles of exosomes in colorectal cancer liver metastasis. Oncol Rep. 2015;33:2445–2453.
Carethers JM, Jung BH. Genetics and genetic biomarkers in sporadic colorectal cancer. Gastroenterology. 2015;149:1177–1190 e1173.
Funding
This study is supported by grants from Public Welfare Foundation of Liaoning Province (No. 2015005002).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All of the authors declare that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, J., Li, H., Zheng, B. et al. Competitive Endogenous RNA (ceRNA) Regulation Network of lncRNA–miRNA–mRNA in Colorectal Carcinogenesis. Dig Dis Sci 64, 1868–1877 (2019). https://doi.org/10.1007/s10620-019-05506-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-019-05506-9