Abstract
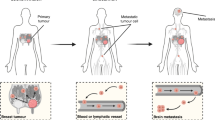
Invasive breast cancer tends to metastasize to lymph nodes and systemic sites. The management of metastasis has evolved by focusing on controlling the growth of the disease in the breast/chest wall, and at metastatic sites, initially by surgery alone, then by a combination of surgery with radiation, and later by adding systemic treatments in the form of chemotherapy, hormone manipulation, targeted therapy, immunotherapy and other treatments aimed at inhibiting the proliferation of cancer cells. It would be valuable for us to know how breast cancer metastasizes; such knowledge would likely encourage the development of therapies that focus on mechanisms of metastasis and might even allow us to avoid toxic therapies that are currently used for this disease. For example, if we had a drug that targeted a gene that is critical for metastasis, we might even be able to cure a vast majority of patients with breast cancer. By bringing together scientists with expertise in molecular aspects of breast cancer metastasis, and those with expertise in the mechanical aspects of metastasis, this paper probes interesting aspects of the metastasis cascade, further enlightening us in our efforts to improve the outcome from breast cancer treatments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
S. David Nathanson and Michael Detmar
The scientifically based management of breast cancer (BC) is dependent upon an understanding of the natural history of the disease. Initially only direct treatment of the diseased breast was possible, and surgery played a predominant role well into the middle of the twentieth century [1]. Treatment by radical mastectomy was offered as an ‘all or nothing’ response; the practitioner could do a drastic operation with the hope that all the cancer had been removed. Patients and their families to this day ask the surgeon whether he/she ‘got it all.’ Until fairly recently that also meant removing all the lymph nodes in the axilla, even when the tumor had not spread to those nodes, exposing the patient to an uncomfortable series of postoperative complications, including severe lymphedema of the arm.
The procedures and processes that played out in the imagination of physicians treating patients with BC were based upon a mechanical/anatomic understanding of metastasis. In a sense, everything in the metastatic process could be related to ‘tubes,’ namely, blood vessels and lymphatics, and their connections in the breast, axilla, and the systemic circulation; tumor cells traveled through these vessels to other parts of the body where they might invade the end organ, such as the lung, liver, bone or brain, forming metastases. Once vital organs had succumbed to these invasive tumors the patient would eventually die of the disease.
When radiation [2] was added to the armamentarium of treatment for BC, it was initially only used for locoregional management on the superficial chest, axilla and supraclavicular areas, but it later became useful as well for direct treatment of metastases in distant organs. The mechanisms of metastasis were still not explored beyond the understanding of an anatomic component; the ‘maps’ of metastasis were known to involve lymphatics, blood vessels, and end points, much like geographic maps that indicate roads, rivers, cities, mountains, coast lines and states.
Systemic treatment for BC, in the form of chemotherapy and endocrine manipulation, advanced the management of BC and was based upon a developing understanding of tumor cell biology, particularly the mechanisms by which tumor cells proliferated [3]. Proliferation did not explain why tumor cells metastasized, although cells that landed in other organs proliferated just like they had in the primary site in the breast. These systemic treatments were often directed at symptoms, such as bone pain.
Evolution and revolution in the management of BC has focused on multidisciplinary treatment and most of the advances have been directed at decreasing metastasis by decreasing proliferation; cells that do not proliferate seem not to metastasize. Many of the large clinical studies of the past few decades [4] have focused attention on preventing metastasis by treating the patient with adjuvant therapies, potentially killing microscopic metastases because that also improves survival. These advances have been accomplished knowing where but not how BC metastasizes.
We have begun a new phase in the systemic treatment of BC by focusing on molecular targets that can be attacked, such as the well-known HER-2/neu, which is based upon molecular aberrations in the tumor cells, but not proven to be involved in how tumors metastasize. Some molecular targets could be functionally important in how tumor cells metastasize without necessarily affecting their ability to proliferate. Drugs targeting functionally important molecules could potentially stop tumors from metastasizing and result in prolonged survival of the patients.
Studies in animal models and in vitro at the dawn of the experimental metastasis era [5] showed the importance of tumor cell invasion into surrounding tissues, with adjusted biological processes already known in cell biology, such as chemotaxis, proteolytic enzyme secretion, expression and de-expression of adhesion molecules, the development of new blood and lymphatic vessels in and around tumors, and immune reactivity [6]. Early pharmacologic studies, using drugs that target biochemical stages in metastasis, show some promise but the field is in its infancy.
The guidelines for BC management, based upon high quality clinical studies have become very dependent upon molecular markers and on statistical models at predicting metastasis [7]. For example, the Oncotype DX test looks at 16 genes, some of which may be functionally important in metastasis, to predict which estrogen-receptor positive tumors that have not metastasized to regional lymph nodes (RLNs) are likely to metastasize to systemic sites [7]. Clinicians frequently use this and other gene studies to decide which patients might benefit from chemotherapy.
The Henry Ford Cancer Institute Mini Symposium on the mechanisms of BC metastasis was held in October 2019 in San Francisco as part of the 8th International Cancer Metastasis Congress. The main objectives of this mini symposium were to look at some recent studies on how tumor cells behave and travel to systemic sites, either through the RLN or directly into the systemic circulation at the site of the original tumor. The more we know about how BC metastasizes at a microscopic, cellular and molecular level, the more likely we are to develop new, more effective ways of treating BC.
The potential role of the sentinel node in systemic metastasis
Timothy P. Padera
The presence of lymph node metastasis (LNM) is associated with worse clinical outcomes for cancer patients than those without LNM [8,9,10]. The question becomes: “Why is this true?” Is it because LNM is just a biomarker for the aggressiveness of the primary cancer, in which more aggressive tumors spread to distant sites and drive patient mortality [1, 11]? Or is it that LNMs themselves can drive cancer progression by serving as a source for distant metastases [12,13,14,15]?
There is new urgency to address these questions as 4 randomized clinical trials (three breast cancer studies-ASCOG-Z0011, IBCSG 23-01, and AMAROS; one melanoma study MSLT-II) have shown that additional lymph node resection in patients beyond the sentinel lymph node does not provide any survival benefit when adjuvant radiation therapy and systemic therapies are used [16,17,18,19,20]. However, implicit in this strategy is the potential to leave cancer bearing lymph nodes in the patient that will require further therapy. Radiation therapy of RLNs has been shown to improve outcomes (disease-free and cancer-specific survival) in early-stage BC [21, 22]. Thus, treatment of lymph nodes benefits patients, highlighting the importance of LNMs in driving cancer progression.
To begin to understand how LNMs could be driving cancer progression, we asked whether it was possible for cancer cells that have colonized lymph nodes to escape the lymph node and seed distant metastatic sites. The main challenge in addressing this question is the ability to identify cancer cells that had been in a lymph node and then were later either circulating in the blood or residing in a distant metastatic site. To overcome this challenge, we relied on a photoconvertible fluorescent protein, Dendra2, which when exposed to specific wavelengths of ultraviolet light can change the color of its emission from green to red [23]. After stably expressing Dendra2 in 3 murine cancer cell lines (4T1 triple negative breast carcinoma, B16F10 melanoma and SCCVII squamous cell carcinoma), each tumor type was grown orthotopically in syngeneic immunocompetent mice [24]. Once the cancer cells spontaneously metastasized from the primary tumor to the lymph node, the primary tumors were surgically resected. We then exposed only the tumor draining lymph node to the photoconverting light and were able to switch the color of the cancer cells in the lymph node from green to red with an efficiency of about 70%. After this photoconversion, the only source of red cancer cells in the animal could be from LNM. We then asked, can we identify red cancer cells from the lymph node circulating in the blood or in distant metastatic sites?
First, we collected all the blood from the animals and using flow cytometry, identified red cancer cells—which must have been in the lymph node at the time of photoconversion—as well as green cancer cells in the blood of animals containing photoconverted 4T1 and B16F10 LNMs, but identified only green cancer cells in the blood of animals containing photoconverted SCVII LNMs [24]. These data provided the first direct evidence that in some models, metastatic cancer cells in lymph nodes can escape the node and enter the blood circulation. Next, we collected the lungs from these animals, made sections through the whole lung and used confocal microscopy to spectrally scan through the tissue. Similar to the blood, we identified red cancer cells, as well as green cancer cells, in the lungs of animals containing photoconverted 4T1 and B16F10 LNMs, but identified only green cancer cells in the lungs of animals containing photoconverted SCVII LNMs [24]. These data provide evidence that the circulating cancer cells that escape the lymph node can disseminate to a distant metastatic organ.
To determine if cancer cells can also escape directly from the primary tumor, we photoconverted a portion of 4T1 primary tumors before they spread to the lymph node and identified circulating red cancer cells in the blood [24]. Thus, there are two sources of circulating cancer cells: those directly from the primary tumor and those from the LNM. Further experiments showed that cancer cells both directly from the primary tumor and those from spontaneous LNM can form metastatic lesions in the lungs [24].
The next question we addressed was how cancer cells escaped the lymph node. There are two possible exit routes. First, cancer cells could leave via the efferent lymphatic vessels, heading to the next echelon nodes upstream. Clinically, it is observed that LNM can often be found in secondary draining lymph node beds and cancer cells have been observed in medullary lymphatic structures in metastatic lymph nodes from extramammary Paget’s disease [25], making escape through the efferent lymphatic vessels plausible. However, by performing intravital microscopy of metastatic lymph nodes, we saw cancer cells migrating toward and interacting with lymph node blood vessels [24]. We established that cancer cells in our mouse model migrate and enter lymph node blood vessels, an additional route of escape for cancer cells out of the lymph node. Finally, looking at a series of LNM from patients with head and neck squamous cell carcinomas, we identified cancer cells inside lymph node blood vessels in 7 out of 19 cases [24], confirming that cancer cells in some patients are able to escape the lymph node through nodal blood vessels.
Our data show it is possible for cancer cells to spontaneously disseminate to lymph nodes and then escape the lymph node to seed another metastatic site. The work also provides evidence that one method of escape from the lymph node is by invasion of the lymph node blood vessels by the cancer cells. Our work does not exclude other methods of dissemination to distant metastasis (e.g., direct invasion of primary tumor blood vessels, which we show is also occurring in the 4T1 model) or other methods of escape from the lymph node (e.g., escape through efferent lymphatic vessels). It also does not suggest that every patient will have distant dissemination occurring from metastatic lymph nodes. However, our data did provide direct evidence for a new route for how cancer cells can spread throughout the body, one that had long been hypothesized [13, 26]. Our work was simultaneously corroborated by data from the laboratories of Dontscho Kerjaschki and Michael Sixt, which showed similar findings using different methods [27].
Other recent work studying LNM in patients also supports the concept that cancer cells can metastasize from lymph node lesions. In a study of 17 colorectal cancer patients, Naxerova et al. built phylogenetic trees to relate the primary tumor, LNMs and distant metastases based on mutational evolution. In this study 6 out 17 patients showed LNM with close mutational relationships to distant metastases [28]. Similarly to our mouse data, this study also shows that some, but not all, patients may have distant metastases seeded by lymph node lesions. In a long-term study of 3329 BC patients that underwent RLN biopsy, Nathanson et al., show that LNMs are predictive of distant metastases, whereas lymphovascular invasion at the primary site is not predictive by itself [29]. These data further suggest that LNM can drive distant cancer progression.
Our work has shown that it is possible for LNM to spread further in the body. However, there is currently no actionable information to identify which patients are at risk for this occurring. Further research into the molecular and physiological mechanisms that drive this process are needed before at-risk patients can be identified and interventions designed. By identifying these mechanisms, we aim to improve outcomes for patients with LNM while balancing the risk of overtreatment by appropriate patient selection.
The genomic evolution of breast cancer metastasis
Lucy Yates
Cancer is primarily caused by DNA damage. The transformation from a normal to cancerous phenotype and from primary to metastatic BC is marked by the accumulation of genetic changes known as somatic mutations. By applying evolutionary principles to genomic sequence data, we have started to uncover the fundamental patterns underlying BC metastasis.
Whole genome sequencing studies have revealed that BC genomes are highly abnormal. In a study of 560 primary BCs, we identified an average of 6214 single base pair substitutions, 665 small insertions and deletions and 140 structural variants per cancer [30]. In a separate study, we found that the progression from primary tumor to the diagnosis of distant metastatic disease was accompanied by an increase in the mutation burden of around a third [31]. Only a tiny fraction of the somatic mutations detected in cancer genomes alter known cancer genes and act as drivers of cancer progression. The majority of mutations are ‘passengers’ with no effect cell fitness; however, they provide great statistical power for identifying mutational signatures and tracing the evolutionary patterns that underlie cancer metastasis.
Cancer subclones
Breast cancers are composed of multiple genetically related subclones [32,33,34,35,36,37].
Subclone composition can vary over space (within the same tumor mass or across metastatic sites) and time (sequential samples). All subclones in a cancer are by definition genetically related, having arisen from a single common ancestor, but individual subclones are distinguished by the existence of private genetic changes. This may be directly observed in single cell studies although technical limitations still mean that these are most reliable for measuring copy number variation rather than point mutations [34, 38]. Subclonal composition can also be inferred indirectly from bulk tissue samples (that typically consist of thousands of millions of mixed cancer cells and normal cells) using bioinformatic approaches that determine the nature of and cellular prevalence of genotypes using features such as variant allele fraction, tumor purity and allele specific copy number information [32, 39]. Multi-region sampling approaches increase our ability to detect subclonal structure. Recent advances in low-input material sequencing (of 100 cells or less) are moving this approach to a new level, providing much higher resolution of subclone composition in both cancer and normal tissues. In the coming years, we expect these approaches to continue to provide important insights into how normal tissue development is subverted during cancer evolution [40, 41].
Different genetic subclones can have different phenotypes and therefore provide the substrate upon which natural selection may act. Indeed, we and others have demonstrated that aggressive cancer traits such as invasion, treatment resistance or metastasis can be mapped back to antecedent subclones earlier in the disease history [31, 32, 41,42,43]. Understanding how these ill-fated subclones relate and differ to their seemingly well-behaved sisters, within the same primary tumor, is a major priority in cancer research. The recent development of a wide range of spatial sequencing approaches is set to provide critical insights into the spatial arrangement and characterization of individual cancer subclones [44,45,46]. Combining these techniques with evolutionary mapping approaches offers an exciting opportunity to isolate and study the most clinically important subclones within the tumor environment.
Trees describe cancer evolution
We can use evolutionary or phylogenetic trees, akin to the ‘tree of life’ famously described by Charles Darwin, to represent diagrammatically, the inferred evolutionary relationships between different cancer subclones (Fig. 1). Phylogenetic tree construction starts with two basic assumptions: mutations can be gained but not lost and each mutation can only occur once [47]. Although there are caveats to these assumptions, these can generally be accommodated. Firstly, we must consider copy number differences between subclones (i.e., where deletions might have ‘lost’ some mutations in one subclone/sample) and secondly, by using as much of the rich genome-wide mutation data as possible rather than relying on limited targeted gene panels that only identify a handful of mutations at best. Using the two principles we can identify ‘clonal’ mutations as those shared by all cancer cells and form the ‘trunk’ of the phylogenenetic tree (Fig. 1). Mutual exclusivity of mutations identifies subclones from different branches, while co-occurrence indicates mutations within the same subclone or a direct lineage. The lengths of branches are typically scaled according to the number of mutations, and therefore, act as a kind of molecular clock that allows us to order branching events in time. Studying the branching patterns that we derive can then help us to understand how different cancers develop and progress.
Metastasis seeding cell dissemination timing
The point at which the Metastasis Seeding Cell (MSC) diverges form the primary tumor has important implications for the use of adjuvant systemic therapy. Take the example where 3 driver mutations have accumulated within a primary tumor up to the point where it is diagnosed. A precision medicine treatment plan would have to be based on the 3 mutations seen in the primary tumor, but we do not know how representative this is of the potential MSCs at this time point. In the situation where the MSC left the primary tumor recently, i.e., the late branching scenario, it is likely that these cells share the same 3 driver mutations and the primary tumor can be considered a good proxy. However, in the case where the MSC departed from the tumor a long time ago, i.e., an early branching scenario, these cells are less likely to contain the same driver mutations seen in the primary tumor and furthermore, they have had significant time to accumulate novel driver mutations that we will not be able to detect. The primary tumor in this situation is a poor proxy, providing inaccurate and incomplete information about the cancer cells that we wish to eliminate.
When we applied this principle to 17 cases of metastatic or relapsed BCs, we found that most MSCs diverged from the primary tumor late in evolution (on average 87% in primary tumor molecular time). This was a phenomenon shared across distant metastases, local relapses and synchronous LNM. Importantly, the majority of driver mutations in the primary tumor were also present in 100% of the cells in the metastasis indicating that targeted therapies based on the primary tumor genomic profile would be relevant to the metastasis.
Some studies appear to contradict our findings, suggesting that cancer cell dissemination occurs, or at least can occur early—even from pre-invasive lesions [48,49,50,51,52]. Notwithstanding the need to extend our analyses to larger sample series and different BC subtypes, we do suspect that one potential confounding factor is that these studies have tended to rely on less robust technologies for inferring clonal relatedness. Indeed, much of the support for early dissemination is derived from disseminated tumor cell (DTC) studies that defined bone marrow cells as DTCs based on morphology and cytokeratin expression. Importantly, recent single cell sequencing of DTCs and exome sequencing/genome-wide copy number profiling of the primary tumor revealed that only a fraction of presumed DTCs are actually clonally related to the primary tumor [53]. Further work linking these DTCs to the subsequent metastasis is needed to determine if real-time characterization could be helpful for treatment scheduling during the relapse free window.
Future perspectives
Our findings to date have identified that metastases typically arise from a detectable subclone in the primary tumor. To date, we have not been able to demonstrate that the driver mutations that dominate the metastasis are pre-selected in the primary tumor. Furthermore, we have had little success in identifying the anatomical localization of the subclones that are most closely related to a future metastasis. A critical question remains to be answered: “Is there something special about the metastasis seeding subclone?”.
The emergence of spatially resolved approaches, such as laser capture microscopy with low-throughput sequencing and in-situ sequencing of somatic mutations, are expected to allow much closer interrogation of primary tumors and pin-pointing of ill-fated subclones [41, 44]. These approaches will be essential for identifying subclone specific associations with tumor micro-environmental factors that are likely to be important determinants of subclone fate.
Mitochondrial-nuclear exchange (MNX) mice reveal contributions of mitochondrial genetics to cancer metastasis
Thomas C. Beadnell, Adam D. Scheid, and Danny R. Welch
Introduction
The ability of cancer cells to metastasize requires coordinated expression of multiple genes [54,55,56,57,58,59,60,61,62,63,64] and the efficiency of metastasis is contributed to by multiple quantitative trait loci (QTL) [65] alleles that influence a trait [66] in combination with environmental factors.
Because of its comparative size, most studies with QTL examine the nuclear genome, ignoring the mitochondrial genome. However, in the context of cancer, mitochondria are key players, which were first identified by Warburg with regard to their roles in metabolism [67,68,69], but the definitive mechanisms by which mitochondrial DNA (mtDNA) contribute to cancer phenotypes has been understudied and underappreciated. It is critical to emphasize that mtDNA polymorphisms are likely metastasis modifiers rather than drivers per se. mtDNA QTL would combine nuclear and mitochondrially-encoded genes to regulate cancer severity and metastasis [70,71,72]. Given the miniscule size of the mitochondrial genome (~ 16 kb) to the nuclear genome (~ 3 × 104 kb), could the hypothetical existence of mtDNA metastasis QTL be reasonable? Despite being relatively understudied, the answer is that it is likely (reviewed in [73,74,75]).
The classic experiment demonstrating the existence of metastasis QTL in the nuclear genome was performed in the laboratory of Kent Hunter, who crossed female mice from multiple Mus musculus strains to male transgenic mice with the oncogenic polyomavirus middle T antigen (PyMT) under the control of the mouse mammary tumor virus (MMTV) promoter (FVB/N-TgN(MMTV-PyMT)) [76]. Critically, these mice were made on the FVB/N genetic background [77]. Crossing inbred strains with FVB/N-TgN(MMTV-PyMT) resulted in differential tumor latency and metastatic burden in the first filial generation [76]. Using backcrosses and comprehensive genetic screens, his group has identified metastasis modifiers [78,79,80], many of which have also been observed in human BCs [81,82,83].
However, an alternative interpretation of their data was recognized because their experimental design crossed females of those various strains to male FVB/N-TgN(MMTV-PyMT) mice. Since mtDNA is maternally transmitted, the possibility existed that mtDNA could also be a metastasis modifier QTL. A growing volume of data implicates mitochondria in cancer and metastasis (reviewed in [73, 74]). Specifically, Kaori Ishikawa and colleagues demonstrated that transfer of mitochondria from highly metastatic cells to poorly metastatic recipient cancer cells enhanced metastatic potential. A reciprocal experiment also lowered metastatic efficiency [84].
We began exploring approaches to study contributions of mtDNA to metastasis. Unfortunately, the unique characteristics of mtDNA presented numerous experimental challenges (reviewed in [73, 85]). Briefly, mtDNA is present in 100s to 1000s of copies per cell, making alteration of all mtDNA copies currently impossible even with the most advanced methods [86]. Therefore, introduction of heterogeneous heteroplasmy would confound effects of the alteration. The use of cybrids or transmitochondrial mice involves prior exposure to mutagens, which could potentially introduce mutations that would complicate interpretation. Likewise, generation of conplastic mice, while not exposing to mutagenic agents, involves backcrossing for at least 10 generations to achieve 99.9% nuclear DNA (nDNA) purity with different mtDNA composition [87]. While conplastic mice mostly eliminate disparate nDNA as a confounding variable, the number of necessary backcrosses increases the likelihood that nDNA recombination has occurred.
To circumvent the issues associated with the above approaches, we developed the mitochondrial-nuclear exchange (MNX) mouse model [88] to assess whether there are metastasis modifier QTLs in the mitochondrial genome. Crossing MNX female mice with mammary cancer transgenic mouse models demonstrated a driver-dependent regulation of metastasis. Moreover, injection of syngeneic cancer cells into MNX mice revealed non-cell autonomous contributions of the mitochondrial genome to metastasis. mtDNA polymorphisms alter immune profiles as well as other changes to the tumor microenvironment, which affect metastatic efficiency. Together, these data demonstrate that mtDNA indeed contains QTL for cancer metastasis.
Generating MNX mice did not involve backcrossing or mutagen exposure. They were made using micropipette transfer of pronuclei from embryos into enucleated cytoplasts from another mouse strain (i.e., having mitochondria from another strain) [88, 89]. The resulting hybrid embryos transplanted into pseudopregnant mice resulted in progeny, which are homoplasmic, phenotypically normal and fertile. The resulting colonies have been maintained for more than a decade and have been very stable.
Mitochondrial DNA alters metastasis in a cell autonomous manner
Female MNX mice on the FVB genetic background having FVB mitochondria (FF), BALB/c mitochondria (FB) or C57BL/6 mitochondria (FC) were crossed with MMTV-PyMT mice. The nuclear genomes were identical except for the transgene. Latency to first tumor formation and metastasis were measured. FF crosses developed tumors as expected while time to first tumor was accelerated in the FB crosses and slowed in FC progeny. Almost identical to the data from the Hunter crosses, pulmonary metastatic burden (i.e., cumulative volume of metastases and metastasis size) differed. The number of metastases arising was relatively unaffected [90].
Subsequently, the same MNX mouse strains were crossed to male FVB/N-Tg(MMTVneu) mice, which overexpress the wild-type Her2/neu oncogene under the control of the MMTV promoter [91]. Tumor latency and metastases were quantified. Results in FF and FC progeny were similar to those seen in the PyMT model. However, Her2 FB mice developed tumors later and had lower metastatic burden than FF mice [91]. This contrasted with the PyMT crosses, where FB mice exhibited more rapid primary tumor formation than FF mice.
Taken together, these two studies demonstrate that mtDNA polymorphisms affect both tumorigenicity and metastasis. The changes are dependent upon contributions of both the nuclear genome and the mitochondrial genome, exactly what one would expect for QTL. Additionally, Brinker and colleagues showed that using male MNX mice crossed to MMTVneu females (i.e., mtDNA from the MNX mice would not be inherited in the progeny) resulted in no difference in the tumor behaviors.
Additional evidence that the mitochondrial genome could impact tumor formation was obtained by Vivian et al. [92]. Mice unexposed to carcinogen treatments were allowed to age naturally, and the incidence and location of spontaneous autochthonous tumors were recorded. Although underpowered for some strains of MNX mice, a tumor protective effect of C57BL/6 mtDNA was observed.
Since metastasis involves multiple genes to be coordinately expressed and only a very small fraction of those genes are encoded in the mtDNA, we reasoned that SNP in mtDNA could influence gene expression in the nuclear genome. By altering the nuclear epigenome via changes in cytosine methylation or modifying histones, gene expression and corresponding phenotypes could be affected. We performed whole genome methylation sequencing coupled with RNA sequencing and determined that there were selective changes in the location of methylation in the mouse genome and that some of those changes occurred concomitant to changes in gene expression [93]. In a more recent study, we measured 4 common histone marks using ChIP-Seq and again found a selectivity in location of histone modifications. Many of the methylation marks and histone marks are in similar regions, suggesting that mtDNA somehow dictates how the nuclear genome operates.
Mitochondrial DNA alters metastasis in a non-cell autonomous manner
Although having previously demonstrated that nuclear [87] and mitochondrial [64, 69] genetics could alter metastatic efficiency, it was recognized that all cells in F1 progeny would inherit mtDNA from the MNX mother. We therefore investigated the role of mtDNA in the tumor microenvironment and its impact on tumor development. Using 2 mammary carcinoma and 2 melanoma cell lines, we were able to keep the mtDNA constant within the cancer cells and injected the cells into either WT or MNX mice. Orthotopic tumor growth was not significantly altered; however, formation of experimental metastasis was significantly affected. These experiments utilized wild-type C57BL/6 (CC), C3H/HeN (HH) and reciprocal MNX mice C57BL/6 nDNA:C3H/HeN mtDNA (CH) and C3H/HeN nDNA:C57BL/6 mtDNA (HC) into which cells were injected either orthotopically or into the lateral tail vein. Critically, wild-type mice and their nuclear-matched MNX counterparts were major histocompatibility matched; so, overt transplant rejection mechanisms were eliminated as an experimental variable. Clearly the mtDNA in stromal compartments influenced behavior of tumor cells [94]. The question is: how?
In all mouse strains utilized for the above studies, only a handful of mtDNA polymorphisms have been identified [71]. Changes have been reported in protein encoding genes as well as in transfer RNA [71, 92]. Polymorphisms in mtDNA genes encoding electron transport chain proteins implicate metabolism and previous studies found oxygen consumption to be different between MNX strain mammary tissue [69] and cardiomyocytes [68]. However, parameters associated with metabolism (i.e., oxygen consumption, extracellular acidification, and mitochondrial load) showed no correlation with changes in tumor cell behavior. Metabolomic analyses comparing the MNX mice also identified metabolite differences among MNX strains. Nothing jumped out as an explanation for the different tumor cell phenotypes observed. The caveat to dismissing differences in metabolism as a mechanism for stromal manipulation of metastasis is that the measurements were done in vitro and in tissues that are not necessarily the most appropriate locations of tumor cells.
A byproduct of electron transport is generation of reactive oxygen species (ROS). We tested the hypothesis that altered ROS in MNX mice might be responsible for the changes in metastatic potential. Using MitoTEMPO, a scavenger for mitochondria-derived ROS, the metastasis-promoting effects of C3H/HeN mtDNA were reduced. While the results are consistent with the hypothesis, reciprocal experiments (i.e., increasing ROS in C57 mice) could not be done because of multiple off-target effects [91].
A key stromal contributor to tumorigenicity and metastasis is the immune system. Based upon linkages between mitochondrial genetics, metabolism, and observations that metabolism is closely linked to immune differentiation and polarization [95], we studied whether baseline immune profiles and functionalities could explain results in the non-cell autonomous studies. Most differences between mouse strain immune profiles were found to be regulated by nDNA. Interestingly, many of the lymphocyte populations were not dramatically altered. However, macrophage differentiation (and perhaps polarization state) was significantly altered by SNP in the mtDNA [96]. Detailed studies are underway to refine the changes and to directly test which macrophage subpopulations are most relevant to the phenotypic changes reported for tumorigenicity and metastasis.
Relevance and perspective
The co-evolution of mtDNA and nDNA [97] demonstrate clear communication between the two genomes. Metabolic adaptations to changing climates encountered by ancient humans migrating from Africa provided selective pressures dictating divergence from the original mtDNA genotype contained in ‘Mitochondrial Eve’ [98]. Those physiological adaptations mediated by mtDNA haplogroups have played critical roles in human evolution. Moreover, relevant to this discussion, they also contribute to disease pathologies and racial health disparities [73, 99]. Individuals with certain mtDNA haplogroups have increased predisposition to certain cancers compared to people with other mtDNA haplogroups [85, 90, 91]. Some adaptive advantages in some mtDNA variants resemble oncogenic mitochondrial functions [100, 101].
Clinicians have long known that patient race/ethnicity contributes to cancer incidence rates and survival. For instance, triple negative BC is more prevalent in African American and Hispanic Caucasian women [102, 103] who exhibit higher proliferation rates, increased angiogenesis markers, higher grade, higher rates of LNM and worse overall survival compared to non-Hispanic women [104,105,106,107,108,109,110,111,112,113]. While economic and modifiable factors contribute to outcome disparities, incidence and progression differences persist even after controlling for socioeconomic factors [106, 109,110,111,112,113]. To date, no nuclear genomic explanations have been found comparing patients from different races [114,115,116,117]. We posit that the impact of contributions from each QTL that define race are masked by uncontrollable variables in analyses. Use of the MNX model, which isolates mtDNA as an experimental variable, allowed us to gain a foothold into the mechanistic underpinnings of mitochondrial contributions to metastasis.
In the context of metastasis, it is critical to remember that tumor cells encounter multiple different microenvironments throughout their journey from the primary tumor to distant sites. Our data (Table 1) exploring the roles of mtDNA in the MNX mice support the notion that mitochondria perform critical functions in the receipt of information as well as the conveyance of neoplastic cell signals to the microenvironment. While the mitochondrial genome is small, it leverages the much larger nuclear genome to affect neoplastic cell behavior. Mitochondria are mediators of outside-in and inside-out cellular communication [118,119,120]. They sense changes in the microenvironment and signal to the nucleus the requirement of a cell to adapt. As yet, the mediators of mitochondrial signaling are poorly defined. Identifying those signals represents an as yet untapped opportunity for prognostic information from the mitochondria and therapeutic targets within the mitochondrial genome. There is currently no direct evidence that mitochondrial genetics play a role in human breast cancer. Efforts to create Synteny maps between mouse mitochondrial and human mitochondrial genomes and a comprehensive map of mtDNA polymorphisms are underway.
Deconstructing collective metastasis: emergent multicellular mechanisms supporting metastatic colonization
Emma D. Wrenn and Kevin J. Cheung
A large number of tumor cells reaching distant sites will die and never grow into a clinically detectable metastasis [121]. Disseminated tumor cells encounter inhospitable stromal matrices, cell types, and paracrine signals different from their organ of origin. In addition, they are actively eliminated by immune cells, such as natural killer (NK) and T-cells. Diverse mechanisms have been described that enable tumor cells to overcome these barriers: entry into a stem cell-like state, epithelial-to-mesenchymal and mesenchymal-to-epithelial transitions, genetic mutation, and co-option of the native microenvironment for example [122,123,124]. Here, we focus on an emerging mechanism by which cancer cells increase their chance of success—metastasizing as cohesive clusters of cells, also known as collective metastasis.
Much research has been conducted on the mechanisms by which single tumor cells metastasize. However, accumulating studies have shown that tumor cells can also metastasize as multicellular aggregates, and do so with much higher efficiency than solitary cancer cells [125, 126]. A key foundation for this concept originates from seminal experimental metastasis studies performed in the mid-twentieth century. Investigators injected lung cancer, melanoma, and fibrosarcoma tumor cells into mice as either clusters or filtrated single cells [127,128,129]. In each instance the clusters had significantly greater success forming new metastases. Since then, a number of studies in different tumor types and metastasis models have demonstrated greater metastasis formation by injected clusters compared to single cells [130,131,132,133], with for example ~ 15-fold higher lung metastasis seeding rates by clusters in pancreatic cancer [134] and up to 500-fold higher rates in BC [135, 136] compared with equal numbers of single cells.
These findings are buttressed by recent studies tracing the clonal composition of spontaneous metastases in mouse models. Multiple groups have used multi-color fluorescent tumor cell models [131, 132, 134, 135, 137,138,139] or deep sequencing [140, 141] to demonstrate that transplantable primary tumors produce metastases that are polyclonal, that is seeded by multiple clones present in the primary tumor. A large fraction of metastases were found to be polyclonal in several of these models; 48–53% using BC cell lines [137], 54% using BC PDXs [131], ~ 80% of large lesions to the peritoneal wall and diaphragm using KPCX pancreatic cancer cells [134], and over 97% using the MMTV-PyMT breast tumor model [135]. A caveat to these experiments is that polyclonal metastases could arise from serial seeding by single cells [142]. Importantly, multiple studies have also conducted experiments excluding large contributions from serial seeding, which indicates that polyclonal metastases in their models arise primarily from tumor cell clusters [132, 134, 135, 137].
Deep sequencing studies comparing primary and metastatic genomic alterations have also supported a role for polyclonal seeding in human tumors. Polyclonal seeding has been reported in colorectal cancer [143,144,145,146], intrahepatic cholangiocarcinoma [147], gastric cancer [148], and prostate cancer [149]. Two recent studies of metastatic BC patients identified polyclonal metastases in 63% to 73% of patients [150, 151]. This clonal diversity raises the possibility that multiple subclones in polyclonal metastases could cooperate to increase one another's fitness, instead of responding to selective pressures from the tumor microenvironment purely on their cell-intrinsic properties. This kind of interclonal cooperativity has been observed in mouse models of cancer [139, 152, 153]. But further research is needed to understand how diverse genetic compositions in tumor cell clusters contribute to different stages of collective metastasis. Importantly, direct isolation of tumor cell clusters (CTCs) from the blood of metastatic patients has provided further clinical evidence for collective metastasis [154,155,156,157]. Moreover, CTC clusters are correlated with poorer patient prognosis across the most common cancer types [137, 158,159,160,161,162,163,164,165,166,167]. Together these diverse experimental and clinical studies indicate that tumor cell clusters are potent metastatic seeds.
Despite these many studies establishing the impact of cluster-based metastasis, the molecular mechanisms underpinning their efficient colonization are much less understood. One possible explanation is physical entrapment; it could be that clusters’ large size simply results in rapid arrest in the circulation. However, a recent study demonstrated that patient-derived CTC clusters in the blood could rearrange into single-file chains, which survive passage through narrow capillaries [168]. Additionally, another study in zebrafish models found no significant difference in the rate of extravasation between single and clustered CTCs [130]. These findings do not rule out the importance of physical entrapment. But, as discussed below, recent studies also show that clustering of tumor cells alters their properties beyond mechanical trapping in ways that support metastasis.
The metastatic potential of a single cell can be conceived as a balance of factors promoting or hindering that cell’s ability to colonize other tissues. But to understand cluster-based metastasis we have to consider an additional dimension, the interactions amongst the cells in that cluster. Indeed, a number of recent studies have coalesced around a common theme: tumor cell clustering induces global shifts in key cell states promoting survival, growth, and ultimately colonization. For instance, clusters may be able to resist attrition in the critical early stage of colonization (Fig. 2). Shortly after tail vein injection, tumor cell clusters arrested in the lungs had lower rates of apoptosis than single BC cells [137]. Recently, we observed that while both clusters and single cells were able to reach the lungs immediately after tail vein injection, clusters persisted while single cells were mostly cleared within 48 h [136]. Additionally, time lapse imaging in the presence of an apoptosis biosensor revealed that clusters generated from 8 of 10 human BC tumor samples were significantly more apoptosis resistant than single cells in 3-dimensional culture [136]. A recent clinical study of CTCs from small-cell lung cancer patients also observed a reduction in apoptosis in CTC clusters vs. single cells [159]. Another study using BC tumor cell clusters from patient-derived xenografts found that homophilic CD44 adhesions promote cluster-based metastasis, possibly through PAK2-mediated increases in cell survival [131]. Together, these suggest that likelihood of survival during colonization is a key difference between clusters and single tumor cells.
Some of the survival advantage of clusters appears to depend upon resisting ROS, a critical challenge for disseminating tumor cells [169, 170]. A recent study found that E-cadherin mitigates TGFb-dependent ROS stress in tumor cell clusters, facilitating their survival at metastatic sites [171]. Clusters have also been shown to induce mitophagy to limit ROS and increase cell survival [172]. Clustering may play a role in non-apoptotic forms of cell death as well. Recent reports show that multicellular aggregates are more resistant to ferroptosis [173, 174], an iron-dependent form of non-apoptotic cell death characterized by ROS accumulation [175]. Tumor cell clusters generated Nectin-4 dependent Src signaling, which buffered against ferroptotic lipid peroxidation [173, 174].
Cell–cell adhesion could also help tumor cell clusters successfully evade attack from the immune system. Tumor cell clusters were recently demonstrated to be more resistant to NK cells than single cells via downregulation of NK cell activating ligands [132]. NK cells play a key role in the immunosurveillance and targeting of metastasis, and NK tumor cell infiltration and activation often correlate with better patient prognosis [176,177,178]. Interestingly, a number of cell–cell adhesion molecules function as NK cell inhibitory ligands, suggesting that NK cells may more effectively target solitary tumor cells or cells that have undergone complete epithelial mesenchymal transition EMT [132, 179, 180]. Clusters may even utilize the immune system to promote survival and growth; heterotypic clustering of CTCs with neutrophils has been found to promote cell cycle entry in tumor cells [181] and clusters may shift NK cells into a more metastasis-promoting state [182].
Beyond survival, clustering generates changes in signaling and cell state supportive of metastatic outgrowth. Tumor cell clusters are more proliferative and more stem cell-like than single tumor cells [131, 135, 136, 183]. How does cell–cell contact generate these pro-metastatic states? An attractive hypothesis is that cell–cell adhesion molecules directly generate pro-proliferative signaling through cross-talk with cellular signaling pathways that regulate growth [184]. On the other hand, many studies have shown that cell–cell adhesion induces contact inhibition, restricting cell proliferation [185, 186]. In fact, the same cell–cell adhesion molecule can either promote or inhibit metastasis depending on context. For example, E-cadherin is required for successful metastasis in a mouse model of BC and promotes cell survival [171]. But in the same mouse model, E-cadherin activating antibodies inhibit metastasis, suggesting that E-cadherin’s role in metastasis may be tunable [187]. At present, the balance between pro- and anti-metastatic signals downstream of cell–cell adhesion molecules remains unresolved and may be highly context-dependent between different cell types, tumor types, and model systems.
Cell–cell adhesions can also promote metastasis without direct intracellular signal transduction. Instead, clustering can induce 3-dimensional architectural changes that generate collective signaling. Our group recently characterized nanolumina within tumor cell clusters (Fig. 3)—hollow spaces between cells, sealed at either end by electron dense cell–cell junctions [136]. They are often lined by microvilli-like protrusions, which can interdigitate between neighboring cells and provide high surface area for intercellular interactions. Nanolumenal junctions were selectively permeable, thereby controlling the composition of nanolumina and shielding them from the microenvironment.
Tumor cell clusters contain intercellular nanolumina that concentrate signaling molecules. Left, transmission electron microscopy of an MMTV-PyMT tumor cell cluster. Between tumor cells we observe intercellular cavities lined by microvilli-like protrusions and sealed by cell–cell junctions. We find that the growth factor epigen is trafficked to and concentrated within nanolumina, resulting in cooperative pro-growth signaling during collective metastasis
Importantly, we found that nanolumina formed sites for intercellular communication. We identified a growth factor, epigen, which was trafficked into nanolumina where it achieved concentrations > 5000-fold higher than those outside the cluster. Importantly, epigen suppression profoundly reduced both primary tumor growth and metastatic outgrowth. By secreting a growth signal into a shared space, clusters create collective, non-cell autonomous signaling to promote proliferation during metastasis. In this instance, cell–cell adhesions do not directly transduce a signal but instead regulate a signaling molecule’s concentration by trapping it within intercellular cavities. This 3-dimensional topology-dependent collective signaling is an emergent feature of clusters, as it is not possible in single cells lacking cell–cell adhesion. Our findings indicate that the physical architecture of a multicellular cluster provides an additional layer of regulation during metastasis, and a mechanism for robust intercellular signaling.
Importantly, we also observed nanolumina in freshly isolated human tumor cell cluster samples. Nanolumina with restricted permeability were also present in an aggressive subset of basal-like 2 (BL2) triple negative BC cell lines which highly express epigen, but not in mesenchymal-like triple negative BC cell clusters. BL2 BCs have poor treatment response and overexpress growth factors and myoepithelial genes, whereas mesenchymal-like BCs overexpress genes related to EMT and cell motility [188,189,190,191]. RNA-sequencing analysis demonstrated that BL2 nanolumina-containing cell lines highly express epithelial genes and genes associated with branching morphogenesis during early development. This suggests that epigen expression and nanolumina may be linked to epithelial identity and could have a role during epithelial development that is exploited by metastatic tumor cell clusters.
Treatments are critically needed to effectively eradicate metastases and suppress the metastatic process [192]. An understanding of the mechanisms underlying tumor cell clusters’ metastatic potency could pave the way toward therapies targeting their metastatic advantages. One approach is to target the adhesion molecules holding clusters together. Indeed, disrupting cell–cell adhesion can repress metastatic potential [131, 137, 171, 183, 193]. However, cell–cell adhesion molecules are often highly expressed in normal tissues, which could narrow the therapeutic window unless tumor specific properties or activation states are identified. Another approach is to target the cooperative signals generated by clusters within nanolumina. However, much remains to be learned about the optimal strategy to target these collective signaling compartments in practice. Our finding that BL2 but not mesenchymal-like BCNot cells contain nanolumina also suggests that the relative contribution and mechanisms of collective metastasis could differ widely between cancers, subtypes, and individual patients. Future studies are essential to expand on these mechanistic insights into collective metastasis, to rigorously assess their relevance to specific subgroups of patients, and ultimately to translate these findings into the clinic.
Conclusions
Breast cancer metastasis is a complex process requiring many interacting components, anatomic, cellular and molecular/genetic. Mechanical factors play a part in sentinel lymph node metastasis, and breast cancer cells may gain direct access to the systemic circulation by invasion into veins in the node. Polyclonal multicellular tumor cell aggregates metastasize with much higher efficiency than solitary cancer cells, partly because large size clusters might simply arrest in the circulation by physical entrapment. DNA studies show that some breast cancers metastasize to systemic sites without first entering lymph nodes, suggesting the secretion of gene-induced functional proteins, cytokines and/or peptides, present in some clones but not others, cause direct invasion into blood vessels in the primary breast tumor. It is more likely that tumor cells invade lymphatic rather than blood vessel capillaries at the primary site.
Identification of activated genes and other molecular markers that are important in systemic breast cancer metastasis would be valuable to clinicians in many ways. For example, a more accurate molecular fingerprint of tumor cells identified by needle biopsy of the primary tumor could subclassify patients into those who might not benefit from sentinel node biopsy. Innovation and adaptation will be necessary for continued relevance of SLN biopsy, which will likely take place in several areas. There would need to be incremental advances helping to optimize an already good technique. Other avenues of research could advance SLN utility even further, including developing more refined criteria for selecting patients for SLN biopsy. Some advances may make SLN biopsy redundant; for example, we can imagine non-surgical therapies that will kill tumor in lymph nodes in which case removing them might become unnecessary. We still don’t know for certain whether removing the SLN treats breast cancer and that will require further research.
In our current practice most patients with no clinical evidence of axillary lymph node metastasis who undergo lumpectomy for invasive disease are eligible for sentinel lymph node biopsy. Only one in four patients in this cohort are found to have node metastasis. This means that 75 of every 100 patients with negative lymph nodes might safely avoid the operation of sentinel node biopsy altogether. Revealing currently unknown molecular markers might help us identify these patients upfront. Seven in every seventy-five patients without node metastases develop systemic metastases. More accurate and advanced molecular signatures might allow us to focus adjuvant systemic therapy on only this small proportion of patients, which would be a major advance on our current use of commercially available molecular subtypes used to decide which patients should be offered chemo, targeted and hormonal therapies. The majority of patients could perhaps safely avoid uncomfortable systemic and loco-regional treatments without jeopardizing their chances of survival.
The development of anti-metastatic therapies is an obvious future direction for researchers who identify metastasis-related molecular markers. Hypothesis-directed research would first need to connect the clinically important molecular markers to the mechanisms of metastasis. For example, although we know that patients with HER-2/neu positive breast cancers are at higher risk for systemic metastasis, there is no direct proof that HER-2/neu is involved in the process of metastasis. Similarly, triple negative breast cancer carries a risk of poorer survival than hormone-receptor rich tumors, but no-one has yet identified any related molecular or cellular mechanisms to account for this difference. The relative contributions and mechanisms of breast cancer metastasis might vary widely between cancers, subtypes, and individual patients and we are at the dawn of those discoveries.
The precise biochemical and molecular/genetic signals that prompt lymphatic or blood vessel invasion, and methods by which tumor cells survive the hazardous journey through the blood stream, and how exactly those cells settle into their new environment within the body, are still under investigation but it seems likely that when we discover more details about these mechanisms we will be better able to target steps in metastasis formation. When we reach those goals we may be able to eliminate metastases altogether, goals that are both urgent and possible and will augur a new era of breast cancer treatment.
Data availability
Obtained through existing institutional guidelines.
References
Fisher B, Jeong JH, Anderson S, Bryant J, Fisher ER, Wolmark N (2002) Twenty-five-year follow-up of a randomized trial comparing radical mastectomy, total mastectomy, and total mastectomy followed by irradiation. N Engl J Med 347:567–575. https://doi.org/10.1056/NEJMoa020128
Baclesse F (1949) Roentgen therapy as the sole method of treatment of cancer of the breast. Am J Roentgenol Radium Ther 62:311–319
Sledge GW, Mamounas EP, Hortobagyi GN, Burstein HJ, Goodwin PJ, Wolff AC (2014) Past, present, and future challenges in breast cancer treatment. J Clin Oncol 32:1979–1986. https://doi.org/10.1200/JCO.2014.55.4139
Fisher B, Jeong JH, Dignam J, Anderson S, Mamounas E, Wickerham DL, Wolmark N (2001) Findings from recent National Surgical Adjuvant Breast and Bowel Project adjuvant studies in stage I breast cancer. J Natl Cancer Inst Monogr. https://doi.org/10.1093/oxfordjournals.jncimonographs.a003463
Fidler IJ (1973) Selection of successive tumour lines for metastasis. Nat New Biol 242:148–149. https://doi.org/10.1038/newbio242148a0
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144:646–674. https://doi.org/10.1016/j.cell.2011.02.013
Park KU, Chen Y, Chitale D, Choi S, Ali H, Nathanson SD, Bensenhaver J, Proctor E, Petersen L, Loutfi R, Simonds A, Kuklinski M, Doyle T, Dabak V, Cole K, Davis M, Newman L (2018) Utilization of the 21-gene recurrence score in a diverse breast cancer patient population: development of a clinicopathologic model to predict high-risk scores and response to neoadjuvant chemotherapy. Ann Surg Oncol 25:1921–1927. https://doi.org/10.1245/s10434-018-6440-7
Edge SB, Byrd DR, Compton CC, Fritz AG, Greene FL, Andrew Trotti I (eds) (2010) AJCC cancer staging manual, 7th edn. Springer, New York
Ferris RL, Lotze MT, Leong SP, Hoon DS, Morton DL (2012) Lymphatics, lymph nodes and the immune system: barriers and gateways for cancer spread. Clin Exp Metastasis 29:729–736. https://doi.org/10.1007/s10585-012-9520-2
Kawada K, Taketo MM (2011) Significance and mechanism of lymph node metastasis in cancer progression. Cancer Res 71:1214–1218. https://doi.org/10.1158/0008-5472.CAN-10-3277
Cady B (2007) Regional lymph node metastases; a singular manifestation of the process of clinical metastases in cancer: contemporary animal research and clinical reports suggest unifying concepts. Ann Surg Oncol 14:1790–1800. https://doi.org/10.1245/s10434-006-9234-2
Cascinelli N, Morabito A, Santinami M, MacKie RM, Belli F (1998) Immediate or delayed dissection of regional nodes in patients with melanoma of the trunk: a randomised trial. WHO Melanoma Program Lancet 351:793–796. https://doi.org/10.1016/s0140-6736(97)08260-3
Halsted WS (1907) I. The results of radical operations for the cure of carcinoma of the breast. Ann Surg 46:1–19. https://doi.org/10.1097/00000658-190707000-00001
Starz H, Balda BR, Kramer KU, Buchels H, Wang H (2001) A micromorphometry-based concept for routine classification of sentinel lymph node metastases and its clinical relevance for patients with melanoma. Cancer 91:2110–2121
Nathanson SD, Shah R, Rosso K (2015) Sentinel lymph node metastases in cancer: causes, detection and their role in disease progression. Semin Cell Dev Biol 38:106–116. https://doi.org/10.1016/j.semcdb.2014.10.002
Giuliano AE, Hunt KK, Ballman KV, Beitsch PD, Whitworth PW, Blumencranz PW, Leitch AM, Saha S, McCall LM, Morrow M (2011) Axillary dissection vs no axillary dissection in women with invasive breast cancer and sentinel node metastasis: a randomized clinical trial. JAMA 305:569–575. https://doi.org/10.1001/jama.2011.90
Galimberti V, Cole BF, Zurrida S, Viale G, Luini A, Veronesi P, Baratella P, Chifu C, Sargenti M, Intra M, Gentilini O, Mastropasqua MG, Mazzarol G, Massarut S, Garbay JR, Zgajnar J, Galatius H, Recalcati A, Littlejohn D, Bamert M, Colleoni M, Price KN, Regan MM, Goldhirsch A, Coates AS, Gelber RD, Veronesi U (2013) Axillary dissection versus no axillary dissection in patients with sentinel-node micrometastases (IBCSG 23–01): a phase 3 randomised controlled trial. Lancet Oncol 14:297–305. https://doi.org/10.1016/S1470-2045(13)70035-4
Jagsi R, Chadha M, Moni J, Ballman K, Laurie F, Buchholz TA, Giuliano A, Haffty BG (2014) Radiation field design in the ACOSOG Z0011 (alliance) Trial. J Clin Oncol 32:3600–3606. https://doi.org/10.1200/JCO.2014.56.5838
Donker M, van Tienhoven G, Straver ME, Meijnen P, van de Velde CJ, Mansel RE, Cataliotti L, Westenberg AH, Klinkenbijl JH, Orzalesi L, Bouma WH, van der Mijle HC, Nieuwenhuijzen GA, Veltkamp SC, Slaets L, Duez NJ, de Graaf PW, van Dalen T, Marinelli A, Rijna H, Snoj M, Bundred NJ, Merkus JW, Belkacemi Y, Petignat P, Schinagl DA, Coens C, Messina CG, Bogaerts J, Rutgers EJ (2014) Radiotherapy or surgery of the axilla after a positive sentinel node in breast cancer (EORTC 10981–22023 AMAROS): a randomised, multicentre, open-label, phase 3 non-inferiority trial. Lancet Oncol 15:1303–1310. https://doi.org/10.1016/S1470-2045(14)70460-7
Faries MB, Thompson JF, Cochran AJ, Andtbacka RH, Mozzillo N, Zager JS, Jahkola T, Bowles TL, Testori A, Beitsch PD, Hoekstra HJ, Moncrieff M, Ingvar C, Wouters M, Sabel MS, Levine EA, Agnese D, Henderson M, Dummer R, Rossi CR, Neves RI, Trocha SD, Wright F, Byrd DR, Matter M, Hsueh E, MacKenzie-Ross A, Johnson DB, Terheyden P, Berger AC, Huston TL, Wayne JD, Smithers BM, Neuman HB, Schneebaum S, Gershenwald JE, Ariyan CE, Desai DC, Jacobs L, McMasters KM, Gesierich A, Hersey P, Bines SD, Kane JM, Barth RJ, McKinnon G, Farma JM, Schultz E, Vidal-Sicart S, Hoefer RA, Lewis JM, Scheri R, Kelley MC, Nieweg OE, Noyes RD, Hoon DSB, Wang HJ, Elashoff DA, Elashoff RM (2017) Completion dissection or observation for sentinel-node metastasis in melanoma. N Engl J Med 376:2211–2222. https://doi.org/10.1056/NEJMoa1613210
Poortmans PM, Collette S, Kirkove C, Van Limbergen E, Budach V, Struikmans H, Collette L, Fourquet A, Maingon P, Valli M, De Winter K, Marnitz S, Barillot I, Scandolaro L, Vonk E, Rodenhuis C, Marsiglia H, Weidner N, van Tienhoven G, Glanzmann C, Kuten A, Arriagada R, Bartelink H, Van den Bogaert W, Oncology ER, Breast Cancer G (2015) Internal mammary and medial supraclavicular irradiation in breast cancer. N Engl J Med 373:317–327. https://doi.org/10.1056/NEJMoa1415369
Whelan TJ, Olivotto IA, Parulekar WR, Ackerman I, Chua BH, Nabid A, Vallis KA, White JR, Rousseau P, Fortin A, Pierce LJ, Manchul L, Chafe S, Nolan MC, Craighead P, Bowen J, McCready DR, Pritchard KI, Gelmon K, Murray Y, Chapman JA, Chen BE, Levine MN, Investigators MAS (2015) Regional nodal irradiation in early-stage breast cancer. N Engl J Med 373:307–316. https://doi.org/10.1056/NEJMoa1415340
Chudakov DM, Lukyanov S, Lukyanov KA (2007) Tracking intracellular protein movements using photoswitchable fluorescent proteins PS-CFP2 and Dendra2. Nat Protoc 2:2024–2032. https://doi.org/10.1038/nprot.2007.291
Pereira ER, Kedrin D, Seano G, Gautier O, Meijer EFJ, Jones D, Chin SM, Kitahara S, Bouta EM, Chang J, Beech E, Jeong HS, Carroll MC, Taghian AG, Padera TP (2018) Lymph node metastases can invade local blood vessels, exit the node, and colonize distant organs in mice. Science 359:1403–1407. https://doi.org/10.1126/science.aal3622
Hirakawa S, Detmar M, Kerjaschki D, Nagamatsu S, Matsuo K, Tanemura A, Kamata N, Higashikawa K, Okazaki H, Kameda K, Nishida-Fukuda H, Mori H, Hanakawa Y, Sayama K, Shirakata Y, Tohyama M, Tokumaru S, Katayama I, Hashimoto K (2009) Nodal lymphangiogenesis and metastasis: role of tumor-induced lymphatic vessel activation in extramammary Paget’s disease. Am J Pathol 175:2235–2248. https://doi.org/10.2353/ajpath.2009.090420
Nathanson SD, Kwon D, Kapke A, Alford SH, Chitale D (2009) The role of lymph node metastasis in the systemic dissemination of breast cancer. Ann Surg Oncol 16:3396–3405. https://doi.org/10.1245/s10434-009-0659-2
Brown M, Assen FP, Leithner A, Abe J, Schachner H, Asfour G, Bago-Horvath Z, Stein JV, Uhrin P, Sixt M, Kerjaschki D (2018) Lymph node blood vessels provide exit routes for metastatic tumor cell dissemination in mice. Science 359:1408–1411. https://doi.org/10.1126/science.aal3662
Naxerova K, Reiter JG, Brachtel E, Lennerz JK, van de Wetering M, Rowan A, Cai T, Clevers H, Swanton C, Nowak MA, Elledge SJ, Jain RK (2017) Origins of lymphatic and distant metastases in human colorectal cancer. Science 357:55–60. https://doi.org/10.1126/science.aai8515
David Nathanson S, Leonard-Murali S, Burmeister C, Susick L, Baker P (2020) Clinicopathological evaluation of the potential anatomic pathways of systemic metastasis from primary breast cancer suggests an orderly spread through the regional lymph nodes. Ann Surg Oncol 27:4810–4818. https://doi.org/10.1245/s10434-020-08904-w
Nik-Zainal S, Davies H, Staaf J, Ramakrishna M, Glodzik D, Zou X, Martincorena I, Alexandrov LB, Martin S, Wedge DC, Van Loo P, Ju YS, Smid M, Brinkman AB, Morganella S, Aure MR, Lingjaerde OC, Langerod A, Ringner M, Ahn SM, Boyault S, Brock JE, Broeks A, Butler A, Desmedt C, Dirix L, Dronov S, Fatima A, Foekens JA, Gerstung M, Hooijer GK, Jang SJ, Jones DR, Kim HY, King TA, Krishnamurthy S, Lee HJ, Lee JY, Li Y, McLaren S, Menzies A, Mustonen V, O’Meara S, Pauporte I, Pivot X, Purdie CA, Raine K, Ramakrishnan K, Rodriguez-Gonzalez FG, Romieu G, Sieuwerts AM, Simpson PT, Shepherd R, Stebbings L, Stefansson OA, Teague J, Tommasi S, Treilleux I, Van den Eynden GG, Vermeulen P, Vincent-Salomon A, Yates L, Caldas C, Veer L, Tutt A, Knappskog S, Tan BK, Jonkers J, Borg A, Ueno NT, Sotiriou C, Viari A, Futreal PA, Campbell PJ, Span PN, Van Laere S, Lakhani SR, Eyfjord JE, Thompson AM, Birney E, Stunnenberg HG, van de Vijver MJ, Martens JW, Borresen-Dale AL, Richardson AL, Kong G, Thomas G, Stratton MR (2016) Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature 534:47–54. https://doi.org/10.1038/nature17676
Yates LR, Knappskog S, Wedge D, Farmery JHR, Gonzalez S, Martincorena I, Alexandrov LB, Van Loo P, Haugland HK, Lilleng PK, Gundem G, Gerstung M, Pappaemmanuil E, Gazinska P, Bhosle SG, Jones D, Raine K, Mudie L, Latimer C, Sawyer E, Desmedt C, Sotiriou C, Stratton MR, Sieuwerts AM, Lynch AG, Martens JW, Richardson AL, Tutt A, Lonning PE, Campbell PJ (2017) Genomic evolution of breast cancer metastasis and relapse. Cancer Cell 32(169–184):e167. https://doi.org/10.1016/j.ccell.2017.07.005
Yates LR, Gerstung M, Knappskog S, Desmedt C, Gundem G, Van Loo P, Aas T, Alexandrov LB, Larsimont D, Davies H, Li Y, Ju YS, Ramakrishna M, Haugland HK, Lilleng PK, Nik-Zainal S, McLaren S, Butler A, Martin S, Glodzik D, Menzies A, Raine K, Hinton J, Jones D, Mudie LJ, Jiang B, Vincent D, Greene-Colozzi A, Adnet PY, Fatima A, Maetens M, Ignatiadis M, Stratton MR, Sotiriou C, Richardson AL, Lonning PE, Wedge DC, Campbell PJ (2015) Subclonal diversification of primary breast cancer revealed by multiregion sequencing. Nat Med 21:751–759. https://doi.org/10.1038/nm.3886
Zare H, Wang J, Hu A, Weber K, Smith J, Nickerson D, Song C, Witten D, Blau CA, Noble WS (2014) Inferring clonal composition from multiple sections of a breast cancer. PLoS Comput Biol 10:e1003703. https://doi.org/10.1371/journal.pcbi.1003703
Wang Y, Waters J, Leung ML, Unruh A, Roh W, Shi X, Chen K, Scheet P, Vattathil S, Liang H, Multani A, Zhang H, Zhao R, Michor F, Meric-Bernstam F, Navin NE (2014) Clonal evolution in breast cancer revealed by single nucleus genome sequencing. Nature 512:155–160. https://doi.org/10.1038/nature13600
Swanton C, Burrell RA, Futreal PA (2011) Breast cancer genome heterogeneity: a challenge to personalised medicine? Breast Cancer Res 13:104. https://doi.org/10.1186/bcr2807
Savas P, Teo ZL, Lefevre C, Flensburg C, Caramia F, Alsop K, Mansour M, Francis PA, Thorne HA, Silva MJ, Kanu N, Dietzen M, Rowan A, Kschischo M, Fox S, Bowtell DD, Dawson SJ, Speed TP, Swanton C, Loi S (2016) The subclonal architecture of metastatic breast cancer: results from a prospective community-based rapid autopsy program “CASCADE.” PLoS Med 13:e1002204. https://doi.org/10.1371/journal.pmed.1002204
Desmedt C, Fumagalli D, Pietri E, Zoppoli G, Brown D, Nik-Zainal S, Gundem G, Rothe F, Majjaj S, Garuti A, Carminati E, Loi S, Van Brussel T, Boeckx B, Maetens M, Mudie L, Vincent D, Kheddoumi N, Serra L, Massa I, Ballestrero A, Amadori D, Salgado R, de Wind A, Lambrechts D, Piccart M, Larsimont D, Campbell PJ, Sotiriou C (2015) Uncovering the genomic heterogeneity of multifocal breast cancer. J Pathol 236:457–466. https://doi.org/10.1002/path.4540
Gao R, Davis A, McDonald TO, Sei E, Shi X, Wang Y, Tsai PC, Casasent A, Waters J, Zhang H, Meric-Bernstam F, Michor F, Navin NE (2016) Punctuated copy number evolution and clonal stasis in triple-negative breast cancer. Nat Genet 48:1119–1130. https://doi.org/10.1038/ng.3641
Roth A, Khattra J, Yap D, Wan A, Laks E, Biele J, Ha G, Aparicio S, Bouchard-Cote A, Shah SP (2014) PyClone: statistical inference of clonal population structure in cancer. Nat Methods 11:396–398. https://doi.org/10.1038/nmeth.2883
Martincorena I, Roshan A, Gerstung M, Ellis P, Van Loo P, McLaren S, Wedge DC, Fullam A, Alexandrov LB, Tubio JM, Stebbings L, Menzies A, Widaa S, Stratton MR, Jones PH, Campbell PJ (2015) Tumor evolution. High burden and pervasive positive selection of somatic mutations in normal human skin. Science 348:880–886. https://doi.org/10.1126/science.aaa6806
Casasent AK, Schalck A, Gao R, Sei E, Long A, Pangburn W, Casasent T, Meric-Bernstam F, Edgerton ME, Navin NE (2018) Multiclonal invasion in breast tumors identified by topographic single cell sequencing. Cell 172(205–217):e212. https://doi.org/10.1016/j.cell.2017.12.007
Weaver JM, Ross-Innes CS, Shannon N, Lynch AG, Forshew T, Barbera M, Murtaza M, Ong CA, Lao-Sirieix P, Dunning MJ, Smith L, Smith ML, Anderson CL, Carvalho B, O’Donovan M, Underwood TJ, May AP, Grehan N, Hardwick R, Davies J, Oloumi A, Aparicio S, Caldas C, Eldridge MD, Edwards PA, Rosenfeld N, Tavare S, Fitzgerald RC, Consortium O (2014) Ordering of mutations in preinvasive disease stages of esophageal carcinogenesis. Nat Genet 46:837–843. https://doi.org/10.1038/ng.3013
Lee JY, Schizas M, Geyer FC, Selenica P, Piscuoglio S, Sakr RA, Ng CKY, Carniello JVS, Towers R, Giri DD, de Andrade VP, Papanastasiou AD, Viale A, Harris RS, Solit DB, Weigelt B, Reis-Filho JS, King TA (2019) Lobular carcinomas in situ display intralesion genetic heterogeneity and clonal evolution in the progression to invasive lobular carcinoma. Clin Cancer Res 25:674–686. https://doi.org/10.1158/1078-0432.CCR-18-1103
Maino N, Hauling T, Cappi G, Madaboosi N, Dupouy DG, Nilsson M (2019) A microfluidic platform towards automated multiplexed in situ sequencing. Sci Rep 9:3542. https://doi.org/10.1038/s41598-019-40026-6
Vickovic S, Eraslan G, Salmen F, Klughammer J, Stenbeck L, Schapiro D, Aijo T, Bonneau R, Bergenstrahle L, Navarro JF, Gould J, Griffin GK, Borg A, Ronaghi M, Frisen J, Lundeberg J, Regev A, Stahl PL (2019) High-definition spatial transcriptomics for in situ tissue profiling. Nat Methods 16:987–990. https://doi.org/10.1038/s41592-019-0548-y
Rodriques SG, Stickels RR, Goeva A, Martin CA, Murray E, Vanderburg CR, Welch J, Chen LM, Chen F, Macosko EZ (2019) Slide-seq: a scalable technology for measuring genome-wide expression at high spatial resolution. Science 363:1463–1467. https://doi.org/10.1126/science.aaw1219
Nik-Zainal S, Van Loo P, Wedge DC, Alexandrov LB, Greenman CD, Lau KW, Raine K, Jones D, Marshall J, Ramakrishna M, Shlien A, Cooke SL, Hinton J, Menzies A, Stebbings LA, Leroy C, Jia M, Rance R, Mudie LJ, Gamble SJ, Stephens PJ, McLaren S, Tarpey PS, Papaemmanuil E, Davies HR, Varela I, McBride DJ, Bignell GR, Leung K, Butler AP, Teague JW, Martin S, Jonsson G, Mariani O, Boyault S, Miron P, Fatima A, Langerod A, Aparicio SA, Tutt A, Sieuwerts AM, Borg A, Thomas G, Salomon AV, Richardson AL, Borresen-Dale AL, Futreal PA, Stratton MR, Campbell PJ (2012) The life history of 21 breast cancers. Cell 149:994–1007. https://doi.org/10.1016/j.cell.2012.04.023
Klein CA, Blankenstein TJ, Schmidt-Kittler O, Petronio M, Polzer B, Stoecklein NH, Riethmuller G (2002) Genetic heterogeneity of single disseminated tumour cells in minimal residual cancer. Lancet 360:683–689. https://doi.org/10.1016/S0140-6736(02)09838-0
Klein CA (2011) Framework models of tumor dormancy from patient-derived observations. Curr Opin Genet Dev 21:42–49. https://doi.org/10.1016/j.gde.2010.10.011
Klein CA (2009) Parallel progression of primary tumours and metastases. Nat Rev Cancer 9:302–312. https://doi.org/10.1038/nrc2627
Husemann Y, Geigl JB, Schubert F, Musiani P, Meyer M, Burghart E, Forni G, Eils R, Fehm T, Riethmuller G, Klein CA (2008) Systemic spread is an early step in breast cancer. Cancer Cell 13:58–68. https://doi.org/10.1016/j.ccr.2007.12.003
Hosseini H, Obradovic MMS, Hoffmann M, Harper KL, Sosa MS, Werner-Klein M, Nanduri LK, Werno C, Ehrl C, Maneck M, Patwary N, Haunschild G, Guzvic M, Reimelt C, Grauvogl M, Eichner N, Weber F, Hartkopf AD, Taran FA, Brucker SY, Fehm T, Rack B, Buchholz S, Spang R, Meister G, Aguirre-Ghiso JA, Klein CA (2016) Early dissemination seeds metastasis in breast cancer. Nature 540:552–558. https://doi.org/10.1038/nature20785
Demeulemeester J, Kumar P, Moller EK, Nord S, Wedge DC, Peterson A, Mathiesen RR, Fjelldal R, Zamani Esteki M, Theunis K, Fernandez Gallardo E, Grundstad AJ, Borgen E, Baumbusch LO, Borresen-Dale AL, White KP, Kristensen VN, Van Loo P, Voet T, Naume B (2016) Tracing the origin of disseminated tumor cells in breast cancer using single-cell sequencing. Genome Biol 17:250. https://doi.org/10.1186/s13059-016-1109-7
Welch DR, Hurst DR (2019) Defining the hallmarks of metastasis. Cancer Res 79:3011–3027. https://doi.org/10.1158/0008-5472.CAN-19-0458
Kang Y, Siegel PM, Shu W, Drobnjak M, Kakonen SM, Cordon-Cardo C, Guise TA, Massague J (2003) A multigenic program mediating breast cancer metastasis to bone. Cancer Cell 3:537–549. https://doi.org/10.1016/s1535-6108(03)00132-6
Welch DR, Chen P, Miele ME, McGary CT, Bower JM, Stanbridge EJ, Weissman BE (1994) Microcell-mediated transfer of chromosome 6 into metastatic human C8161 melanoma cells suppresses metastasis but does not inhibit tumorigenicity. Oncogene 9:255–262
Phillips KK, Welch DR, Miele ME, Lee JH, Wei LL, Weissman BE (1996) Suppression of MDA-MB-435 breast carcinoma cell metastasis following the introduction of human chromosome 11. Cancer Res 56:1222–1227
Lee JH, Miele ME, Hicks DJ, Phillips KK, Trent JM, Weissman BE, Welch DR (1996) KiSS-1, a novel human malignant melanoma metastasis-suppressor gene. J Natl Cancer Inst 88:1731–1737. https://doi.org/10.1093/jnci/88.23.1731
Lee JH, Welch DR (1997) Identification of highly expressed genes in metastasis-suppressed chromosome 6/human malignant melanoma hybrid cells using subtractive hybridization and differential display. Int J Cancer 71:1035–1044. https://doi.org/10.1002/(sici)1097-0215(19970611)71:6%3c1035::aid-ijc20%3e3.0.co;2-b
Seraj MJ, Samant RS, Verderame MF, Welch DR (2000) Functional evidence for a novel human breast carcinoma metastasis suppressor, BRMS1, encoded at chromosome 11q13. Cancer Res 60:2764–2769
Goldberg SF, Miele ME, Hatta N, Takata M, Paquette-Straub C, Freedman LP, Welch DR (2003) Melanoma metastasis suppression by chromosome 6: evidence for a pathway regulated by CRSP3 and TXNIP. Cancer Res 63:432–440
Weinstein IB, Joe AK (2006) Mechanisms of disease: oncogene addiction–a rationale for molecular targeting in cancer therapy. Nat Clin Pract Oncol 3:448–457. https://doi.org/10.1038/ncponc0558
Esquela-Kerscher A, Slack FJ (2006) Oncomirs - microRNAs with a role in cancer. Nat Rev Cancer 6:259–269. https://doi.org/10.1038/nrc1840
Steeg PS, Theodorescu D (2008) Metastasis: a therapeutic target for cancer. Nat Clin Pract Oncol 5:206–219. https://doi.org/10.1038/ncponc1066
Neumeyer S, Hemani G, Zeggini E (2020) Strengthening causal inference for complex disease using molecular quantitative trait loci. Trends Mol Med 26:232–241. https://doi.org/10.1016/j.molmed.2019.10.004
Abiola O, Angel JM, Avner P, Bachmanov AA, Belknap JK, Bennett B, Blankenhorn EP, Blizard DA, Bolivar V, Brockmann GA, Buck KJ, Bureau JF, Casley WL, Chesler EJ, Cheverud JM, Churchill GA, Cook M, Crabbe JC, Crusio WE, Darvasi A, de Haan G, Dermant P, Doerge RW, Elliot RW, Farber CR, Flaherty L, Flint J, Gershenfeld H, Gibson JP, Gu J, Gu W, Himmelbauer H, Hitzemann R, Hsu HC, Hunter K, Iraqi FF, Jansen RC, Johnson TE, Jones BC, Kempermann G, Lammert F, Lu L, Manly KF, Matthews DB, Medrano JF, Mehrabian M, Mittlemann G, Mock BA, Mogil JS, Montagutelli X, Morahan G, Mountz JD, Nagase H, Nowakowski RS, O’Hara BF, Osadchuk AV, Paigen B, Palmer AA, Peirce JL, Pomp D, Rosemann M, Rosen GD, Schalkwyk LC, Seltzer Z, Settle S, Shimomura K, Shou S, Sikela JM, Siracusa LD, Spearow JL, Teuscher C, Threadgill DW, Toth LA, Toye AA, Vadasz C, Van Zant G, Wakeland E, Williams RW, Zhang HG, Zou F, Complex Trait C (2003) The nature and identification of quantitative trait loci: a community’s view. Nat Rev Genet 4:911–916. https://doi.org/10.1038/nrg1206
Brandon M, Baldi P, Wallace DC (2006) Mitochondrial mutations in cancer. Oncogene 25:4647–4662. https://doi.org/10.1038/sj.onc.1209607
Chandra D, Singh KK (2011) Genetic insights into OXPHOS defect and its role in cancer. Biochim Biophys Acta 1807:620–625. https://doi.org/10.1016/j.bbabio.2010.10.023
Vyas S, Zaganjor E, Haigis MC (2016) Mitochondria and cancer. Cell 166:555–566. https://doi.org/10.1016/j.cell.2016.07.002
Zhang W, Bojorquez-Gomez A, Velez DO, Xu G, Sanchez KS, Shen JP, Chen K, Licon K, Melton C, Olson KM, Yu MK, Huang JK, Carter H, Farley EK, Snyder M, Fraley SI, Kreisberg JF, Ideker T (2018) A global transcriptional network connecting noncoding mutations to changes in tumor gene expression. Nat Genet 50:613–620. https://doi.org/10.1038/s41588-018-0091-2
Cookson W, Liang L, Abecasis G, Moffatt M, Lathrop M (2009) Mapping complex disease traits with global gene expression. Nat Rev Genet 10:184–194. https://doi.org/10.1038/nrg2537
Hunter KW, Amin R, Deasy S, Ha NH, Wakefield L (2018) Genetic insights into the morass of metastatic heterogeneity. Nat Rev Cancer 18:211–223. https://doi.org/10.1038/nrc.2017.126
Scheid AD, Beadnell TC, Welch DR (2019) The second genome: effects of the mitochondrial genome on cancer progression. Adv Cancer Res 142:63–105. https://doi.org/10.1016/bs.acr.2019.01.001
Beadnell TC, Scheid AD, Vivian CJ, Welch DR (2018) Roles of the mitochondrial genetics in cancer metastasis: not to be ignored any longer. Cancer Metastasis Rev 37:615–632. https://doi.org/10.1007/s10555-018-9772-7
Scheid AD, Beadnell TC, Welch DR (2021) Roles of mitochondria in the hallmarks of metastasis. Br J Cancer 124:124–135. https://doi.org/10.1038/s41416-020-01125-8
Lifsted T, Le Voyer T, Williams M, Muller W, Klein-Szanto A, Buetow KH, Hunter KW (1998) Identification of inbred mouse strains harboring genetic modifiers of mammary tumor age of onset and metastatic progression. Int J Cancer 77:640–644. https://doi.org/10.1002/(sici)1097-0215(19980812)77:4%3c640::aid-ijc26%3e3.0.co;2-8
Guy CT, Cardiff RD, Muller WJ (1992) Induction of mammary tumors by expression of polyomavirus middle T oncogene: a transgenic mouse model for metastatic disease. Mol Cell Biol 12:954–961. https://doi.org/10.1128/mcb.12.3.954
Ha NH, Long J, Cai Q, Shu XO, Hunter KW (2016) The circadian rhythm gene Arntl2 is a metastasis susceptibility gene for estrogen receptor-negative breast cancer. PLoS Genet 12:e1006267. https://doi.org/10.1371/journal.pgen.1006267
Faraji F, Hu Y, Wu G, Goldberger NE, Walker RC, Zhang J, Hunter KW (2014) An integrated systems genetics screen reveals the transcriptional structure of inherited predisposition to metastatic disease. Genome Res 24:227–240. https://doi.org/10.1101/gr.166223.113
Hunter KW (2012) Mouse models of cancer: does the strain matter? Nat Rev Cancer 12:144–149. https://doi.org/10.1038/nrc3206
Hsieh SM, Look MP, Sieuwerts AM, Foekens JA, Hunter KW (2009) Distinct inherited metastasis susceptibility exists for different breast cancer subtypes: a prognosis study. Breast Cancer Res 11:R75. https://doi.org/10.1186/bcr2412
Pfefferle AD, Herschkowitz JI, Usary J, Harrell JC, Spike BT, Adams JR, Torres-Arzayus MI, Brown M, Egan SE, Wahl GM, Rosen JM, Perou CM (2013) Transcriptomic classification of genetically engineered mouse models of breast cancer identifies human subtype counterparts. Genome Biol 14:R125. https://doi.org/10.1186/gb-2013-14-11-r125
Herschkowitz JI, Simin K, Weigman VJ, Mikaelian I, Usary J, Hu Z, Rasmussen KE, Jones LP, Assefnia S, Chandrasekharan S, Backlund MG, Yin Y, Khramtsov AI, Bastein R, Quackenbush J, Glazer RI, Brown PH, Green JE, Kopelovich L, Furth PA, Palazzo JP, Olopade OI, Bernard PS, Churchill GA, Van Dyke T, Perou CM (2007) Identification of conserved gene expression features between murine mammary carcinoma models and human breast tumors. Genome Biol 8:R76. https://doi.org/10.1186/gb-2007-8-5-r76
Ishikawa K, Takenaga K, Akimoto M, Koshikawa N, Yamaguchi A, Imanishi H, Nakada K, Honma Y, Hayashi J (2008) ROS-generating mitochondrial DNA mutations can regulate tumor cell metastasis. Science 320:661–664. https://doi.org/10.1126/science.1156906
Bussard KM, Siracusa LD (2017) Understanding mitochondrial polymorphisms in cancer. Cancer Res 77:6051–6059. https://doi.org/10.1158/0008-5472.CAN-17-1939
Mok BY, de Moraes MH, Zeng J, Bosch DE, Kotrys AV, Raguram A, Hsu F, Radey MC, Peterson SB, Mootha VK, Mougous JD, Liu DR (2020) A bacterial cytidine deaminase toxin enables CRISPR-free mitochondrial base editing. Nature 583:631–637. https://doi.org/10.1038/s41586-020-2477-4
Markel P, Shu P, Ebeling C, Carlson GA, Nagle DL, Smutko JS, Moore KJ (1997) Theoretical and empirical issues for marker-assisted breeding of congenic mouse strains. Nat Genet 17:280–284. https://doi.org/10.1038/ng1197-280
Kesterson RA, Johnson LW, Lambert LJ, Vivian JL, Welch DR, Ballinger SW (2016) Generation of mitochondrial-nuclear eXchange mice via pronuclear transfer. Bio Protoc 6:e1976. https://doi.org/10.21769/BioProtoc.1976
Fetterman JL, Zelickson BR, Johnson LW, Moellering DR, Westbrook DG, Pompilius M, Sammy MJ, Johnson M, Dunham-Snary KJ, Cao X, Bradley WE, Zhang J, Wei CC, Chacko B, Schurr TG, Kesterson RA, Dell’italia LJ, Darley-Usmar VM, Welch DR, Ballinger SW (2013) Mitochondrial genetic background modulates bioenergetics and susceptibility to acute cardiac volume overload. Biochem J 455:157–167. https://doi.org/10.1042/BJ20130029
Feeley KP, Bray AW, Westbrook DG, Johnson LW, Kesterson RA, Ballinger SW, Welch DR (2015) Mitochondrial genetics regulate breast cancer tumorigenicity and metastatic potential. Cancer Res 75:4429–4436. https://doi.org/10.1158/0008-5472.CAN-15-0074
Brinker AE, Vivian CJ, Koestler DC, Tsue TT, Jensen RA, Welch DR (2017) Mitochondrial haplotype alters mammary cancer tumorigenicity and metastasis in an oncogenic driver-dependent manner. Cancer Res 77:6941–6949. https://doi.org/10.1158/0008-5472.CAN-17-2194
Vivian CJ, Hagedorn TM, Jensen RA, Brinker AE, Welch DR (2018) Mitochondrial polymorphisms contribute to aging phenotypes in MNX mouse models. Cancer Metastasis Rev 37:633–642. https://doi.org/10.1007/s10555-018-9773-6
Vivian CJ, Brinker AE, Graw S, Koestler DC, Legendre C, Gooden GC, Salhia B, Welch DR (2017) Mitochondrial genomic backgrounds affect nuclear DNA methylation and gene expression. Cancer Res 77:6202–6214. https://doi.org/10.1158/0008-5472.CAN-17-1473
Brinker AE, Vivian CJ, Beadnell TC, Koestler DC, Teoh ST, Lunt SY, Welch DR (2020) Mitochondrial haplotype of the host stromal microenvironment alters metastasis in a non-cell autonomous manner. Cancer Res 80:1118–1129. https://doi.org/10.1158/0008-5472.CAN-19-2481
Belkaid Y, Hand TW (2014) Role of the microbiota in immunity and inflammation. Cell 157:121–141. https://doi.org/10.1016/j.cell.2014.03.011
Beadnell TC, Fain C, Vivian CJ, King JCG, Hastings R, Markiewicz MA, Welch DR (2020) Mitochondrial genetics cooperate with nuclear genetics to selectively alter immune cell development/trafficking. Biochim Biophys Acta Mol Basis Dis 1866:165648. https://doi.org/10.1016/j.bbadis.2019.165648
Gammage PA, Frezza C (2019) Mitochondrial DNA: the overlooked oncogenome? BMC Biol 17:53. https://doi.org/10.1186/s12915-019-0668-y
Wallace DC (2015) Mitochondrial DNA variation in human radiation and disease. Cell 163:33–38. https://doi.org/10.1016/j.cell.2015.08.067
Choudhury AR, Singh KK (2017) Mitochondrial determinants of cancer health disparities. Semin Cancer Biol 47:125–146. https://doi.org/10.1016/j.semcancer.2017.05.001
van der Walt JM, Nicodemus KK, Martin ER, Scott WK, Nance MA, Watts RL, Hubble JP, Haines JL, Koller WC, Lyons K, Pahwa R, Stern MB, Colcher A, Hiner BC, Jankovic J, Ondo WG, Allen FH Jr, Goetz CG, Small GW, Mastaglia F, Stajich JM, McLaurin AC, Middleton LT, Scott BL, Schmechel DE, Pericak-Vance MA, Vance JM (2003) Mitochondrial polymorphisms significantly reduce the risk of Parkinson disease. Am J Hum Genet 72:804–811. https://doi.org/10.1086/373937
Ross DT, Perou CM (2001) A comparison of gene expression signatures from breast tumors and breast tissue derived cell lines. Dis Markers 17:99–109. https://doi.org/10.1155/2001/850531
Carey KD, Garton AJ, Romero MS, Kahler J, Thomson S, Ross S, Park F, Haley JD, Gibson N, Sliwkowski MX (2006) Kinetic analysis of epidermal growth factor receptor somatic mutant proteins shows increased sensitivity to the epidermal growth factor receptor tyrosine kinase inhibitor, erlotinib. Cancer Res 66:8163–8171. https://doi.org/10.1158/0008-5472.CAN-06-0453
Cancer Genome Atlas Network (2012) Comprehensive molecular characterization of human colon and rectal cancer. Nature 487:330–337. https://doi.org/10.1038/nature11252
van Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AA, Mao M, Peterse HL, van der Kooy K, Marton MJ, Witteveen AT, Schreiber GJ, Kerkhoven RM, Roberts C, Linsley PS, Bernards R, Friend SH (2002) Gene expression profiling predicts clinical outcome of breast cancer. Nature 415:530–536. https://doi.org/10.1038/415530a
Slamon DJ, Clark GM, Wong SG, Levin WJ, Ullrich A, McGuire WL (1987) Human breast cancer: correlation of relapse and survival with amplification of the HER-2/neu oncogene. Science 235:177–182. https://doi.org/10.1126/science.3798106
Paik S, Shak S, Tang G, Kim C, Baker J, Cronin M, Baehner FL, Walker MG, Watson D, Park T, Hiller W, Fisher ER, Wickerham DL, Bryant J, Wolmark N (2004) A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med 351:2817–2826. https://doi.org/10.1056/NEJMoa041588
Bertucci F, Ng CKY, Patsouris A, Droin N, Piscuoglio S, Carbuccia N, Soria JC, Dien AT, Adnani Y, Kamal M, Garnier S, Meurice G, Jimenez M, Dogan S, Verret B, Chaffanet M, Bachelot T, Campone M, Lefeuvre C, Bonnefoi H, Dalenc F, Jacquet A, De Filippo MR, Babbar N, Birnbaum D, Filleron T, Le Tourneau C, Andre F (2019) Genomic characterization of metastatic breast cancers. Nature 569:560–564. https://doi.org/10.1038/s41586-019-1056-z
Yuan J, Hu Z, Mahal BA, Zhao SD, Kensler KH, Pi J, Hu X, Zhang Y, Wang Y, Jiang J, Li C, Zhong X, Montone KT, Guan G, Tanyi JL, Fan Y, Xu X, Morgan MA, Long M, Zhang Y, Zhang R, Sood AK, Rebbeck TR, Dang CV, Zhang L (2018) Integrated analysis of genetic ancestry and genomic alterations across cancers. Cancer Cell 34(549–560):e549. https://doi.org/10.1016/j.ccell.2018.08.019
Dietze EC, Sistrunk C, Miranda-Carboni G, O’Regan R, Seewaldt VL (2015) Triple-negative breast cancer in African-American women: disparities versus biology. Nat Rev Cancer 15:248–254. https://doi.org/10.1038/nrc3896
Dietz JR, Partridge AH, Gemignani ML, Javid SH, Kuerer HM (2015) Breast cancer management updates: young and older, pregnant, or male. Ann Surg Oncol 22:3219–3224. https://doi.org/10.1245/s10434-015-4755-1
Carey LA, Perou CM, Livasy CA, Dressler LG, Cowan D, Conway K, Karaca G, Troester MA, Tse CK, Edmiston S, Deming SL, Geradts J, Cheang MC, Nielsen TO, Moorman PG, Earp HS, Millikan RC (2006) Race, breast cancer subtypes, and survival in the Carolina Breast Cancer Study. JAMA 295:2492–2502. https://doi.org/10.1001/jama.295.21.2492
Perou CM, Borresen-Dale AL (2011) Systems biology and genomics of breast cancer. Cold Spring Harb Perspect Biol 3:a003293. https://doi.org/10.1101/cshperspect.a003293
Perou CM (2011) Molecular stratification of triple-negative breast cancers. Oncologist 16:61–70. https://doi.org/10.1634/theoncologist.2011-S1-61
Fejerman L, Hu D, Huntsman S, John EM, Stern MC, Haiman CA, Perez-Stable EJ, Ziv E (2013) Genetic ancestry and risk of mortality among U.S Latinas with breast cancer. Cancer Res 73:7243–7253. https://doi.org/10.1158/0008-5472.CAN-13-2014
Ademuyiwa FO, Edge SB, Erwin DO, Orom H, Ambrosone CB, Underwood W 3rd (2011) Breast cancer racial disparities: unanswered questions. Cancer Res 71:640–644. https://doi.org/10.1158/0008-5472.CAN-10-3021
Fejerman L, Haiman CA, Reich D, Tandon A, Deo RC, John EM, Ingles SA, Ambrosone CB, Bovbjerg DH, Jandorf LH, Davis W, Ciupak G, Whittemore AS, Press MF, Ursin G, Bernstein L, Huntsman S, Henderson BE, Ziv E, Freedman ML (2009) An admixture scan in 1,484 African American women with breast cancer. Cancer Epidemiol Biomarkers Prev 18:3110–3117. https://doi.org/10.1158/1055-9965.EPI-09-0464
Ademuyiwa FO, Tao Y, Luo J, Weilbaecher K, Ma CX (2017) Differences in the mutational landscape of triple-negative breast cancer in African Americans and Caucasians. Breast Cancer Res Treat 161:491–499. https://doi.org/10.1007/s10549-016-4062-y
Macleod KF (2020) Mitophagy and mitochondrial dysfunction in cancer. Annu Rev Cancer Biol 4:41–60. https://doi.org/10.1146/annurev-cancerbio-030419-033405
Arnould T, Michel S, Renard P (2015) Mitochondria retrograde signaling and the UPR mt: where are we in mammals? Int J Mol Sci 16:18224–18251. https://doi.org/10.3390/ijms160818224
Guerra F, Guaragnella N, Arbini AA, Bucci C, Giannattasio S, Moro L (2017) Mitochondrial dysfunction: a novel potential driver of epithelial-to-mesenchymal transition in cancer. Front Oncol 7:295. https://doi.org/10.3389/fonc.2017.00295
Vanharanta S, Massague J (2013) Origins of metastatic traits. Cancer Cell 24:410–421. https://doi.org/10.1016/j.ccr.2013.09.007
Celia-Terrassa T, Kang Y (2016) Distinctive properties of metastasis-initiating cells. Genes Dev 30:892–908. https://doi.org/10.1101/gad.277681.116
Lambert AW, Pattabiraman DR, Weinberg RA (2017) Emerging biological principles of metastasis. Cell 168:670–691. https://doi.org/10.1016/j.cell.2016.11.037
Massague J, Obenauf AC (2016) Metastatic colonization by circulating tumour cells. Nature 529:298–306. https://doi.org/10.1038/nature17038
Cheung KJ, Ewald AJ (2016) A collective route to metastasis: seeding by tumor cell clusters. Science 352:167–169. https://doi.org/10.1126/science.aaf6546
Cheung KJ, Gabrielson E, Werb Z, Ewald AJ (2013) Collective invasion in breast cancer requires a conserved basal epithelial program. Cell 155:1639–1651. https://doi.org/10.1016/j.cell.2013.11.029
Fidler IJ (1973) The relationship of embolic homogeneity, number, size and viability to the incidence of experimental metastasis. Eur J Cancer 9:223–227. https://doi.org/10.1016/s0014-2964(73)80022-2
Liotta LA, Saidel MG, Kleinerman J (1976) The significance of hematogenous tumor cell clumps in the metastatic process. Cancer Res 36:889–894
Watanabe S (1954) The metastasizability of tumor cells. Cancer 7:215–223. https://doi.org/10.1002/1097-0142(195403)7:2%3c215::aid-cncr2820070203%3e3.0.co;2-6
Allen TA, Asad D, Amu E, Hensley MT, Cores J, Vandergriff A, Tang J, Dinh PU, Shen D, Qiao L, Su T, Hu S, Liang H, Shive H, Harrell E, Campbell C, Peng X, Yoder JA, Cheng K (2019) Circulating tumor cells exit circulation while maintaining multicellularity, augmenting metastatic potential. J Cell Sci 132:jcs231563. https://doi.org/10.1242/jcs.231563
Liu X, Taftaf R, Kawaguchi M, Chang YF, Chen W, Entenberg D, Zhang Y, Gerratana L, Huang S, Patel DB, Tsui E, Adorno-Cruz V, Chirieleison SM, Cao Y, Harney AS, Patel S, Patsialou A, Shen Y, Avril S, Gilmore HL, Lathia JD, Abbott DW, Cristofanilli M, Condeelis JS, Liu H (2019) Homophilic CD44 interactions mediate tumor cell aggregation and polyclonal metastasis in patient-derived breast cancer models. Cancer Discov 9:96–113. https://doi.org/10.1158/2159-8290.Cd-18-0065
Lo HC, Xu Z, Kim IS, Pingel B, Aguirre S, Kodali S, Liu J, Zhang W, Muscarella AM, Hein SM, Krupnick AS, Neilson JR, Paust S, Rosen JM, Wang H, Zhang XHF (2020) Resistance to natural killer cell immunosurveillance confers a selective advantage to polyclonal metastasis. Nat Cancer 1:709–722. https://doi.org/10.1038/s43018-020-0068-9
Zajac O, Raingeaud J, Libanje F, Lefebvre C, Sabino D, Martins I, Roy P, Benatar C, Canet-Jourdan C, Azorin P, Polrot M, Gonin P, Benbarche S, Souquere S, Pierron G, Nowak D, Bigot L, Ducreux M, Malka D, Lobry C, Scoazec JY, Eveno C, Pocard M, Perfettini JL, Elias D, Dartigues P, Goere D, Jaulin F (2018) Tumour spheres with inverted polarity drive the formation of peritoneal metastases in patients with hypermethylated colorectal carcinomas. Nat Cell Biol 20:296–306. https://doi.org/10.1038/s41556-017-0027-6
Maddipati R, Stanger BZ (2015) Pancreatic cancer metastases harbor evidence of polyclonality. Cancer Discov 5:1086–1097. https://doi.org/10.1158/2159-8290.Cd-15-0120
Cheung KJ, Padmanaban V, Silvestri V, Schipper K, Cohen JD, Fairchild AN, Gorin MA, Verdone JE, Pienta KJ, Bader JS, Ewald AJ (2016) Polyclonal breast cancer metastases arise from collective dissemination of keratin 14-expressing tumor cell clusters. Proc Natl Acad Sci USA 113:E854-863. https://doi.org/10.1073/pnas.1508541113
Wrenn ED, Yamamoto A, Moore BM, Huang Y, McBirney M, Thomas AJ, Greenwood E, Rabena YF, Rahbar H, Partridge SC, Cheung KJ (2020) Regulation of collective metastasis by nanolumenal signaling. Cell 183:395-410.e319. https://doi.org/10.1016/j.cell.2020.08.045
Aceto N, Bardia A, Miyamoto DT, Donaldson MC, Wittner BS, Spencer JA, Yu M, Pely A, Engstrom A, Zhu H, Brannigan BW, Kapur R, Stott SL, Shioda T, Ramaswamy S, Ting DT, Lin CP, Toner M, Haber DA, Maheswaran S (2014) Circulating tumor cell clusters are oligoclonal precursors of breast cancer metastasis. Cell 158:1110–1122. https://doi.org/10.1016/j.cell.2014.07.013
Gambera S, Abarrategi A, Gonzalez-Camacho F, Morales-Molina A, Roma J, Alfranca A, Garcia-Castro J (2018) Clonal dynamics in osteosarcoma defined by RGB marking. Nat Commun 9:3994. https://doi.org/10.1038/s41467-018-06401-z
Janiszewska M, Tabassum DP, Castano Z, Cristea S, Yamamoto KN, Kingston NL, Murphy KC, Shu S, Harper NW, Del Alcazar CG, Aleckovic M, Ekram MB, Cohen O, Kwak M, Qin Y, Laszewski T, Luoma A, Marusyk A, Wucherpfennig KW, Wagle N, Fan R, Michor F, McAllister SS, Polyak K (2019) Subclonal cooperation drives metastasis by modulating local and systemic immune microenvironments. Nat Cell Biol 21:879–888. https://doi.org/10.1038/s41556-019-0346-x
Echeverria GV, Powell E, Seth S, Ge Z, Carugo A, Bristow C, Peoples M, Robinson F, Qiu H, Shao J, Jeter-Jones SL, Zhang X, Ramamoorthy V, Cai S, Wu W, Draetta G, Moulder SL, Symmans WF, Chang JT, Heffernan TP, Piwnica-Worms H (2018) High-resolution clonal mapping of multi-organ metastasis in triple negative breast cancer. Nat Commun 9:5079. https://doi.org/10.1038/s41467-018-07406-4
McFadden DG, Papagiannakopoulos T, Taylor-Weiner A, Stewart C, Carter SL, Cibulskis K, Bhutkar A, McKenna A, Dooley A, Vernon A, Sougnez C, Malstrom S, Heimann M, Park J, Chen F, Farago AF, Dayton T, Shefler E, Gabriel S, Getz G, Jacks T (2014) Genetic and clonal dissection of murine small cell lung carcinoma progression by genome sequencing. Cell 156:1298–1311. https://doi.org/10.1016/j.cell.2014.02.031
Kim MY, Oskarsson T, Acharyya S, Nguyen DX, Zhang XH, Norton L, Massague J (2009) Tumor self-seeding by circulating cancer cells. Cell 139:1315–1326. https://doi.org/10.1016/j.cell.2009.11.025
Dang HX, Krasnick BA, White BS, Grossman JG, Strand MS, Zhang J, Cabanski CR, Miller CA, Fulton RS, Goedegebuure SP, Fronick CC, Griffith M, Larson DE, Goetz BD, Walker JR, Hawkins WG, Strasberg SM, Linehan DC, Lim KH, Lockhart AC, Mardis ER, Wilson RK, Ley TJ, Maher CA, Fields RC (2020) The clonal evolution of metastatic colorectal cancer. Sci Adv 6:eaay9691. https://doi.org/10.1126/sciadv.aay9691
Leung ML, Davis A, Gao R, Casasent A, Wang Y, Sei E, Vilar E, Maru D, Kopetz S, Navin NE (2017) Single-cell DNA sequencing reveals a late-dissemination model in metastatic colorectal cancer. Genome Res 27:1287–1299. https://doi.org/10.1101/gr.209973.116
Ulintz PJ, Greenson JK, Wu R, Fearon ER, Hardiman KM (2018) Lymph node metastases in colon cancer are polyclonal. Clin Cancer Res 24:2214–2224. https://doi.org/10.1158/1078-0432.Ccr-17-1425
Wei Q, Ye Z, Zhong X, Li L, Wang C, Myers RE, Palazzo JP, Fortuna D, Yan A, Waldman SA, Chen X, Posey JA, Basu-Mallick A, Jiang BH, Hou L, Shu J, Sun Y, Xing J, Li B, Yang H (2017) Multiregion whole-exome sequencing of matched primary and metastatic tumors revealed genomic heterogeneity and suggested polyclonal seeding in colorectal cancer metastasis. Ann Oncol 28:2135–2141. https://doi.org/10.1093/annonc/mdx278
Dong LQ, Shi Y, Ma LJ, Yang LX, Wang XY, Zhang S, Wang ZC, Duan M, Zhang Z, Liu LZ, Zheng BH, Ding ZB, Ke AW, Gao DM, Yuan K, Zhou J, Fan J, Xi R, Gao Q (2018) Spatial and temporal clonal evolution of intrahepatic cholangiocarcinoma. J Hepatol 69:89–98. https://doi.org/10.1016/j.jhep.2018.02.029
Hirotsu Y, Hada M, Amemiya K, Oyama T, Mochizuki H, Omata M (2020) Multi-regional sequencing reveals clonal and polyclonal seeding from primary tumor to metastases in advanced gastric cancer. J Gastroenterol 55:553–564. https://doi.org/10.1007/s00535-019-01659-6
Gundem G, Van Loo P, Kremeyer B, Alexandrov LB, Tubio JMC, Papaemmanuil E, Brewer DS, Kallio HML, Hognas G, Annala M, Kivinummi K, Goody V, Latimer C, O’Meara S, Dawson KJ, Isaacs W, Emmert-Buck MR, Nykter M, Foster C, Kote-Jarai Z, Easton D, Whitaker HC, Neal DE, Cooper CS, Eeles RA, Visakorpi T, Campbell PJ, McDermott U, Wedge DC, Bova GS (2015) The evolutionary history of lethal metastatic prostate cancer. Nature 520:353–357. https://doi.org/10.1038/nature14347
Siegel MB, He X, Hoadley KA, Hoyle A, Pearce JB, Garrett AL, Kumar S, Moylan VJ, Brady CM, Van Swearingen AE, Marron D, Gupta GP, Thorne LB, Kieran N, Livasy C, Mardis ER, Parker JS, Chen M, Anders CK, Carey LA, Perou CM (2018) Integrated RNA and DNA sequencing reveals early drivers of metastatic breast cancer. J Clin Invest 128:1371–1383. https://doi.org/10.1172/JCI96153
Ullah I, Karthik GM, Alkodsi A, Kjallquist U, Stalhammar G, Lovrot J, Martinez NF, Lagergren J, Hautaniemi S, Hartman J, Bergh J (2018) Evolutionary history of metastatic breast cancer reveals minimal seeding from axillary lymph nodes. J Clin Invest 128:1355–1370. https://doi.org/10.1172/JCI96149
Cleary AS, Leonard TL, Gestl SA, Gunther EJ (2014) Tumour cell heterogeneity maintained by cooperating subclones in Wnt-driven mammary cancers. Nature 508:113–117. https://doi.org/10.1038/nature13187
Tabassum DP, Polyak K (2015) Tumorigenesis: it takes a village. Nat Rev Cancer 15:473–483. https://doi.org/10.1038/nrc3971
Au SH, Edd J, Haber DA, Maheswaran S, Stott SL, Toner M (2017) Clusters of circulating tumor cells: a biophysical and technological perspective. Curr Opin Biomed Eng 3:13–19. https://doi.org/10.1016/j.cobme.2017.08.001
Pantel K, Alix-Panabieres C (2019) Liquid biopsy and minimal residual disease - latest advances and implications for cure. Nat Rev Clin Oncol 16:409–424. https://doi.org/10.1038/s41571-019-0187-3
Thiele JA, Bethel K, Králíčková M, Kuhn P (2017) Circulating tumor cells: fluid surrogates of solid tumors. Annu Rev Pathol 12:419–447. https://doi.org/10.1146/annurev-pathol-052016-100256
Yu M, Stott S, Toner M, Maheswaran S, Haber DA (2011) Circulating tumor cells: approaches to isolation and characterization. J Cell Biol 192:373–382. https://doi.org/10.1083/jcb.201010021
Chang MC, Chang YT, Chen JY, Jeng YM, Yang CY, Tien YW, Yang SH, Chen HL, Liang TY, Wang CF, Lee EY, Chang YC, Lee WH (2016) Clinical significance of circulating tumor microemboli as a prognostic marker in patients with pancreatic ductal adenocarcinoma. Clin Chem 62:505–513. https://doi.org/10.1373/clinchem.2015.248260
Hou JM, Krebs MG, Lancashire L, Sloane R, Backen A, Swain RK, Priest LJ, Greystoke A, Zhou C, Morris K, Ward T, Blackhall FH, Dive C (2012) Clinical significance and molecular characteristics of circulating tumor cells and circulating tumor microemboli in patients with small-cell lung cancer. J Clin Oncol 30:525–532. https://doi.org/10.1200/JCO.2010.33.3716
Lee M, Kim EJ, Cho Y, Kim S, Chung HH, Park NH, Song YS (2017) Predictive value of circulating tumor cells (CTCs) captured by microfluidic device in patients with epithelial ovarian cancer. Gynecol Oncol 145:361–365. https://doi.org/10.1016/j.ygyno.2017.02.042
Long E, Ilie M, Bence C, Butori C, Selva E, Lalvee S, Bonnetaud C, Poissonnet G, Lacour JP, Bahadoran P, Brest P, Gilson E, Ballotti R, Hofman V, Hofman P (2016) High expression of TRF2, SOX10, and CD10 in circulating tumor microemboli detected in metastatic melanoma patients. A potential impact for the assessment of disease aggressiveness. Cancer Med 5:1022–1030. https://doi.org/10.1002/cam4.661
Mu Z, Wang C, Ye Z, Austin L, Civan J, Hyslop T, Palazzo JP, Jaslow R, Li B, Myers RE, Jiang J, Xing J, Yang H, Cristofanilli M (2015) Prospective assessment of the prognostic value of circulating tumor cells and their clusters in patients with advanced-stage breast cancer. Breast Cancer Res Treat 154:563–571. https://doi.org/10.1007/s10549-015-3636-4
Paoletti C, Li Y, Muniz MC, Kidwell KM, Aung K, Thomas DG, Brown ME, Abramson VG, Irvin WJ, Lin NU, Liu MC, Nanda R, Nangia JR, Storniolo AM, Traina TA, Vaklavas C, Van Poznak CH, Wolff AC, Forero-Torres A, Hayes DF (2015) Significance of circulating tumor cells in metastatic triple-negative breast cancer patients within a randomized, phase II trial: TBCRC 019. Clin Cancer Res 21:2771–2779. https://doi.org/10.1158/1078-0432.CCR-14-2781
Vona G, Estepa L, Beroud C, Damotte D, Capron F, Nalpas B, Mineur A, Franco D, Lacour B, Pol S, Brechot C, Paterlini-Brechot P (2004) Impact of cytomorphological detection of circulating tumor cells in patients with liver cancer. Hepatology 39:792–797. https://doi.org/10.1002/hep.20091
Wang C, Mu Z, Chervoneva I, Austin L, Ye Z, Rossi G, Palazzo JP, Sun C, Abu-Khalaf M, Myers RE, Zhu Z, Ba Y, Li B, Hou L, Cristofanilli M, Yang H (2017) Longitudinally collected CTCs and CTC-clusters and clinical outcomes of metastatic breast cancer. Breast Cancer Res Treat 161:83–94. https://doi.org/10.1007/s10549-016-4026-2
Zhang D, Zhao L, Zhou P, Ma H, Huang F, Jin M, Dai X, Zheng X, Huang S, Zhang T (2017) Circulating tumor microemboli (CTM) and vimentin+ circulating tumor cells (CTCs) detected by a size-based platform predict worse prognosis in advanced colorectal cancer patients during chemotherapy. Cancer Cell Int 17:6. https://doi.org/10.1186/s12935-016-0373-7
Zheng X, Fan L, Zhou P, Ma H, Huang S, Yu D, Zhao L, Yang S, Liu J, Huang A, Cai C, Dai X, Zhang T (2017) Detection of circulating tumor cells and circulating tumor microemboli in gastric cancer. Transl Oncol 10:431–441. https://doi.org/10.1016/j.tranon.2017.02.007
Au SH, Storey BD, Moore JC, Tang Q, Chen YL, Javaid S, Sarioglu AF, Sullivan R, Madden MW, O’Keefe R, Haber DA, Maheswaran S, Langenau DM, Stott SL, Toner M (2016) Clusters of circulating tumor cells traverse capillary-sized vessels. Proc Natl Acad Sci USA 113:4947–4952. https://doi.org/10.1073/pnas.1524448113
Piskounova E, Agathocleous M, Murphy MM, Hu Z, Huddlestun SE, Zhao Z, Leitch AM, Johnson TM, DeBerardinis RJ, Morrison SJ (2015) Oxidative stress inhibits distant metastasis by human melanoma cells. Nature 527:186–191. https://doi.org/10.1038/nature15726
Tasdogan A, Faubert B, Ramesh V, Ubellacker JM, Shen B, Solmonson A, Murphy MM, Gu Z, Gu W, Martin M, Kasitinon SY, Vandergriff T, Mathews TP, Zhao Z, Schadendorf D, DeBerardinis RJ, Morrison SJ (2020) Metabolic heterogeneity confers differences in melanoma metastatic potential. Nature 577:115–120. https://doi.org/10.1038/s41586-019-1847-2
Padmanaban V, Krol I, Suhail Y, Szczerba BM, Aceto N, Bader JS, Ewald AJ (2019) E-cadherin is required for metastasis in multiple models of breast cancer. Nature 573:439–444. https://doi.org/10.1038/s41586-019-1526-3
Labuschagne CF, Cheung EC, Blagih J, Domart MC, Vousden KH (2019) Cell clustering promotes a metabolic switch that supports metastatic colonization. Cell Metab 30(720–734):e725. https://doi.org/10.1016/j.cmet.2019.07.014
Brown CW, Amante JJ, Mercurio AM (2018) Cell clustering mediated by the adhesion protein PVRL4 is necessary for alpha6beta4 integrin-promoted ferroptosis resistance in matrix-detached cells. J Biol Chem 293:12741–12748. https://doi.org/10.1074/jbc.RA118.003017
Pavlova NN, Pallasch C, Elia AEH, Braun CJ, Westbrook TF, Hemann M, Elledge SJ (2013) A role for PVRL4-driven cell-cell interactions in tumorigenesis. Elife 2:e00358–e00358. https://doi.org/10.7554/eLife.00358
Dixon SJ, Stockwell BR (2019) The hallmarks of ferroptosis. Ann Rev Cancer Biol 3:35–54. https://doi.org/10.1146/annurev-cancerbio-030518-055844
Chiossone L, Dumas PY, Vienne M, Vivier E (2018) Natural killer cells and other innate lymphoid cells in cancer. Nat Rev Immunol 18:671–688. https://doi.org/10.1038/s41577-018-0061-z
Daher M, Rezvani K (2018) Next generation natural killer cells for cancer immunotherapy: the promise of genetic engineering. Curr Opin Immunol 51:146–153. https://doi.org/10.1016/j.coi.2018.03.013
López-Soto A, Gonzalez S, Smyth MJ, Galluzzi L (2017) Control of metastasis by NK cells. Cancer Cell 32:135–154. https://doi.org/10.1016/j.ccell.2017.06.009
Lopez-Soto A, Huergo-Zapico L, Galvan JA, Rodrigo L, de Herreros AG, Astudillo A, Gonzalez S (2013) Epithelial-mesenchymal transition induces an antitumor immune response mediated by NKG2D receptor. J Immunol 190:4408–4419. https://doi.org/10.4049/jimmunol.1202950
Souza-Fonseca-Guimaraes F, Cursons J, Huntington ND (2019) The emergence of natural killer cells as a major target in cancer immunotherapy. Trends Immunol 40:142–158. https://doi.org/10.1016/j.it.2018.12.003
Szczerba BM, Castro-Giner F, Vetter M, Krol I, Gkountela S, Landin J, Scheidmann MC, Donato C, Scherrer R, Singer J, Beisel C, Kurzeder C, Heinzelmann-Schwarz V, Rochlitz C, Weber WP, Beerenwinkel N, Aceto N (2019) Neutrophils escort circulating tumour cells to enable cell cycle progression. Nature 566:553–557. https://doi.org/10.1038/s41586-019-0915-y
Chan IS, Knutsdottir H, Ramakrishnan G, Padmanaban V, Warrier M, Ramirez JC, Dunworth M, Zhang H, Jaffee EM, Bader JS, Ewald AJ (2020) Cancer cells educate natural killer cells to a metastasis-promoting cell state. J Cell Biol 219:e202001134. https://doi.org/10.1083/jcb.202001134
Gkountela S, Castro-Giner F, Szczerba BM, Vetter M, Landin J, Scherrer R, Krol I, Scheidmann MC, Beisel C, Stirnimann CU, Kurzeder C, Heinzelmann-Schwarz V, Rochlitz C, Weber WP, Aceto N (2019) Circulating tumor cell clustering shapes DNA methylation to enable metastasis seeding. Cell 176:98-112.e114. https://doi.org/10.1016/j.cell.2018.11.046
Klezovitch O, Vasioukhin V (2015) Cadherin signaling: keeping cells in touch. F1000Res 4:550. https://doi.org/10.12688/f1000research.6445.1
Kim NG, Koh E, Chen X, Gumbiner BM (2011) E-cadherin mediates contact inhibition of proliferation through Hippo signaling-pathway components. Proc Natl Acad Sci USA 108:11930–11935. https://doi.org/10.1073/pnas.1103345108
Mendonsa AM, Na TY, Gumbiner BM (2018) E-cadherin in contact inhibition and cancer. Oncogene 37:4769–4780. https://doi.org/10.1038/s41388-018-0304-2
Na TY, Schecterson L, Mendonsa AM, Gumbiner BM (2020) The functional activity of E-cadherin controls tumor cell metastasis at multiple steps. Proc Natl Acad Sci USA 117:5931–5937. https://doi.org/10.1073/pnas.1918167117
Lehmann BD, Bauer JA, Chen X, Sanders ME, Chakravarthy AB, Shyr Y, Pietenpol JA (2011) Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J Clin Invest 121:2750–2767. https://doi.org/10.1172/JCI45014
Masuda H, Baggerly KA, Wang Y, Zhang Y, Gonzalez-Angulo AM, Meric-Bernstam F, Valero V, Lehmann BD, Pietenpol JA, Hortobagyi GN, Symmans WF, Ueno NT (2013) Differential response to neoadjuvant chemotherapy among 7 triple-negative breast cancer molecular subtypes. Clin Cancer Res 19:5533–5540. https://doi.org/10.1158/1078-0432.CCR-13-0799
Ring BZ, Hout DR, Morris SW, Lawrence K, Schweitzer BL, Bailey DB, Lehmann BD, Pietenpol JA, Seitz RS (2016) Generation of an algorithm based on minimal gene sets to clinically subtype triple negative breast cancer patients. BMC Cancer 16:143. https://doi.org/10.1186/s12885-016-2198-0
Wang DY, Jiang Z, Ben-David Y, Woodgett JR, Zacksenhaus E (2019) Molecular stratification within triple-negative breast cancer subtypes. Sci Rep 9:19107. https://doi.org/10.1038/s41598-019-55710-w
Steeg PS (2016) Targeting metastasis. Nat Rev Cancer 16:201–218. https://doi.org/10.1038/nrc.2016.25
Choi JW, Kim JK, Yang YJ, Kim P, Yoon KH, Yun SH (2015) Urokinase exerts antimetastatic effects by dissociating clusters of circulating tumor cells. Cancer Res 75:4474–4482. https://doi.org/10.1158/0008-5472.CAN-15-0684
Acknowledgements
The Symposium was funded in part by the Team Angels Organization, Sterling Heights, Michigan and the Nathanson/Rands Chair in Breast Cancer Research.
Funding
The symposium was funded by the Nathanson/Rands Breast Cancer research chair; TP is funded by NIH R01CA214913 and NIH R01HL128168. L.R.Y is funded by a Wellcome Trust Clinical Research Career Development Fellowship ref: 214584/Z/18/Z. DW is funded by Susan G. Komen for the Cure (Grant No. SAC110037) and the National Foundation for Cancer Research. Additional funding support was provided by U.S. Army Medical Research Defense Command, W81XWH-18-1-0450 (TCB); National Cancer Institute P30-CA168524 (DRW) and National Institutes of Health GM103418 (TCB, ADS and DRW). KP and EW are funded by The Department of Defense Breast Cancer Research Program (BCRP; W81XWH-18–1–0098), the NIH/NCI (Grant No. R37CA234488), Susan G. Komen Foundation (Career Catalyst Research Grant), and the Burroughs Wellcome Fund Career Award for Medical Scientists.
Author information
Authors and Affiliations
Contributions
All sections were written by the contributors and edited by the first author.
Corresponding author
Ethics declarations
Conflict of interest
The authors all declare no conflicts of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee guidelines and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Consent to participate
Not applicable. Internal Review Board approval obtained where necessary.
Consent for publication
IRB approval obtained where necessary.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nathanson, S.D., Detmar, M., Padera, T.P. et al. Mechanisms of breast cancer metastasis. Clin Exp Metastasis 39, 117–137 (2022). https://doi.org/10.1007/s10585-021-10090-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10585-021-10090-2