Abstract
Over 70% of North American freshwater mussel species (families Unionidae and Margaritiferidae) are listed as threatened or endangered. Knowledge of the genetic structure of target species is essential for the development of effective conservation plans. Because Ambelma plicata is a common species, its population genetic structure is likely to be relatively intact, making it a logical model species for investigations of freshwater mussel population genetics. Using mtDNA and allozymes, we determined the genotypes of 170+ individuals in each of three distinct drainages: Lake Erie, Ohio River, and the Lower Mississippi River. Overall, within-population variation increased significantly from north to south, with unique haplotypes and allele frequencies in the Kiamichi River (Lower Mississippi River drainage). Genetic diversity was relatively low in the Strawberry River (Lower Mississippi River drainage), and in the Lake Erie drainage. We calculated significant among-population structure using both molecular markers (A.p. Φst = 0.15, θst = 0.12). Using a hierarchical approach, we found low genetic structure among rivers and drainages separated by large geographic distances, indicating high effective population size and/or highly vagile fish hosts for this species. Genetic structure in the Lake Erie drainage was similar to that in the Ohio River, and indicates that northern populations were founded from at least two glacial refugia following the Pleistocene. Conservation of genetic diversity in Amblema plicata and other mussel species with similar genetic structure should focus on protection of a number of individual populations, especially those in southern rivers.
Similar content being viewed by others
References
Avise JC (2000) Phylogeography: A history of formation of species. Harvard University Press, Cambridge, Massachusetts
Berendzen PB, Simons AM, Wood RM (2003) Phylogeography of the northern hogsucker, Hypentelium nigricans (Teleostei: Cypriniformes): Genetic evidence for the existence of the ancient Teays River. J Biogeogr 30:1139–1152
Berg DJ, Berg PH (2000) Conservation genetics of freshwater mussels: Comments on Mulvey et al. Conserv Biol 14:1920–1923
Berg DJ, Cantonwine EG, Hoeh WR, Guttman SI (1998) Genetic structure of Quadrula quadrula, Bivalvia : Unionidae.: Little variation across large distances. J Shell Res 17:1365–1373
Berg DJ, Haag WR, Guttman SI, Sickel JB (1995) Mantle biopsy: A technique for nondestructive tissue sampling of freshwater mussels. J N Amer Benthol Soc 14:577–581
Bernatchez L, Wilson CC (1998) Comparative phylogeography of nearctic and palearctic fishes. Mol Ecol 7:431–452
Bierne N, Launey S, Naciri-Graven Y, Bonhomme F (1998) Early effect of inbreeding as revealed by microsatellite analyses on Ostrea edulis larvae. Genetics 148:1893–1906
Bilton DT, Freeland JR, Okamura B (2001) Dispersal in freshwater invertebrates. Annu Rev Ecol Syst 32:159–181
Bogan AE (1993) Fresh-water bivalve extinctions, Mollusca, Unionoida.; A search for causes. Amer Zool 33:599–609
Bunn SE, Hughes JM (1997) Dispersal and recruitment in streams: Evidence from genetic studies. J N Amer Benth Soc 16:338–346
Clement M, Posada D, Crandall K (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1660
Crandall KA (1996) Multiple interspecies transmissions of human and simian T-cell leukemia/lymphoma virus type I sequences. Mol Biol Evol 13:115–131
Crandall KA (1998) Conservation phylogenetics of Ozark crayfishes: Assigning priorities for aquatic habitat protection. Biol Conserv 84:107–117
Cummings KS, Mayer CA (1992) Field guide to freshwater mussels of the Midwest. Illinois Natural History Survey, Champaign, Illinois
Curole JP, Foltz DW, Brown KA (2004) Extensive allozyme monomorphism in a threatened species of freshwater mussel, Margaritifera hembeli Conrad, Bivalvia : Margaritiferidae. Conserv Genet 5:271–278
Elderkin CL, Klerks PL, Theriot E (2001) Shifts in allele and genotype frequencies in zebra mussels, Dreissena polymorpha, along the latitudinal gradient formed by the Mississippi River. J N Am Bethol Soc 20:595–605
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: Application to human mitochondrial DNA restriction data. Genetics 131:479–491
Fetzner JW, Crandall KA (2003) Linear habitats and the nested clade analysis: An empirical evaluation of geographic versus river distances using an Ozark crayfish (Decapoda: Cambaridae). Evolution 57:2101–2118
Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R (1994) DNA primers for amplification of mitochondrial cytochrome oxidase subunit I from diverse metazoan invertebrates. Mol Mar Biol Biotech 3:294–299
Foltz DW (1986) Null alleles as a possible cause of heterozygote deficiencies in the oyster Crassostrea virginica and other bivalves. Evolution 40:869–870
Frankham R (2003) Genetics and conservation biology. Comptes Rendus Biologies 326:S22–S29
Fraser DJ, Bernatchez L (2001) Adaptive evolutionary conservation: Towards a unified concept for defining conservation units. Mol Ecol 10:2741–2752
Gaffney PM, Scott TM, Koehn RK, Diehl WJ (1990) Interrelationships of heterozygosity, growth rate and heterozygote deficiencies in the coot clam, Mulinia lateralis. Genetics 124:687–699
Graf DL (2002) Historical biogeography and late glacial origin of the freshwater pearly mussel (Bivalvia: Unionidae) faunas of Lake Erie, North America. Occasional Papers on Mollusks 6:175–211
Grobler JP, Jones JW, Johnson NA, Beaty B, Struthers J, Neves RJ, Hallerman EM (2006) Patterns of genetic differentiation in the slabside pearlymussel, Lexingtonia dollabelloides Lea 1840, in the Tennessee drainage. J Molluscan Stud 72:65–75
Haag WR, Berg DJ, Garton DW, Farris JL (1993) Reduced survival and fitness in native bivalves in response to fouling by the introduced zebra mussel, Dreissena polymorpha. in western Lake Erie. Can J Fish Aquat Sci 50:13–19
Hartl DL, Clark AG (1997) Principles of population genetics, 3rd edn. Sinauer Associates, Inc., Sunderland, Massachusetts
Hebert PDN, Beaton MJ (1993) Methodologies for allozyme analysis using cellulose acetate electrophoresis. Helena Laboratories, Beaumont, Texas
Hewitt GM (1996) Some genetic consequences of ice ages, and their role in divergence and speciation. Biol J Linn Soc 58:247–276
Hilbish TJ, Koehn RK (1985) Dominance in physiological phenotypes and fitness at an enzyme locus. Science 229:52–54
Hoeh WR, Stewart DT, Guttman SI (2002) High fidelity of mitochondrial genome transmission under the doubly uniparental mode of inheritance in freshwater mussels (Bivalvia: Unionoidea). Evolution 56:2252–2261
Hoeh WR, Stewart DT, Sutherland BW, Zouros E (1996) Cytochrome oxidase sequence comparisons suggest an unusually high rate of mitochondrial DNA evolution in Mytilus (Mollusca: Bivalvia). Mol Biol Evol 13:418–421
Hoggarth MA (1995–1996) The Unionidae (Mollusca: Bivalvia) of the Walhonding River, Coshocton county, Ohio, including the federally endangered catspaw, Epioblasma obliquata obliquata, fanshell, Eyprogenia stegaria, and clubshell, Pleurobema clava mussels. Walkerana 8:149–176
Hughes JM, Bunn SE, Cleary C, Hurwood DA (2000) A hierarchical analysis of the genetic structure of an aquatic insect Bungona (Baetidae: Ephemeroptera). Heredity 85:561–570
Hutchison DW, Templeton AR (1999) Correlation of pairwise genetic and geographic distance measures: Inferring the relative influences of gene flow and drift on the distribution of genetic variability. Evolution 53:1898–1914
Johnson RL, Liang FQ, Farris JL (1998) Genetic diversity among four Amblemini species (Bivalvia: Unionidae) in the Cache and White rivers, Arkansas. Southwest Nat 43:321–332
Kimura M, Weiss GH (1964) The stepping stone model of population structure and the decrease of genetic correlation with distance. Genetics 49:561–576
King TL, Eackles MS, Gjetvaj B, Hoeh WR (1999) Intraspecific phylogeography of Lasmigona subviridis (Bivalvia: Unionidae): Conservation implications of range discontinuity. Mol Ecol 8:S65–S78
Lessios HA (1992) Testing electrophoretic data for agreement with Hardy-Weinberg expectations. Mar Biol 112:517–523
Lewis PO, Zaykin D (1999) Genetic Data Analysis: Computer program for the analysis of allelic data. Version 1.0, d13. http://www.lewis.eeb.uconn.edu/lewishome/software.html
Lyons J, Kanehl P (2002) Seasonal movements of smallmouth bass in streams. In: Philipp DP, Ridgeway MS (eds) Black bass: Ecology, conservation, and management, symposium 31. American Fisheries Society, Bethesda, Maryland, pp␣149–160
Machordom A, Araujo R, Erpenbeck D, Ramos MA (2003) Phylogeography and conservation genetics of endangered European Margaritiferidae, Bivalvia : Unionoidea. Biol J Linn Soc 78:235–252
Maddison DR, Maddison WP (2001) MacClade 4: Analysis of phylogeny and character evolution. Sunderland Associates, Sunderland, Massachusetts
Mandrak NE, Crossman EJ (1992) Postglacial dispersal of freshwater fishes into Ontario. Can J Zool 70:2247–2259
Mayden RL (1985) Biogeography of Ouachita highland fishes. Southwest Nat 30:195–211
Mayden RL (1988) Vicariance biogeography, parsimony, and evolution in North American freshwater fishes. Syst Zool 37:329–355
McMahon RF (1991) Mollusca: Bivalvia. In: Thorpe JH, Covich AP (eds) Ecology and classification of North American freshwater invertebrates. Academic Press, San Diego, pp␣315–399
Meffe GK, Vrijenhoek RC (1988) Conservation genetics in the management of desert fishes. Conserv Biol 2:157–169
Metcalfe-Smith JL, Staton SK, Mackie GL, Lane NM (1998) Changes in the biodiversity of freshwater mussels in the Canadian waters of the lower Great Lakes drainage basin over the past 140 years. J Great Lakes Res 24:845–858
Miller MP (1997) Tools for population genetic analyses, TFPGA. a Windows program for the analysis of allozyme and molecular population genetic data. Available from the author: M.P. Miller. http://www.bioweb.usu.edu/mpmbio/
Miller MP, Blinn DW, Keim P (2002) Correlations between observed dispersal capabilities and patterns of genetic differentiation in populations of four aquatic insect species from the Arizona White Mountains, U.S.A. Freshw Biol 47:1660–1673
Moritz C (1994a) Applications of mitochondrial DNA analysis in conservation: A critical review. Mol Ecol 3:401–411
Moritz C (1994b) Defining evolutionarily significant units for conservation. Trends Ecol Evol 9:373–375
Moritz C (2002) Strategies to protect biological diversity and the␣evolutionary processes that sustain it. Syst Biol 51:238–254
Mulvey M, Lydeard C, Pyer DL, Hicks KM, Brim-Box J, Williams JD, Butler RS (1997) Conservation genetics of North American freshwater mussels Amblema and Megalonaias. Conserv Biol 11:868–878
Nagel KO, Badino G, Alessandria B (1996) Population genetics of European Anodontinae (Bivalvia: Unionidae). J Mollus Stud 62:343–357
Nei M (1978) Estimating average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
Neves RJ (1999) Conservation and commerce: Management of freshwater mussel, Bivalvia : Unionoidea, resources in the United States. Malacologia 41:461–474
Newman D, Pilson D (1997) Increased probability of extinction due to decreased genetic effective population size: Experimental populations of Clarkia pulchella. Evolution 51:354–362
Reed DH, Frankham R (2003) Correlation between fitness and genetic diversity. Conserv Biol 17:230–237
Richardson BJ, Baverstock PR, Adams M (1986) Allozyme electrophoresis: A handbook for animal and population studies. Academic Press, New York
Roe KJ, Hartfield PD, Lydeard C (2001) Phylogeographic analysis of the threatened and endangered superconglutinate producing mussels of the genus Lampsilis (Bivalvia: Unionidae). Mol Ecol 10:2225–2234
Roe KJ, Lydeard C (1998) Molecular systematics of the freshwater mussel genus Potamilus (Bivalvia: Unionidae). Malacologia 39:195–205
Rousset F, Raymond M (1995) Testing heterozygote excess and deficiency. Genetics 140:1413–1419
Saccheri I, Kuussaari M, Kankare M, Vikman P, Fortelius W, Hanski I (1998) Inbreeding and extinction in a butterfly metapopulation. Nature 392:491–494
Schmidt PS, Rand DM (1999) Intertidal microhabitat and selection at Mpi: Interlocus contrasts in the northern acorn barnacle, Semibalanus balanoides. Evolution 53:135–146
Schneider SH, Kueffer J, Roessli D, Excoffier L (2000) Arlequin: A software for population genetic data analysis, Version 2.0. Geneva, Switzerland
Schultheis AS, Weigt LA, Hendricks AC (2002) Gene flow, dispersal, and nested clade analysis among populations of the stonefly Peltoperla tarteri in the southern Appalachians. Mol Ecol 11:317–327
Serb JM, Buhay JE, Lydeard C (2003) Molecular systematics of the North American freshwater bivalve genus Quadrula (Unionidae: Ambleminae) based on mitochondrial ND1 sequences. Mol Phylogen Evol 28:1–11
Slatkin M (1981) Estimating levels of gene flow in natural populations. Genetics 99:323–335
Sokal RR, Sneath PHA (1963) Principles of Numerical Taxonomy. W. H. Freeman, San Francisco, CA
Stiven AE, Alderman J (1992) Genetic similarities among certain freshwater mussel populations of the Lampsilis genus in North Carolina. Malacologia 34:355–369
Swofford DL (2000) PAUP*; phylogenetic analysis using parsimony (* and other methods). Sinauer Associates, Sunderland, MA
Swofford DL, Selander RB (1981) BIOSYS-1: a FORTRAN program for the comparative analysis of electrophoretic data in population genetics and systematics. J Hered 72:281–283
Tamura K, Nei M (1993) Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol 10:512–526
Templeton AR (1998) Nested clade analyses of phylogeographic data: Testing hypotheses about gene flow and population history. Mol Ecol 7:381–397
Templeton AR, Sing CF (1993) A cladistic analysis of phenotypic associations with haplotypes inferred from restriction endonuclease mapping 4. Nested analyses with cladogram uncertainty and recombination. Genetics 134:659–669
Turner TF, Trexler JC, Harris JL, Haynes JL (2000) Nested cladistic analysis indicates population fragmentation shapes genetic diversity in a freshwater mussel. Genetics 154:777–785
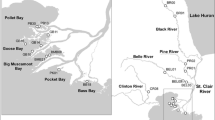
Vaughn CC, Mather CM, Pyron M, Mehlhop P, Miller EK (1996) The current and historical mussel fauna of the Kiamichi River, Oklahoma. Southwest Nat 41:325–328
Vaughn CC, Spooner DE (2004) Status of the mussel fauna of the Poteau River and implications for commercial harvest. Amer Mid Nat 152:336–346
Waples RS (1998) Separating the wheat from the chaff: Patterns of genetic differentiation in high gene flow species. J Hered 89:438–450
Wares JP, Turner TF (2003) Phylogeography and diversification in aquatic mollusks. In: Lydeard C, Lindberg DR (eds) Molecular systematics and phylogeography of mollusks. Smithsonian Books, Washington, D.C, pp 229–269
Watt WB (1977) Adaptation at specific loci. 1. Natural selection on phosphoglucose isomerase of Colias butterflies: Biochemical and population aspects. Genetics 87:177–194
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Whitlock MC, McCauley DE (1999) Indirect measures of gene flow and migration: Fst≠1/(4 Nm + 1). Heredity 82:117–125
Williams JD, Warren ML, Cummings KS, Harris JL, Neves RJ (1993) Conservation status of freshwater mussels of the United States and Canada. Fisheries 18:6–21
Wright S (1942) Isolation by distance. Genetics 28:114–138
Wright S (1965) The interpretation of population structure by F-statistics with special regard to systems of mating. Evolution 19:395–420
Zouros E, Foltz D (1984) Possible explanations of heterozygote deficiency in bivalve mollusks. Malacologia 25:583–591
Acknowledgements
We thank S. Guttman and W. Hoeh for assistance and advice with methodologies. We thank the following field assistants: J. Di Maio, S. Staton, S. Roark, K. Elderkin , J. Harris, S. Chordas , C. Meier, and D. Spooner. B. Jack, J. Trybula, S. Roark, V. Gervasio, and numerous undergraduate students assisted with the laboratory work. E. Cunningham calculated geographic distances among populations. This manuscript was greatly improved by the comments of R. Waples and two anonymous reviewers. Mussels were collected under multiple permits from Division of Wildlife, Ohio Department of Natural Resources, and Oklahoma Department of Wildlife Conservation. Tissue collection by C. Vaughn was supported by NSF grants DEB-9306687 and DEB-9870092. This project was supported by Ohio’s Lake Erie Protection Fund (grant 00–01).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Elderkin, C.L., Christian, A.D., Vaughn, C.C. et al. Population genetics of the freshwater mussel, Amblema plicata (Say 1817) (Bivalvia: Unionidae): Evidence of high dispersal and post-glacial colonization. Conserv Genet 8, 355–372 (2007). https://doi.org/10.1007/s10592-006-9175-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-006-9175-0