Abstract
Background
Circulating microRNA expression profiles may be promising biomarkers for diagnosis and assessment of the prognosis of cancer patients. Quantitative polymerase chain reaction (qPCR) is a sensitive technique for estimating expression levels of circulating microRNAs. However, there is no current consensus on the reference genes for qPCR analysis of circulating microRNAs.
Aims
In this study we tried to identify suitable reference genes for qPCR analysis of serum microRNA in gastric cancer patients and healthy individuals.
Methods
Six microRNAs (let-7a, miR-16, miR-93, miR-103, miR-192, and miR-451) and RNU6B were chosen as candidate reference genes on the basis of the literature. Expression levels of these candidates were analyzed by qPCR in serum samples from 40 gastric cancer patients and 20 healthy volunteers. The geNorm, Normfinder, bestkeeper, and comparative delta-Ct method algorithms were used to select the most suitable reference gene from the seven candidates. This was then validated by normalizing the expression levels of serum miR-21 across all gastric cancer patients and healthy volunteers.
Results
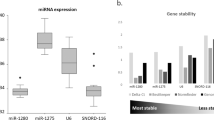
The algorithms revealed miR-16 and miR-93 were the most stably expressed reference genes, with stability values of 1.778 and 2.213, respectively, for serum microRNA analysis across all the patients and healthy controls. The effect of different normalization strategies was compared; when normalized to the serum volume there were no significant differences between patients and controls. However, when the data were normalized to miR-93, miR-16, or miR-93 and miR-16 combined, significant differences were detected.
Conclusions
Our results demonstrated that reference gene choice for qPCR data analysis has a great effect on the study outcome, and that it is necessary to choose a suitable reference for reliable expression data. We recommend miR-16 and miR-93 as suitable reference genes for serum miRNA analysis for gastric cancer patients and healthy controls.




Similar content being viewed by others
References
Esquela-Kerscher A, Slack FJ. Oncomirs: microRNAs with a role in cancer. Nat Rev Cancer. 2006;6:259–269.
Small EM, Frost RJ, Olson EN. MicroRNAs add a new dimension to cardiovascular disease. Circulation. 2010;121:1022–1032.
Kolfschoten IG, Roggli E, Nesca V, Regazzi R. Role and therapeutic potential of microRNAs in diabetes. Diabetes Obes Metab. 2009;11:118–129.
Ng EK, Chong WW, Jin H, et al. Differential expression of microRNAs in plasma of patients with colorectal cancer: a potential marker for colorectal cancer screening. Gut. 2009;58:1375–1381.
Ueda T, Volinia S, Okumura H, et al. Relation between microRNA expression and progression and prognosis of gastric cancer: a microRNA expression analysis. Lancet Oncol. 2010;11:136–146.
Chen X, Ba Y, Ma L, et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008;18:997–1006.
Mitchell PS, Parkin RK, Kroh EM, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci USA. 2008;105:10513–10518.
Kosaka N, Iguchi H, Ochiya T. Circulating microRNA in body fluid: a new potential biomarker for cancer diagnosis and prognosis. Cancer Sci. 2010;101:2087–2092.
Lawrie CH, Gal S, Dunlop HM, et al. Detection of elevated levels of tumour-associated microRNAs in serum of patients with diffuse large B-cell lymphoma. Br J Haematol. 2008;141:672–675.
Tsujiura M, Ichikawa D, Komatsu S, et al. Circulating microRNAs in plasma of patients with gastric cancers. Br J Cancer. 2010;102:1174–1179.
Liu R, Zhang C, Hu Z, et al. A five-microRNA signature identified from genome-wide serum microRNA expression profiling serves as a fingerprint for gastric cancer diagnosis. Eur J Cancer. 2011;47:784–791.
Asaga S, Kuo C, Nguyen T, et al. Direct serum assay for microRNA-21 concentrations in early and advanced breast cancer. Clin Chem. 2011;57:84–91.
Kanemaru H, Fukushima S, Yamashita J, et al. The circulating microRNA-221 level in patients with malignant melanoma as a new tumor marker. J Dermatol Sci. 2011;61:187–193.
Kreth S, Heyn J, Grau S, et al. Identification of valid endogenous control genes for determining gene expression in human glioma. Neuro Oncol. 2010;12:570–579.
Davoren PA, McNeill RE, Lowery AJ, et al. Identification of suitable endogenous control genes for microRNA gene expression analysis in human breast cancer. BMC Mol Biol. 2008;9:76.
Zhu HT, Dong QZ, Wang G et al. (2011) Identification of Suitable Reference Genes for qPCR Analysis of Circulating microRNAs in Hepatitis B Virus-Infected Patients. Mol Biotechnol. (Epub ahead of print). doi:10.1007/s12033-011-9414-6.
McNeill RE, Miller N, Kerin MJ. Evaluation and validation of candidate endogenous control genes for real-time quantitative PCR studies of breast cancer. BMC Mol Biol. 2007;8:107.
Profiling of microRNA in blood serum/plasma. Guidelines for the miRCURY LNATM Universal RT microRNA PCR System. Available at: http://www.exiqon.com/. Accessed 11 July 2011.
Andersen CL, Jensen JL, Orntoft TF. Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 2004;64:5245–5250.
Vandesompele J, De Preter K, Pattyn F et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002; 3: RESEARCH0034.
Pfaffl MW, Tichopad A, Prgomet C, Neuvians TP. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper–Excel-based tool using pair-wise correlations. Biotechnol Lett. 2004;26:509–515.
Thein SL, Silver N, Best S, Jiang J. Selection of housekeeping genes for gene expression studies in human reticulocytes using real-time PCR. BMC Mol Biol. 2006; 7.
Chen C, Ridzon DA, Broomer AJ et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res. 2005; 33: e179.
Tang F, Hajkova P, Barton SC et al. MicroRNA expression profiling of single whole embryonic stem cells. Nucleic Acids Res. 2006; 34: e9.
Peltier HJ, Latham GJ. Normalization of microRNA expression levels in quantitative RT-PCR assays: identification of suitable reference RNA targets in normal and cancerous human solid tissues. RNA. 2008;14:844–852.
Kroh EM, Parkin RK, Mitchell PS, Tewari M. Analysis of circulating microRNA biomarkers in plasma and serum using quantitative reverse transcription-PCR (qPCR). Methods. 2010;50:298–301.
Chan SH, Wu CW, Li AF, et al. miR-21 microRNA expression in human gastric carcinomas and its clinical association. Anticancer Res. 2008;28:907–911.
Si ML, Zhu S, Wu H, et al. miR-21-mediated tumor growth. Oncogene. 2007;26:2799–2803.
Zhang Z, Li Z, Gao C, et al. miR-21 plays a pivotal role in gastric cancer pathogenesis and progression. Lab Invest. 2008;88:1358–1366.
Motoyama K, Inoue H, Mimori K, et al. Clinicopathological and prognostic significance of PDCD4 and microRNA-21 in human gastric cancer. Int J Oncol. 2010;36:1089–1095.
Johnson SM, Grosshans H, Shingara J, et al. RAS is regulated by the let-7 MicroRNA family. Cell. 2005;120:635–647.
Cimmino A, Calin GA, Fabbri M, et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc Natl Acad Sci USA. 2005;102:13944–13949.
Petrocca F, Visone R, Onelli MR, et al. E2F1-regulated microRNAs impair TGFbeta-dependent cell-cycle arrest and apoptosis in gastric cancer. Cancer Cell. 2008;13:272–286.
Piccolo S, Martello G, Rosato A, et al. A MicroRNA targeting dicer for metastasis control. Cell. 2010;141:U1195–U1276.
Ju JF, Song B, Wang Y, et al. miR-192 regulates dihydrofolate reductase and cellular proliferation through the p53-microRNA Circuit. Clin Cancer Res. 2008;14:8080–8086.
Bandres E, Bitarte N, Arias F, et al. microRNA-451 regulates macrophage migration inhibitory factor production and proliferation of gastrointestinal cancer cells. Clin Cancer Res. 2009;15:2281–2290.
Kiss T. Small nucleolar RNA-guided post-transcriptional modification of cellular RNAs. EMBO J. 2001;20:3617–3622.
Conflict of interest
No benefits in any form have been received or will be received from a commercial party related directly or indirectly to the subject of this article.
Author information
Authors and Affiliations
Corresponding author
Additional information
Jianning Song and Zhigang Bai contributed equally to this work.
Rights and permissions
About this article
Cite this article
Song, J., Bai, Z., Han, W. et al. Identification of Suitable Reference Genes for qPCR Analysis of Serum microRNA in Gastric Cancer Patients. Dig Dis Sci 57, 897–904 (2012). https://doi.org/10.1007/s10620-011-1981-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-011-1981-7