Abstract
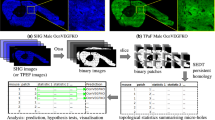
Pathological examination of a biopsy is the most reliable and widely used technique to diagnose bone cancer. However, it suffers from both inter- and intra- observer subjectivity. Techniques for automated tissue modeling and classification can reduce this subjectivity and increases the accuracy of bone cancer diagnosis. This paper presents a graph theoretical method, called extracellular matrix (ECM)-aware cell-graph mining, that combines the ECM formation with the distribution of cells in hematoxylin and eosin stained histopathological images of bone tissues samples. This method can identify different types of cells that coexist in the same tissue as a result of its functional state. Thus, it models the structure-function relationships more precisely and classifies bone tissue samples accurately for cancer diagnosis. The tissue images are segmented, using the eigenvalues of the Hessian matrix, to compute spatial coordinates of cell nuclei as the nodes of corresponding cell-graph. Upon segmentation a color code is assigned to each node based on the composition of its surrounding ECM. An edge is hypothesized (and established) between a pair of nodes if the corresponding cell membranes are in physical contact and if they share the same color. Hence, multiple colored-cell-graphs coexist in a tissue each modeling a different cell-type organization. Both topological and spectral features of ECM-aware cell-graphs are computed to quantify the structural properties of tissue samples and classify their different functional states as healthy, fractured, or cancerous using support vector machines. Classification accuracy comparison to related work shows that the ECM-aware cell-graph approach yields 90.0% whereas Delaunay triangulation and the simple cell-graph approach achieves 75.0 and 81.1% accuracy, respectively.
Similar content being viewed by others
References
Becker WM, Kleinsmith LJ, Hardin J (2000) The world of the cell. Benjamin/Cummings, Menlo Park, CA
Ben-Dor A, Bruhn L, Friedman N, Nachman I, Schummer M, Yakhini Z (2000) Tissue classification with gene expression profiles. J Comput Biol 7(3–4): 559–583
Bilgin C, Demir C, Nagi C, Yener B (2007) Cell-graph mining for breast tissue modeling and classification. In: Engineering in medicine and biology society, 2007. EMBS 2007. 29th Annual international conference of the IEEE, pp 5311–5314
Byun H, Lee SW (2002) Applications of support vector machines for pattern recognition: a survey. Lecture notes in computer science, pp 213–236
Chang CC, Lin CJ (2001) LIBSVM: a library for support vector machines. Software available at http://www.csie.ntu.edu.tw/cjlin/libsvm, 80:604–611
Chen Y, Lin C (2006) Combining SVMs with various feature selection strategies. Stud Fuzziness Soft Comput 207: 315
Chung FRK (1997) Spectral graph theory. American Mathematical Society, Providence
Demir C, Yener B (2005) Automated cancer diagnosis based on histopathological images: a systematic survey. Rensselaer Polytechnic Institute, Technical Report
Doyle S, Hwang M, Shah K, Madabhushi A, Feldman M, Tomaszeweski J (2007) Automated grading of prostate cancer using architectural and textural image features. In: Biomedical imaging: from nano to macro, 2007. ISBI 2007. 4th IEEE international symposium on, pp 1284–1287
Einstein AJ, Wu HS, Sanchez M, Gil J (1998) Fractal characterization of chromatin appearance for diagnosis in breast cytology. J Pathol 185(4): 366–381
Ersoy I, Bunyak F, Mackey MA, Palaniappan K (2008) Cell segmentation using Hessian-based detection and contour evolution with directional derivatIves. In: 15th IEEE international conference on image processing, 2008. ICIP 2008, pp 1804–1807
Esgiar AN, Naguib RNG, Bennett MK, Murray A (1998) Automated feature extraction and identification of colon carcinoma. J Anal Quant Cytol Histol 20(4): 297–301
Esgiar AN, Naguib RNG, Sharif BS, Bennett MK, Murray A (2002) Fractal analysis in the detection of colonic cancer images. Inf Technol Biomed IEEE Trans 6(1): 54–58
Frangi AF, Niessen WJ, Vincken KL, Viergever MA (1998) Multiscale vessel enhancement filtering. Lecture notes in computer science, pp 130–137
Furey TS, Cristianini N, Duffy N, Bednarski DW, Schummer M, Haussler D (2000) Support vector machine classification and validation of cancer tissue samples using microarray expression data. Bioinformatics 16(10): 906–914
Ganster H, Pinz P, Rohrer R, Wildling E, Binder M, Kittler H (2001) Automated melanoma recognition. Med Imaging IEEE Trans 20(3): 233–239
Glotsos D, Spyridonos P, Petalas P, Nikiforidis G, Cavouras D, Ravazoula P, Dadioti P, Lekka I (2003) Support vector machines for classification of histopathological images of brain tumour astrocytomas. In: Proceedings of the international conference on computational methods in sciences and engineering, pp 192–195. World Scientific Publishing Co., Inc. River Edge, NJ, USA
Golub TR, Slonim DK, Tamayo P, Huard C, Gaasenbeek M, Mesirov JP, Coller H, Loh ML, Downing JR, Caligiuri MA et al (1999) Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286(5439): 531
Gunduz C, Yener B, Gultekin SH (2004) The cell graphs of cancer. Bioinformatics 20(1): i145–51
Guyon I, Weston J, Barnhill S, Vapnik V (2002) Gene selection for cancer classification using support vector machines. Mach Learn 46(1): 389–422
Hamilton PW, Bartels PH, Thompson D, Anderson NH, Montironi R, Sloan JM (1997) Automated location of dysplastic fields in colorectal histology using image texture analysis. J Pathol 182(1): 68–75
Hamilton PWD, Allen DC, Watt PCH, Patterson CC, Biggart JD (1987) Classification of normal colorectal mucosa and adenocarcinoma by morphometry. Histopathology 11(9): 901–911
Hladuvka J, Konig A, Groller E (2001) Exploiting eigenvalues of the Hessian matrix for volume decimation. 9th International conference in Central Europe on computer graphics, visualization, and computer vision WSCG, pp 124–129
Hodneland E, Tai XC, Gerdes HH (2009) Four-color theorem and level set methods for watershed segmentation. Int J Comput Vis 82: 264–283
Hsu CW, Chang CC, Lin CJ et al (2003) A practical guide to support vector classification
Ibanez L, Schroeder W, Ng L, Cates J (2005) The ITK software guide, 2nd edn. Kitware, Inc. ISBN 1-930934-15-7, http://www.itk.org/ItkSoftwareGuide.pdf
Jain R, Abraham A (2004) A comparative study of fuzzy classifiers on breast cancer data. Australas Phys Eng Sci Med 27: 147–152
Keenan SJ, Diamond J, McCluggage WG, Bharucha H, Thompson D, Bartels PH, Hamilton PW (2000) An automated machine vision system for the histological grading of cervical intraepithelial neoplasia (CIN). J Pathol 192(3): 351–362
Keerthi SS, Lin CJ (2003) Asymptotic Behaviors of support vector machines with Gaussian Kernel. Neural Comput 15(7): 1667–1689
Lin HT, Lin CJ, Weng RC (2007) A note on Platt’s probabilistic outputs for support vector machines. Mach Learn 68(3): 267–276
Malmstrom PU (1997) Image analysis based grading of bladder carcinoma. Comparison of object, texture and graph based methods and their reproducibility. Anal Cell Pathol 15(1): 1–18
Mangasarian OL, Street WN, Wolberg WH (1995) Breast cancer diagnosis and prognosis via linear programming. Oper Res Baltimore 43: 570–570
Peña-Reyes CA, Sipper M (1999) A fuzzy-genetic approach to breast cancer diagnosis. Artif Intell Med 17(2): 131–155
Platt J (2000) Probabilistic outputs for support vector machines and comparison to regularized likelihood methods. Advances in large margin classifiers, pp 61–74
Schnorrenberg F, Pattichis CS, Schizas CN, Kyriacou K, Vassiliou M (1996) Computer-aided classification of breast cancer nuclei. Technol Health Care 4(2): 147–161
Siek J, Lee LQ, Lumsdaine A (2002) The boost graph library: user guide and reference manual. Addison-Wesley, Reading
Tasoulis DK, Spyridonos P, Pavlidis NG, Cavouras D, Ravazoula P Nikiforidis G, Vrahatis MN (2003) Urinary bladder tumor grade diagnosis using on-line trained neural networks. Lecture notes in computer science, pp 199–206
Todman AG, Naguib RNG, Bennett MK (2001) Orientational coherence metrics: classification of colonic cancerimages based on human form perception. In: Electrical and computer engineering, 2001. Canadian Conference on, vol 2
TOC View (1998) Microscopic image analysis for quantitative measurement and featureidentification of normal and cancerous colonic mucosa. Inf Technol Biomed IEEE Trans 2(3): 197–203
Weyn B, van de Wouwer G, Kumar-Singh S, van Daele A, Scheunders P, van Marck E, Jacob W (1999) Computer-assisted differential diagnosis of malignant mesothelioma based on syntactic Structure analysis. Cytometry 35: 23–29
Wolberg WH, Street WN, Heisey DM, Mangasarian OL (1995) Computer-derived nuclear features distinguish malignant from benign breast cytology. Hum Pathol 26(7): 792–796
Wu B, Abbott T, Fishman D, McMurray W, Mor G, Stone K, Ward D, Williams K, Zhao H (2003) Comparison of statistical methods for classification of ovarian cancer using mass spectrometry data. Bioinformatics 19(13): 1636–1643
Zhou ZH, Jiang Y, Yang YB, Chen SF (2002) Lung cancer cell identification based on artificial neural network ensembles. Artif Intell Med 24(1): 25–36
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: R. Bharat Rao and Romer Rosales.
Rights and permissions
About this article
Cite this article
Bilgin, C.C., Bullough, P., Plopper, G.E. et al. ECM-aware cell-graph mining for bone tissue modeling and classification. Data Min Knowl Disc 20, 416–438 (2010). https://doi.org/10.1007/s10618-009-0153-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10618-009-0153-2