Abstract
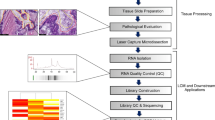
Genome-wide expression profiling has expedited our molecular understanding of the different subtypes of breast cancers, as well as defined the differences among genes expressed in primary tumors and their metastases. Laser-capture microdissection (LCM) coupled to gene expression analysis allows us to understand how specific cell types contribute to the total cancer gene expression signature. Expression profiling was used to define genes that contribute to breast cancer spread into and/or growth within draining lymph nodes (LN). Whole tumor xenografts and their matched whole LN metastases were compared to LCM captured cancer cells from the same tumors and matched LN metastases. One-thousand nine-hundred thirty genes were identified by the whole organ method alone, and 1,281 genes by the LCM method alone. However, less than 1% (30 genes) of genes that changed between tumors and LN metastases were common to both methods. Several of these genes have previously been implicated in cancer aggressiveness. Our data show that whole-organ and LCM based gene expression profiling yield distinctly different lists of metastasis-promoting genes. Contamination of the tumor cells, and cross reactivity of mouse RNA to human-specific chips may explain these differences, and suggests that LCM-derived data may be more accurate.



Similar content being viewed by others
Abbreviations
- CK18:
-
Cytokeratin 18
- ER:
-
Estrogen receptor
- H & E:
-
Hematoxylin and eosin
- LCM:
-
Laser-capture microdissection
- LN:
-
Lymph node
References
van de Rijn M, Perou CM, Tibshirani R et al (2002) Expression of cytokeratins 17 and 5 identifies a group of breast carcinomas with poor clinical outcome. Am J Pathol 161(6):1991–1996
Zajchowski DA, Bartholdi MF, Gong Y et al (2001) Identification of gene expression profiles that predict the aggressive behavior of breast cancer cells. Cancer Res 61(13):5168–5178
Livasy CA, Perou CM, Karaca G et al (2007) Identification of a basal-like subtype of breast ductal carcinoma in situ. Hum Pathol 38(2):197–204
Chang HY, Nuyten DS, Sneddon JB et al (2005) Robustness, scalability, and integration of a wound-response gene expression signature in predicting breast cancer survival. Proc Natl Acad Sci USA 102(10):3738–3743
Chang HY, Sneddon JB, Alizadeh AA et al (2004) Gene expression signature of fibroblast serum response predicts human cancer progression: similarities between tumors and wounds. PLoS Biol 2(2):E7
Rouzier R, Perou CM, Symmans WF et al (2005) Breast cancer molecular subtypes respond differently to preoperative chemotherapy. Clin Cancer Res 11(16):5678–5685
Weigelt B, Hu Z, He X et al (2005) Molecular portraits and 70-gene prognosis signature are preserved throughout the metastatic process of breast cancer. Cancer Res 65(20):9155–9158
Perou CM, Sorlie T, Eisen MB et al (2000) Molecular portraits of human breast tumours. Nature 406(6797):747–752
Shipitsin M, Campbell LL, Argani P et al (2007) Molecular definition of breast tumor heterogeneity. Cancer Cell 11(3):259–273
Fan C, Oh DS, Wessels L et al (2006) Concordance among gene-expression-based predictors for breast cancer. N Engl J Med 355(6):560–569
Harrell JC, Dye WW, Allred DC et al (2006) Estrogen receptor positive breast cancer metastasis: altered hormonal sensitivity and tumor aggressiveness in lymphatic vessels and lymph nodes. Cancer Res 66(18):9308–9315
Sartorius CA, Shen T, Horwitz KB (2003) Progesterone receptors A and B differentially affect the growth of estrogen-dependent human breast tumor xenografts. Breast Cancer Res Treat 79(3):287–299
Weigelt B, Wessels LF, Bosma AJ et al (2005) No common denominator for breast cancer lymph node metastasis. Br J Cancer 93(8):924–932
Montel V, Mose ES, Tarin D (2006) Tumor-stromal interactions reciprocally modulate gene expression patterns during carcinogenesis, metastasis. Int J Cancer 119(2):251–263
Yang F, Foekens JA, Yu J et al (2006) Laser microdissection and microarray analysis of breast tumors reveal ER-alpha related genes and pathways. Oncogene 25(9):1413–1419
Luzzi V, Mahadevappa M, Raja R, Warrington JA, Watson MA (2003) Accurate and reproducible gene expression profiles from laser capture microdissection, transcript amplification, and high density oligonucleotide microarray analysis. J Mol Diagn 5(1):9–14
Naef F, Huelsken J (2005) Cell-type-specific transcriptomics in chimeric models using transcriptome-based masks. Nucleic Acids Res 33(13):e111
Bianco NR, Montano MM (2002) Regulation of prothymosin alpha by estrogen receptor alpha: molecular mechanisms and relevance in estrogen-mediated breast cell growth. Oncogene 21(34):5233–5244
Pan G, Ni J, Wei YF, Yu G, Gentz R, Dixit VM (1997) An antagonist decoy receptor and a death domain-containing receptor for TRAIL. Science 277(5327):815–818
Force WR, Cheung TC, Ware CF (1997) Dominant negative mutants of TRAF3 reveal an important role for the coiled coil domains in cell death signaling by the lymphotoxin-beta receptor. J Biol Chem 272(49):30835–30840
Lin ZP, Belcourt MF, Cory JG, Sartorelli AC (2004) Stable suppression of the R2 subunit of ribonucleotide reductase by R2-targeted short interference RNA sensitizes p53(-/-) HCT-116 colon cancer cells to DNA-damaging agents and ribonucleotide reductase inhibitors. J Biol Chem 279(26):27030–27038
Barrons R (2004) Evaluation of personal digital assistant software for drug interactions. Am J Health Syst Pharm 61(4):380–385
Fiucci G, Ravid D, Reich R, Liscovitch M (2002) Caveolin-1 inhibits anchorage-independent growth, anoikis and invasiveness in MCF-7 human breast cancer cells. Oncogene 21(15):2365–2375
Hasebe T, Konishi M, Iwasaki M et al (2005) Histological characteristics of tumor cells and stromal cells in vessels and lymph nodes are important prognostic parameters of extrahepatic bile duct carcinoma: a prospective study. Hum Pathol 36(6):655–664
Acknowledgements
These studies were supported by a Department of Defense Predoctoral Breast Cancer Training Grant BC050889 to JCH; and by NIH Grant CA26869, the National Foundation for Cancer Research, the Avon Foundation and the Breast Cancer Research Foundation grants to KBH. The authors would like to thank the UCHSC Laser-Capture and Microarray Core Laboratories.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Harrell, J.C., Dye, W.W., Harvell, D.M.E. et al. Contaminating cells alter gene signatures in whole organ versus laser capture microdissected tumors: a comparison of experimental breast cancers and their lymph node metastases. Clin Exp Metastasis 25, 81–88 (2008). https://doi.org/10.1007/s10585-007-9105-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10585-007-9105-7