Abstract
Background
Liver biopsy remains the gold standard to assess hepatic fibrosis. It is desirable to predict hepatic fibrosis without the need for invasive liver biopsy. Proteomic techniques allow unbiased assessment of proteins and might be useful to identify proteins related to hepatic fibrosis.
Aims
We utilized two different proteomic methods to identify serum proteins as candidate biomarkers to predict hepatic fibrosis stage in patients with chronic hepatitis C virus (HCV) infection.
Methods
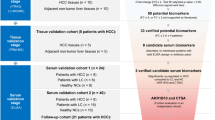
Serum was obtained from 24 people with chronic HCV at time of liver biopsy and from 6 normals. Liver biopsy fibrosis was staged 1–4 (Batts–Ludwig). Pooled serum samples (six in each of four fibrosis groups and controls) were analyzed with 4- and 8-plex isobaric tags for relative and absolute quantitation (iTRAQ), determining protein identification (ID) and ratios of relative protein abundance. Nonpooled samples were analyzed with two-dimensional (2-D) gels and difference in gel electrophoresis (DIGE) comparing different samples on the same gel and across gels. Spots varying among groups were measured with densitometry, excised, digested, and submitted for tandem mass spectrometry (MS/MS) protein ID.
Results
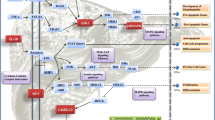
iTRAQ identified 305 proteins (minimum 99% ID confidence); 66 were increased or decreased compared with controls. Some proteins were increased or decreased for specific fibrosis scores. From 704 DIGE protein spots, 66 were chosen, 41 excised, and 135 proteins identified, since one gel spot often identified more than one protein.
Conclusions
Both proteomic methods identified two proteins as biomarker candidates for predicting hepatic fibrosis: complement C4-A and inter-alpha-trypsin inhibitor heavy chain H4.





Similar content being viewed by others
Abbreviations
- iTRAQ:
-
Isobaric tags for relative and absolute quantitation
- DIGE:
-
Difference in gel electrophoresis
- HCV:
-
Hepatitis C virus
References
Liang TJ, Reherman B, Seeff LB, Hoofnagle JH. Pathogenesis, natural history, treatment and prevention of hepatitis C. Ann Intern Med. 2000;132:296–305.
Poynard T, Bedossa P, Opolon P. Natural history of liver fibrosis progression in patients with chronic hepatitis C. Lancet. 1997;349:825–832.
Seaberg EC, Belle SH, Beringer KC, et al. Liver transplantation in the United States from 1987–1998: updated results from the Pitt-UNOS Liver Transplant Registry. In: Cecka M, Terasaki PI, eds. Clinical Transplants. Los Angeles: UCLA Tissue Typing Laboratory; 1998.
Ghany MG, Kleiner DE, Alter H, et al. Progression of fibrosis in chronic hepatitis C. Gastroenterology. 2003;124:97–104.
Neal KR, Ramsay S, Thomson BJ, Irving WL. Excess mortality rates in a cohort of patients infected with the hepatitis C virus: a prospective study. Gut. 2007;56:1098–1104.
Schuppan D, Afdhal NH. Liver cirrhosis. Lancet. 2008;371:838–851.
Poynard T, McHutchison J, Manns M, Myers RP, Albrecht J. Biochemical surrogate markers of liver fibrosis and activity in a randomized trial of peginterferon alfa-2b and ribavirin. Hepatology. 2003;38:481–492.
Thein HH, Yi Q, Dore GJ, Krahn MD. Estimation of stage-specific fibrosis progression rates in chronic hepatitis C virus infection: a meta-analysis and meta-regression. Hepatology. 2008;48:418–431.
Yano M, Kumada H, Kage M, et al. The long-term pathological evolution of chronic hepatitis C. Hepatology. 1996;23:1334–1340.
Janiec DJ, Jacobson ER, Freeth A, Spaulding L, Blaszyk H. Histologic variation of grade and stage of non-alcoholic fatty liver disease in liver biopsies. Obes Surg. 2005;15:497–501.
Piccinino F, Sagnelli E, Pasquale G, Giusti G. Complications following percutaneous liver biopsy. A multicentre retrospective study on 68, 276 biopsies. J Hepatol. 1986;2:165–173.
Anonymous. National institutes of health consensus development conference statement: management of hepatitis C 2002. Gastroenterology. 2002;123:2082–2099.
Guha IN, Parkes J, Roderick P, et al. Noninvasive markers of fibrosis in nonalcoholic fatty liver disease: validating the European liver fibrosis panel and exploring simple markers. Hepatology. 2008;47:455–460.
Popov Y, Schuppan D. Targeting liver fibrosis: strategies for development and validation of antifibrotic therapies. Hepatology. 2009;50:1294–1306.
Thome-Kromer B, Bonk I, Klatt M, et al. Toward the identification of liver toxicity markers: a proteome study in human cell culture and rats. Proteomics. 2003;3:1835–1862.
Marko-Varga G, Berglund M, Malmstrom J, Lindberg H, Fehniger TE. Targeting hepatocytes from liver tissue by laser capture microdissection and proteomics expression profiling. Electrophoresis. 2003;24:3800–3805.
Cyranoski D. China takes centre stage for liver proteome. Nature. 2003;425:441.
Adkins JN, Varnum SM, Auberry KJ, et al. Toward a human blood serum proteome: analysis by multidimensional separation coupled with mass spectrometry. Molecular & Cellular Proteomics. 2002;1:947–955.
Gangadharan B, Antrobus R, Dwek RA, Zitzmann N. Novel serum biomarker candidates for liver fibrosis in hepatitis C patients. Clin Chem. 2007;53:1792–1799.
Burtis CA, Ashwood ER. Tietz Fundamentals of Clinical Chemistry. Philadelphia: W.B. Saunders; 2001.
Wrotnowski C. The future of plasma proteins. Genet Eng News. 1998;18:14.
Anderson NL, Anderson NG. The human plasma proteome: history, character, and diagnostic prospects. Mol Cell Proteomics. 2003;1:845–867.
Marouga R, David S, Hawkins E. The development of the DIGE system: 2D fluorescence difference gel analysis technology. Anal Bioanal Chem. 2005;382:669–678.
Aggarwal K, Choe LH, Lee KH. Shotgun proteomics using the iTRAQ isobaric tags. Brief Funct Genomic Proteomic. 2006;5:112–120.
Batts KP, Ludwig J. Chronic Hepatitis: an update on terminology and reporting. Am J Surg Pathol. 1995;19:1409–1417.
Yates JR III, Eng JK, McCormack AL, Schieltz D. Method to correlate tandem mass spectra of modified peptides to amino acid sequences in the protein database. Anal Chem. 1995;67:1426–1436.
Gasteiger E, Gattiker A, Hoogland C, Ivanyi I, Appel RD, Bairoch A. ExPASy: the proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res. 2003;31:3784–3788.
Tabb DL, McDonald WH, Yates JR III. DTASelect and Contrast: tools for assembling and comparing protein identifications from shotgun proteomics. J Proteome Res. 2002;1:21–26.
Breiman L, Friedman J, Olshen R, et al. Classification and Regression Trees. Belmont, CA: Wadsworth and Brooks; 1984.
Shen Y, Jacobs JM, Camp DG, et al. Ultra-high-efficiency strong cation exchange LC/RPLC/MS/MS for high dynamic range characterization of the human plasma proteome. Anal Chem. 2004;76:1134–1144.
Imbert-Bismut F, Ratziu V, Pieroni L, Charlotte F, Benhamou Y, Poynard T. Biochemical markers of liver fibrosis in patients with hepatitis C virus infection: a prospective study. Lancet. 2001;357:1069–1075.
Bedossa P, Poynard T. An algorithm for the grading of activity in chronic hepatitis C. The METAVIR Cooperative Study Group. The METAVIR Cooperative Study Group. Hepatology. 1996;24:289–293.
The French METAVIR Cooperative Study Group. Intraobserver and interobserver variations in liver biopsy interpretation in patients with chronic hepatitis C. Hepatology. 1994;20:15–20.
Halfon P, Imbert-Bismut F, Messous D, et al. A prospective assessment of the inter-laboratory variability of biochemical markers of fibrosis (FibroTest) and activity (ActiTest) in patients with chronic liver disease. Compar Hepatol. 2002;1:3.
Rossi E, Adams L, Prins A, et al. Validation of the FibroTest biochemical markers score in assessing liver fibrosis in hepatitis C patients. Clin Chem. 2003;49:450–454.
Wai CT, Greenson JK, Fontana RJ, et al. A simple noninvasive index can predict both significant fibrosis and cirrhosis in patients with chronic hepatitis C. Hepatology. 2003;38:518–526.
Le Calvez S, Thabut D, Messous D, et al. The predictive value of Fibrotest vs. APRI for the diagnosis of fibrosis in chronic hepatitis C. Hepatology. 2004;39:862–863.
Gressner OA, Gao C, Siluschek M, Kim P, Gressner AM. Inverse association between serum concentrations of actin-free vitamin D-binding protein and the histopathological extent of fibrogenic liver disease or hepatocellular carcinoma. Eur J Gastroenterol Hepatol. 2009;21:990–995.
Kawai N, Matsumoto H. Vitamin-D binding protein levels in liver cirrhosis, chronic hepatitis, and rheumatoid arthritis. Nihon Hoigaku Zasshi. 1984;38:797–803.
Ben Ari Z, Osman E, Hutton RA, Burroughs AK. Disseminated intravascular coagulation in liver cirrhosis: fact or fiction? Am J Gastroenterol. 1999;94:2977–2982.
Papatheodoridis GV, Papakonstantinou E, Andrioti E, et al. Thrombotic risk factors and extent of liver fibrosis in chronic viral hepatitis. Gut. 2003;52:404–409.
Mouelhi L, Mekki H, Debbeche R, Salem M, Najjar T. Thrombosis in inflammatory bowel disease: mechanisms and risk factors. Tunis Med. 2009;87:307–310.
Chrusciel P, Goch A, Banach M, Mikhailidis DP, Rysz J, Goch JH. Circadian changes in the hemostatic system in healthy men and patients with cardiovascular diseases. Med Sci Monit. 2009;15:RA203–RA208.
Maksan SM, Ulger Z, Gebhard MM, Schmidt J. Impact of antithrombin III on hepatic and intestinal microcirculation in experimental liver cirrhosis and bowel inflammation: an in vivo analysis. World J Gastroenterol. 2005;11:4997–5001.
Calvaruso V, Maimone S, Gatt A, et al. Coagulation and fibrosis in chronic liver disease. Gut. 2008;57:1722–1727.
Szilagyi A, Blasko B, Ronai Z, Fust G, Sasvari-Szekely M, Guttman A. Rapid quantification of human complement component C4A and C4B genes by capillary gel electrophoresis. Electrophoresis. 2006;27:1437–1443.
Zaman A, Rosen HR, Ingram K, Corless CL, Oh E, Smith K. Assessment of FIBROSpect II to detect hepatic fibrosis in chronic hepatitis C patients. Am J Med. 2007;120:280.e9–14.
Gressner OA, Beer N, Jodlowski A, Gressner AM. Impact of quality control accepted inter-laboratory variations on calculated Fibrotest/Actitest scores for the non-invasive biochemical assessment of liver fibrosis. Clin Chim Acta. 2009;409:90–95.
Van den Bergh G, Arckens L. Fluorescent two-dimensional difference gel electrophoresis unveils the potential of gel-based proteomics. Curr Opin Biotechnol. 2004;15:38–43.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yang, L., Rudser, K.D., Higgins, L. et al. Novel Biomarker Candidates to Predict Hepatic Fibrosis in Hepatitis C Identified by Serum Proteomics. Dig Dis Sci 56, 3305–3315 (2011). https://doi.org/10.1007/s10620-011-1745-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-011-1745-4