Abstract
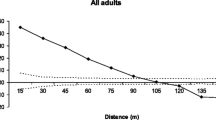
Fragmentation is predicted to increase inbreeding depression and lower the evolutionary potential of organisms by disrupting dispersal. Trees may be more resilient to fragmentation effects due to potential long-distance dispersal mechanisms that genetically connect fragments. Polylepis woodlands in the high Andes are highly fragmented and are currently the focus of reforestation and conservation efforts. Polylepis multijuga Plige (Rosaceae) is a threatened, endemic tree species in the northern Andes of Peru. Samples were collected from 371 adult trees in nine forest fragments separated by 0.5–80 km and genotyped at amplified fragment length polymorphism loci (AFLP) and chloroplast intergenic regions to determine the connectedness of fragments and their suitability for collecting seed for restoration efforts. P. multijuga is wind-pollinated and dispersed; however, genetic diversity in P. multijuga was about half that reported for other wind-pollinated species. Genetic spatial autocorrelation and patterns of chloroplast and AFLP diversity suggest seed dispersal is very limited and that wind dispersed pollen does not effectively connect all fragments. Conservation of this species will require reforestation efforts and possibly augmentation of some fragments to increase their genetic diversity. Collecting seed from multiple large fragments and from individuals separated by at least 25 m within fragments would maximize the genetic diversity of seed collections for reforestation or augmentation. Future studies of this and other Polylepis species should determine how complex topography may affect wind mediated dispersal between fragments and patterns of genetic diversity.






Similar content being viewed by others
References
Aguilar R, Quesada M, Ashworth L, Herrerias DY, Lobo J (2008) Genetic consequences of habitat fragmentation in plant populations: susceptible signals in plant traits and methodological approaches. Mol Ecol 17:5177–5188
AMPA, Amazonicos por La Amazonia NGO (2008) Plan de manejo de la concesión para conservación Alto Huayabamba. Technical Report AMPA Peru NGO, San Martin, Peru
Aragundi S, Hamrick JL, Parker KC (2011) Genetic insights into the historical distribution of Polylepis pauta (Rosaceae) in the northeastern Cordillera Oriental of Ecuador. Conserv Genet 12:607–618
Bischoff A, Steinger T, Müller-Schärer H (2010) The importance of plant provenance and genotypic diversity of seed material used for ecological restoration. Restor Ecol 18:338–348
Broadhurst LM, Lowe A, Coates DJ, Cunningham SA, McDonald M, Vesk PA, Yates C (2008) Seed supply for broadscale restoration: maximizing evolutionary potential. Evol Appl 1:587–597
Chung S-M, Staub JE (2003) The development and evaluation of consensus chloroplast primer pairs that possess highly variable sequence regions in a diverse array of plant taxa. Theoret Appl Genet 107:757–767
Dubreuil M, Riba M, González-Martínez SC, Vendramin GG, Sebastiani F, Mayol M (2010) Genetic effects of chronic habitat fragmentation revisited: strong genetic structure in a temperate tree, Taxus baccata (Taxaceae), with great dispersal capability. Am J Bot 97:303–310
Ficetola GF, Garner TWJ, De Bernardi F (2007) Genetic diversity, but not hatching success, is jointly affected by post glacial colonization and isolation in the threatened frog, Rana latastei. Mol Ecol 16:1787–1797
Fjeldså J (1993) The avifauna of the Polylepis woodlands of the Andean highlands: the efficiency of basing conservation priorities on patterns of endemism. Bird Conserv Internat 3:37–55
Fjeldså J (2002) Polylepis forests—vestiges of a vanishing ecosystem in the Andes. Ecotropica 8:11–123
Fjeldså J, Kessler M (1996) Conserving the biological diversity of Polylepis woodlands of the highland of Peru and Bolivia: a contribution to sustainable natural resource management in the Andes. NORDECO, Copenhagen
Frankham R, Ballou JD, Briscoe DA (2010) Introduction to conservation genetics. Cambridge University Press, New York
Gosling WD, Hanselman JA, Knox C, Valencia B, Bush MB (2009) Long-term drivers of change in Polylepis woodland distribution in the central Andes. J Veg Sci 20:1041–1052
Guillot G, Estoup A, Mortier F, Cosson JF (2005a) A spatial statistical model for landscape genetics. Genetics 170:1261–1280
Guillot G, Mortier F, Estoup A (2005b) Geneland: a computer package for landscape genetics. Mol Ecol Notes 5:708–711
Hamrick JL (2004) Response of forest trees to global environmental changes. Forest Ecol Manag 197:323–335
Hamrick JL, Godt MJW (1990) Allozyme diversity in plant species. In: Brown AHD, Clegg MT, Kahler AL, Weir BS (eds) Plant population genetics, breeding and genetic resources. Sinauer, Sunderland, pp 43–63
Hansen BCS, Wright HE, Bradbury JP (1984) Pollen studies in the Junín area, central Peruvian Andes. Geo Soc Am Bull 95:1454–1465
Hansen BCS, Seltzer GO, Wright HE (1994) Late quaternary vegetational change in the central Peruvian Andes. Palaeogeogr Palaeoclimatol Palaeoecol 109:263–285
Hensen I, Cierjacks A, Hirsch H, Kessler M, Romoleroux K, Renison D, Wesche K (2011) Historic and recent fragmentation coupled with altitude affect the genetic population structure of one of the world’s highest tropical tree line species. Global Ecol Biogeog (online early). doi:10.1111/j.1466-8238.2011.00691.x
Honnay O, Jacquemyn H (2007) Susceptibility of common and rare plant species to the genetic consequences of habitat fragmentation. Conserv Biol 21:823–831
Hu LJ, Uchiyama K, Shen HL, Ide Y (2010) Multiple-scaled spatial genetic structures of Fraxinus mandshurica over a riparian-mountain landscape in Northeast China. Conserv Genet 11:77–87
Huff DR, Peakall R, Smouse PE (1993) RAPD variation within and among natural populations of outcrossing buffalograss Buchloe dactyloides (Nutt.) Engelm. Theor Appl Genet 86:927–934
IUCN, The World Conservation Union (2008) The IUCN Red list of threatened species. Available at http://www.iucnredlist.org/apps/redlist/search
Jump AS, Pañuelas J (2006) Genetic effects of chronic habitat fragmentation in a wind-pollinated tree. Proc Natl Acad Sci USA 103:8096–8100
Karron JD (1987) A comparison of levels of genetic polymorphism and self-compatibility in geographically restricted and widespread plant congeners. Evol Ecol 1:47–58
Kauffman F (2001) Momias ocultas en las selvas de los Chachapoya(s). Revista Copé 8:1–8
Kettle CJ, Hollingsworth PM, Jaffré T, Moran B, Ennos RA (2007) Identifying the early genetic consequences of habitat degradation in a highly threatened tropical conifer, Araucaria nemorosa Laubenfels. Mol Ecol 16:3581–3591
Kramer AT, Ison JL, Ashley MV, Howe HF (2008) The paradox of forest fragmentation genetics. Conserv Biol 22:878–885
Kremer A, Caron H, Cavers S, Colpaert N, Gheysen G, Gribel R, Lemes M, Lowe AJ, Margis R, Navarro C, Salgueiro F (2005) Monitoring genetic diversity in tropical trees with multilocus dominant markers. Heredity 95:274–280
León B, Pitman N, Roque J (2006) Introducción a las plantas endémicas del Perú. In: León B et al. (eds) El libro rojo de las plantas endémicas del Perú, Revista Peruana de Biología. Número especial 13:9s–22s
Lloyd H, Marsden SJ (2008) Bird community variation across Polylepis woodland fragments and matrix habitats: implications for conservation within a high Andean landscape. Biodivers Conserv 17:2645–2660
Lynch M, Milligan BG (1994) Analysis of population genetic structure with RAPD markers. Mol Ecol 3:91–99
Macbride JF (1938) Flora of Peru, vol 13, Part II. Publications of the Field Museum of Natural History Botanical Series, Chicago, pp 1036–1119
Mendoza W, León B (2006) Rosaceae endémicas del Perú. In: León B et al. (eds), El libro rojo de las plantas endémicas del Perú, Revista Peruana de Biología. Número especial 13:583s–585s
Montoya D, Zavala MA, Rodríguez MA, Purves DW (2008) Animal versus wind dispersal and the robustness of tree species to deforestation. Science 320:1502–1504
Nybom H (2004) Comparison of different nuclear DNA markers for estimating intraspecific genetic diversity in plants. Mol Ecol 13:1143–1155
Ohtani M, Terauchi H, Nishihiro J, Ueno S, Tsumura Y, Washitani I (2008) Towards a legal framework for systematic conservation: identification and development of allele-specific PCR markers for conspecific varieties of an endangered perennial herb Primula kisoana Miquel based on sequence variation of chloroplast DNA. Conserv Genet 9:1173–1181
Parker KC, Hamrick JL, Parker AJ, Nason JD (2001) Fine-scale genetic structure in Pinus clausa (Pinaceae) populations, effects of disturbance history. Heredity 87:99–113
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Peakall R, Ruibal M, Lindenmayer DB (2003) Spatial autocorrelation analysis offers new insights into gene flow in the Australian bush rat, Rattus fuscipes. Evol 57:1182–1195
Purcell J, Brelsford A, Kessler M (2004) The world’s highest forest. Bioscience 92:454–461
Renison D, Cingolani AM, Suarez R (2002) Effects of fire on a Polylepis australis (Rosaceae) woodland in the mountains of Cordoba, Argentina. Rev Chilena Hist Nat 75:719–727
Renison D, Hensen I, Suarez R, Cingolani AM (2006) Cover and growth habit of Polylepis woodlands and shrublands in the mountains of central Argentina: human or environmental influence? J Biogeogr 33:876–887
Ruiz LH, Pavon JA (1794) Florae Peruvianae et Chilensis Prodromus. Imprenta de Sancha, Madrid
Schmidt-Lebuhn AN, Kessler M, Kumar M (2006) Promiscuity in the Andes: species relationships in Polylepis (Rosaceae, Sanguisorbeae) based on AFLP and morphology. Syst Bot 31:547–559
Schmidt-Lebuhn AN, Seltmann P, Kessler M (2007) Consequences of the pollination system on genetic structure and patterns of species distribution in the Andean genus Polylepis (Rosaceae): a comparative study. Plant Syst Evol 266:91–103
Seltmann S, Renison D, Cocucci A, Hensen I, Jung K (2007) Fragment size, pollination efficiency and reproductive success in natural populations of wind-pollinated Polylepis australis (Rosaceae) trees. Flora 202:547–554
Seltmann P, Renison D, Cierjacks A, Hensen I, Cocucci A (2009a) Mating system, outcrossing distance effects and pollen availability in the wind-pollinated treeline species Polylepis australis BITT. (Rosaceae). Basic Appl Ecol 10:52–60
Seltmann P, Hensen I, Renison D, Wesche K, Ploch S, Duenas JR, Cocucci A, Jung K (2009b) Biparental inbreeding depression, genetic relatedness and progeny vigour in a wind-pollinated treeline species in Argentina. Plant Ecol 205:155–164
Shaw J, Lickey EB, Schilling EE, Small RL (2007) Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: the tortoise and the hare III. Am J Bot 94:275–288
Simpson BB (1979) A revision of the genus Polylepis (Rosaceae: Sanguisorbeae). Sm C Bot 43:1–62
Sork VL, Smouse PE (2006) Genetic analysis of landscape connectivity in tree populations. Landscape Ecol 21:821–836
Teich I, Cingolani AM, Renison D, Hensen I, Giorgis MA (2005) Do domestic herbivores retard Polylepis australis Bitt. woodland recovery in the mountains of Cordoba, Argentina? Forest Ecol Manag 219:229–241
Torres RC, Renison D, Hensen I, Suarez R, Enrico L (2008) Polylepis australis’ regeneration niche in relation to seed dispersal, site characteristics and livestock density. Forest Ecol Manag 254:255–260
UNEP-WCMC (2004) United Nations Environment Programme. World Conservation Monitoring Center, available at http://www.unep-wcmc.org
Vanden-Broeck A, Gruwez R, Cox K, Adriaesnssens S, Michalczyk IM, Verheyen K (2011) Genetic structure and seed-mediated dispersal rates of an endangered shrub in a fragmented landscape: a case study for Juniperus communis in northwestern Europe. BMC Genetics 12:73
Vekemans X (2002) AFLP-SURV version 1.0. Distributed by the author. Laboratoire de Génétique et Ecologie Végétale, Université Libre de Bruxelles, Belgium
Vekemans X, Hardy OJ (2004) New insights from fine-scale spatial genetic structure analyses in plant populations. Mol Ecol 13:921–935
Vos P, Hogers R, Bleeker M, Reijans M, Van De Lee T, Hornes M, Freuters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP, a new technique for DNA fingerprinting. Nucleic Acids Res 23:4407–4414
Vuilleumier F, Monasterio M (eds) (1986) High altitude tropical biogeography. Oxford University Press, Oxford
Weising K, Gardner RC (1999) A set of conserved PCR primers for the analysis of simple sequence repeat polymorphisms in chloroplast genomes of dicotyledonous angiosperms. Genome 42:9–19
Whitlock R, Hipperson H, Mannarelli M, Butlin RK, Burke T (2008) An objective, rapid and reproducible method for scoring AFLP peak-height data that minimizes genotyping error. Mol Ecol Resour 8:725–735
Young AG, Clarke GM (eds) (2000) Genetics, demography and viability of fragmented populations. Cambridge University Press, Cambridge
Zhivotovsky LA (1999) Estimating population structure in diploids with multilocus dominant DNA markers. Mol Ecol 8:907–913
Aknowledgments
We thank Tony Burgess, Matt Chumchal, Amanda Hale, and Tammy Morgan for help and valuable suggestions. Special thanks to our collaborators Jhon Panduro Cometivos, Karina Pinasco Vela, Abnet Cusquipoma, and Manuel from Amazonicos por la Amazonia for help with all aspects of this study. We also thank the Muños Ruiz family for letting us sample on their land “Los Gevaras” and especially to Edwin and Jobito Muños for their help. Thanks to the people of Direccion General Forestal y de Fauna Silvestre for the research and exportation permits. Thanks also to Mercedes Flores Pimentel, director of the “MOL” herbarium; Keri McNew, Tiana Franklin, and John Janovec from the Botanical Research Institute of Texas (BRIT). Funding was provided by Texas Christian University’s Department of Biology Adkins Fellowship and the Department of Environmental Science.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Quinteros-Casaverde, N., Flores-Negrón, C.F. & Williams, D.A. Low genetic diversity and fragmentation effects in a wind-pollinated tree, Polylepis multijuga Plige (Rosaceae) in the high Andes. Conserv Genet 13, 593–603 (2012). https://doi.org/10.1007/s10592-011-0310-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-011-0310-1