Abstract
Argonaute-1 (Ago-1) plays a crucial role in gene regulation and genome stability via biogenesis of small non-coding RNAs. Two “Argonaute” family genes, piwi and Ago-2 in Drosophila are involved in multiple silencing mechanisms in the nucleus, transgene cosuppression, long-distant chromosome interaction, nuclear organization and heterochromatin formation. To investigate whether Ago-1 also plays a similar role, we have generated a series of Ago-1 mutations by excising P element, inserted in the Ago-1 promoter (Ago-1 k08121). AGO-1 protein is distributed uniformly in the nucleus and cytosol in early embryos but accumulated predominantly in the cytoplasm during the gastrulation stage. Repeat induced silencing produced by the mini-white (mw) array and transcriptional cosuppression of non-homologous transgenes Adh-w/w-Adh was disrupted by Ago-1 mutation. These effects of Ago-1 are distict from its role in microRNA processing because Dicer-1, a critical enzyme for miRNA biogenesis, has no role on the above silencing. Reduction of AGO-1 protein dislodged the POLYCOMB, EZ (enhancer of zeste) and H3me3K27 binding at the cosuppressed Adh-w transgene insertion sites suggesting its role in Polycomb dependent cosuppression. An overall reduction of methylated histone H3me2K9 and H3me3K27 from the polytene nuclei precisely from the mw promoters was also found that leads to concomitant changes in the chromatin structure. These results suggest a prominent role of Ago-1 in chromatin organization and transgene silencing and demonstrate a critical link between transcriptional transgene cosuppression, heterochromatin formation and chromatin organization. We propose Drosophila Ago-1 as a multifunctional RNAi component that interconnects at least two unrelated events, chromatin organization in the nucleus and microRNA processing in the cytoplasm, which may be extended to the other systems.
Similar content being viewed by others
Introduction
RNA interference (RNAi) and silencing mechanisms control gene expression at both transcriptional and post-transcriptional level (Hanon 2002; Mazke and Birchler 2005). Small regulatory RNA molecules, siRNA and endogenous miRNA, play a central role in RNAi as a guide. During post-transcriptional gene silencing (PTGS) different classes of small regulatory RNA, are loaded onto the RNA induced silencing complex (RISC) containing a conserved Argonaute protein, which binds siRNAs and also directly cleaves target mRNA sequences (Ghildiyal and Zamore 2009). On the other hand, RNAi can also trigger DNA methylation and/or chromatin modifications that lead to transcriptional silencing and heterochromatin formation. These RNA-directed chromatin modifications are best studied in fission yeast Schizosaccharomyces pombe (Bernstein and Allis 2005; Buhler and Moazed 2007).
In fission yeast, siRNAs are derived from centromeric repetitive DNA (CenH) and the formation of heterochromatin at these repeats requires components of the RNAi pathways that initiate H3K9 methylation followed by the recruitment of SWI6, a Drosophila homologue of HP1 (Hall et al. 2002). The Ago-1 containing RNA-induced transcriptional silencing complex is required for establishing heterochromatin assembly at the centromeres (Verdel et al. 2004, Hall et al. 2002). These results suggest that Ago-1 is involved in the establishment of chromatin organization especially via histone tail modifications. It is also plausible that similar function of Ago-1 is conserved in higher eukaryotes, along with its contribution in miRNA biogenesis.
It has been shown that few RNAi factors are involved in repeat induced gene silencing and heterochromatin formation at the centromere and telomere repeats in Drosophila. The modifiers of centromeric silencing often do not influence telomeric silencing (Boivin et al. 2003), because centromeric heterochromatin is functionally distinct from the condensed track of chromatin located elsewhere. piwi, an Ago family gene suppresses centromeric and pericentric silencing but caters an opposite effect in telomeric silencing associated with PIWI bound small RNAs (piRNA) (Yin and Lin 2007). Further, a physical interaction between PIWI and HP1 proposed an alternate pathway for heterochromatin silencing (Brower-Toland et al. 2007) that is different from conventional H3me2K9 dependent heterochromatin assembly (Fanti et al. 1998). The PIWI protein is associated with PIWI-interacting RNAs (piRNAs) strongly and interacts with heterochromatin protein 1a (HP1a) (Brower-Toland et al. 2007). Later it was found that heterochromatin formation is independent of piRNA or endo-siRNA pathways (Moshkovich and Lei 2010). Apart from heterochromatin silencing, piwi and aubergine are also involved in certain aspects of Polycomb dependent transgene cosuppression and PTGS in Drosophila (Pal-Bhadra et al. 2002; Aravin et al. 2004). Both piwi and Ago-1 were also shown to be required for clustering of PRE containing Pc-G targets in the embryonic nuclei as shown by the overlapping Pc-G and PIWI or Pc-G and AGO-1 binding (Grimaud et al. 2006). It is reasonable to anticipate that similar to piwi, Ago-1 might be involved in PcG-mediated transgene cosuppression and PTGS in flies. The present study was carried out to investigate the role of Ago-1 on transgene silencing. Here, we have generated a series of Ago-1 mutations. Using these mutations, we have evaluated the exact role of microRNA processor Ago-1 in chromatin packaging, transcriptional gene silencing and the recruitment of the chromatin bound proteins at the transgene insertion sites.
Materials and methods
Fly stocks and generation of Ago-1 excision lines
Flies were cultured in standard Drosophila food media at 25°C. Majority of the fly markers are noted in FLYBASE (http://flybase.org) unless otherwise described. Mutations of the Argonaute-1 gene, {y 1 w 67c23 ; P{w[+mC] = lacW}Ago-1 k08121 /CyO) were obtained from Bloomington Stock Centre (http://flystocks.bio.indiana.edu, Williams and Rubin 2002). Two independent alleles Ago-1 72 (Ago-1a) and Ago-1 99 (Ago-1b) were generated earlier by the mobilization of single P element on the Ago-1 regulatory region using a transposase stock (Δ2-3Sb/TM3 Ser, Grimaud et al. 2006). Using same genetic schemes, a series of imprecise excision lines were generated that were primarily screened against the loss of adult eye colour produced by the mini-w marker gene of the P element (Fig. S1).
Initially, we have generated 67 independent Ago-1 mutations by excising the P-element of two Ago-1 independent P element insertion alleles (Ago-1 72 and Ago-1 99) (Table S1). As described earlier the P element in Ago-1 72 (Ago-1a) was located to 5 bp upstream to the transcriptional start site (Fig. 1a), whereas P element in Ago-1 99 (Ago-1b) is inserted into 27 bp away from transcriptional start site of the Ago-1 promoter.
Generation of excision lines of Ago-1 and their expression by western blot analysis were generated. a A schematic diagram showing structure of the 5′ of the Ago-1 gene with P element insert (inverted triangle). The sequences of the insertion site of the P element (blank triangle) as reported earlier (top) and the present study (bottom) were marked. b Western blot analysis of AGO-1 protein (Kataoka et al. 2001) extracted from adult female flies carrying Ago-1 excision mutation. Wild-type, heterozygous and heteroallelic Ago-1 mutant combinations were used and β-actin acts as gel loading control. Three independent Western blots were conducted for each genotype. Each blot showed a similar profile, first lane (CS) was taken from a different lane of the same gel and merged
The Ago-1 45 is derived from Ago-1 72 in which excision of the P element has retained one P foot and has deleted the neighboring sequence, including the transcriptional start site (Grimaud et al. 2006). We recovered three more excision lines (e-28, e-37, e-105) from the same cross in which P element retains both feet but eliminate mini-w marker genes. Similarly, three excision lines (e-189, e-196, e-242) derived from Ago-1 99 (Ago-1b) insert were selected. They also retain the feet of the P element but mini-w sequence is deleted. The stocks with these mutations were balanced over CyO chromosome. Females of each excision lines, recovered from excision of P element from two parental stocks (Ago-1a and Ago-1b) were further crossed to the heterozygous males from Ago-1 deficiency stock [Df (2R)50 C-107/CyO]. We did not rescue any viable homozygous escapers. It indicates that Ago-1 sequence is functionally disrupted in those alleles (Table S2). Six of these isolated stocks (e-28, e-37, e-105, e-189, e-196, e-242) were allelic to each other as they failed to complement fully the recessive lethality. Only in rare combinations they generate heteroallelic escapers at low frequencies (Table S3), which are used for further studies.
Genetic crosses
The w-Adh transgenic stock contains 2.5 kb w regulatory sequence fused to 1.9 kb Adh structural gene under Adh fn6 mutant background. The reciprocal Adh-w stock contains a 6.2 kb w structural sequence joined to 1.8 kb Adh promoter fragment in the w minus background (Pal-Bhadra et al. 1997, 2002). Therefore w-Adh and Adh-w transcripts in the transgenic stocks are the sole source of the Adh and w mRNA, respectively.
To examine the effect of Ago-1 mutation on the Adh-w/w-Adh transgene silencing (Pal-Bhadra et al. 1999), two independent Ago-1 excision alleles (Ago-1 e-28 and Ago-1 e-37) were crossed to produce adult heteroallelic escapers with white eyes. One Ago-1 allele (Ago-1 e-28) was combined with a single Adh-w transgene inserted in the X chromosome. Subsequently, another Ago-1 e-37 allele was combined with w-Adh transgene inserted in the chromosome 3. The females Adh-w/Adh-w; Ago-1 e-28 /CyO were further crossed to the Ago-1 e-37 /CyO; w-Adh/w-Adh males. The eye colour of the Adh-w/Y, w-Adh/+ male siblings that are heterozygous (Ago-1 e-28 /+ or Ago-1 e-37 /+) and/or hetero-allelic for the Ago1 mutations (Ago1 e-28 /Ago-1 e-37) were compared. The eye pigment as well as w and Adh transcripts from each genotype were also estimated (Pal-Bhadra et al. 1999).
To study the effect of the Ago-1 mutation on the repeat induced silencing, three mini-w transgenic stocks were used. The mw 6-2 is a transgenic P[lacW] stock, in which single copy of Drosophila mw transgene was inserted at the 50 C2 site of the chromosome 2, which is away from the centric and pericentric heterochromatin (Dorer and Henikoff 1994). BX2 stock contains seven tandem copies of same mw transgene inserted at the same location and DX1 contains seven tandem copies of mw transgenes in which one copy is inverted. The BX2 and DX1 stocks show variegated eye colour phenotype. All these stocks were crossed separately with two Ago-1 excision (Ago-1 e-28, Ago1 e-37) alleles.
To test the effect of Ago-1 mutation on the position effect variegation, the same Ago-1 (Ago-1 e-28, Ago-1 e-37) alleles were combined with y 3p variegated mutation and In(1)w m4h inverted chromosomes separately (Bhadra et al. 1997). In In(1)w m4h stock, a large inversion juxtaposes w gene next to the centromeric heterochromatin.
Pigment assay
For eye pigment assay, 50 heads of adult male flies were dissected manually from each (3–4 days post-eclosion) genotype and homogenized in 0.5 ml of 0.01 M HCl in ethanol. The homogenate was incubated at 4°C overnight and further warmed at 50°C for 5 min. The samples were centrifuged, and the OD of the supernatant was recorded at 480 nm (Ephrussi and Herold 1944).
Fly genomic DNA isolation and PCR
Genomic DNA was extracted from each Ago-1 excision line and parental P strains as described earlier (Huang et al. 2000). To determine the sequence deleted from the ends of the P(lacW) element, the sequence at the junction of the P element and Ago-1 promoter from each excision line was amplified using a P element specific forward primer (forward primer-5′-ACAACCTTTCCTCTCAACAAGC-3′) and Ago-1 gene specific reverse primer (reverse primer-5′-TTTTTGTGCACCAAACACGTTCG-3′) The polymerase chain reaction (PCR) product was sequenced (Applied Biosystems 3730) using gene-specific reverse primer as noted above. The sequences were aligned with P(lacW) and fly genomic sequences carrying wild type Ago-1 gene using NCBI BLAST program.
RT-PCR and Western blot analysis
For semi-quantitative reverse transcription Polymerase chain reaction, 1 μg of clean RNA, isolated by the TRIzol method (Invitrogen, USA) was used for each reaction. Ten picomoles of the gene-specific reverse primer and 18S rRNA reverse primer (internal control) were used for converting RNA into cDNA using superscript reverse transcriptase enzyme. The cDNA was amplified using standard PCR. The PCR products were loaded on 1% agarose gel. The primers used for Ago-1 and 18S rRNA are as follows:
For Ago-1
-
Forward primer (FP): 5′-ATA ATA CCT CGT TCG CAA CT-3′
-
Reverse primer (RP): 5′-TAA TGA CAA CAA GGA TGC AA-3′
and 18S rRNA
-
FP: 5′-CCT TAT GGG ACG TGT GCT TT-3′
-
RP: 5′-CCT GCT GCC TTC CTT AGA TG-3′
For Western blot, nearly 50 adult flies were homogenized in lysis buffer [6% SDS, 1 mM EDTA, 2 mM PMSF, 10 μg/ml aprotinin, 10 μg/ml leupeptin, 10 μg/ml pepstatin] and boiled at 95°C for 5 min. The amount of total proteins of each genotype was measured using Bradford assay (Bradford 1976). Western blot was carried out using anti-rabbit AGO-1 (1:500) antibody (Kataoka et al. 2001). The blots were re-probed with β-actin antibody (1:2,000) as a gel loading control.
The basic histone proteins from wild-type and Ago-1 mutant flies were extracted following acid extraction method (Pal-Bhadra et al. 2007). Western analysis was carried out in two separate sets of blots using anti-rabbit H3me2K9 (1:3000) and anti-rabbit H3me3K27 (1:1500) antibodies. The blots were re-probed with anti-mouse histone H3 (1:2000) antibody that serves as gel loading control.
Northern blot analysis
Total RNA was extracted from adult flies using Trizol. Northern blots were carried out as described earlier (Pal-Bhadra et al. 1997). Fifteen microgras per lane of RNA was used for Northern gel and probed with P32 radioactive labelled w and Adh antisense RNAs. The blots were reprobed with antisense ß-tub RNA as a gel loading control.
Embryo staining
Drosophila embryos were collected in small embryo collection baskets and washed thoroughly with water. The embryos were dechorionized with 50% commercial bleach and fixed with 4% paraformaldehyde. Fixed embryos were transferred to glass scintillation vials containing 1.6 ml of 0.1 M Hepes pH 6.9, 2 mM magnesium sulphate, 1 mM EGTA and 0.4 ml of 20% paraformaldehyde. The vials were gently stirred to maintain an effective emulsion of organic and water phases for 15 to 20 min. The lower phase was separated and 10 ml methanol was added. The embryos were therein transferred to a solution of 90% methanol + 10% 0.5 M EGTA followed by washes in PBT (PBS + 0.1% Tween 20) three times of 5 min each. Thereafter, the embryos were treated with RNase A. For blocking, embryos were incubated in PBT containing 2% goat or animal serum for 30 min followed by overnight incubation with primary rabbit anti-AGO-1 (1:50) (Kataoka et al. 2001) and rabbit anti-Pc (1:100 dilution) (Grimaud et al. 2006) antibodies at 4°C. The embryos were further incubated in FITC conjugated or Cy5 conjugated secondary antibodies (1:50 dilution) for 2 h at room temperature followed by washes in PBS for two times of 5 mins each. The embryos were then mounted in propidium iodide containing VECTASHIELD Mounting Media. The photographs of intact embryos and the Z-sections of embryos were viewed under confocal microscope (Zeiss - 20X and 100X lens [LSM510]).
In situ hybridization
In situ hybridization (FISH) was carried out as described earlier (Pal-Bhadra et al. 1999) to determine the cytological location of Adh-w at 16B and mini-w at the 50 C2 region.
Polytene chromosome preparation
Polytene chromosomes were prepared from the salivary glands of the well-fed third instar Ago-1 mutant and wild-type larvae (Canton S) as described earlier (Pal-Bhadra et al. 2002). The chromosomes were incubated with anti-H3me2K9 (1:25) (Upstate, USA), anti-H3me3K27 (1:25) (Upstate, USA), anti-EZ (1:100) (Grimaud et al. 2006) and anti-Pc (1:50) (Grimaud et al. 2006) antibodies, respectively.
Chromatin immunoprecipitation
Chromatin immunoprecipitation (ChIP) assay was carried out as per protocol described earlier (Cavalli et al. 1999) using H3me2K9 and H3me3K27 antibodies (Upstate, USA) from the immunoprecipitated chromatin of the third instar wild-type and Ago-1 mutant larvae. The primer sets used for amplification of specified regions of the w gene are summarized in Table S4. Relative enrichments were calculated as the ratio of product (w promoter/w second exon) in IP over input. Histograms represent data from three biological replicates analysed in parallel.
Results
Expression of Ago-1 excision alleles
To test whether Ago-1 excision alleles have reduced transcripts and/or protein, semi-quantitative RT-PCR and Western blot analysis was carried out using six excision lines (e-28, e-37, e-105, e-189, e-196, e-242). Total RNA was isolated from each line and quantitative RT-PCR was carried out. Amplified ribosomal (18S rRNA) RNA from the same sample served as an internal control. The relative ratio of Ago-1/18S rRNA from each line showed a dramatic reduction relative to the same ratio (Ago-1/18S rRNA) from the parental w minus flies (Fig. S2). The difference of Ago-1 mRNA level in each line revealed that excision lines show marked reduction in Ago-1 transcript.
Since excision lines are recessive lethal, only adult flies heterozygous for each Ago-1 line and a heteroallelic combination of two parental mutations (Ago-1a, Ago-1b) were further used. Total protein from adult flies of each allele was isolated and Western blot analysis was carried out using AGO-1 antibody (Kataoka et al. 2001), while β-actin served as a gel loading control. Similar to mRNA level, each excision line showed a profound reduction in the AGO-1 protein relative to wild type (Fig. 1b) as described earlier (Okamura et al. 2004).
Distribution of AGO-1 protein in Drosophila embryos
To show AGO-1 protein distribution during development, we immunostained early syncytial blastoderm and mature gastrula embryos with anti-AGO-1 antibody (Kataoka et al. 2001). A moderate amount of AGO-1 protein was accumulated initially in the nucleus and cytoplasm of the blastoderm embryos, nearly 2.5 h after egg laying (AEL) which is above the background level (Fig. 2; stage 5). However, in Ago-1 heterozygous mutant embryos localization of the AGO-1 protein was reduced (Fig. 2; stage 5). We further compared differential distribution patterns of AGO-1 protein in the mature gastrula stages. A marginal increase of staining intensity was noticed at the anterior end of the ventral furrow, the midgut plate and posterior part of the elongated germ band during the gastrulation stage nearly 3–4 h AEL (Fig. 2; stage 9). In gastrula embryos, the protein was mostly localized in the cytoplasm of the cells (Fig. 2; enlarged view, stage 9), which is distinct from the uniform distribution of AGO-1 protein in the nucleus and cytoplasm during the blastoderm stage (stage 5, enlarged view). This clear shift of AGO-1 protein from nucleus to cytoplasm is unique for RNAi factor because majority of the RNAi proteins are localized in the cytoplasm.
Distribution of AGO-1 protein in the wild-type Drosophila embryos. The AGO-1 protein is uniformly distributed during the blastoderm stage (stage 5) significant above the background level (enlarged view in few nuclei), while its accumulation is intense in the cytoplasm during the gastrula stage (stage 9 embryos and enlarged view). Loss of AGO-1 protein in heterozygous Ago-1 combinations (Ago-1a/+) has shown a dramatic loss of AGO-1 protein localization either in embryos or in an enlarge view of the nuclei (stage 5). Scale 50 μm in embryos and scale 10 μm in enlarge nuclei
Role of Ago-1 mutation on transcriptional transgene silencing
Earlier, we have shown that single copy of w-Adh is sufficient to reduce the expression of one copy of Adh-w transgene close to 50% of its normal level (Pal-Bhadra et al. 1999). To examine the role of Ago-1 mutation on the Adh-w/w-Adh transgene cosuppression, the expression of Adh-w transgene was carried out in the w-Adh/Adh-w flies carrying Ago-1 heterozygous or heteroallelic mutations (Ago-1 e-28 /Ago-1 e-37). The Ago-1 excision alleles produced a considerable number of heteroallelic w minus progeny. Therefore, Adh-w transgene is the sole source for red-eye pigment and w transcripts when heteroallelic Ago-1 escapers are combined with Adh-w transgene. The effect of Ago-1 mutation on Adh-w transgenes was detected in the adult eye colour phenotype and by quantitative northern blot analysis using w and Adh radiolabelled probes. Initially, two independent stocks Adh-w/Y; Ago-1 e-28 /CyO; w-Adh/+ and Adh-w/Y; Ago-1 e-37 /In(2LR)Gla; w-Adh/+ were generated by employing multiple crossing schemes (Supplementary Materials). Each Ago-1 allele or alleles in heteroallelic combination did not show any change in the eye colour of the Adh-w males in the absence of the w-Adh construct. However, a mild but consistent increase in the eye colour of Adh-w/w-Adh flies in the heterozygous Ago-1 mutant flies was observed relative to the wild type that was restored close to a normal level in heteroallelic Ago-1 mutant (Ago-1 e-28 /Ago-1 e-37) flies (Fig. 3a, b). Thus, an elevation in the Adh-w/w-Adh eye colour in the presence of Ago-1 mutation indicates that transgene silencing is relieved from the adult eyes, which suggests that Ago-1 mutation represses non-homologous transgene cosuppression (Pal-Bhadra et al. 1999).
Role of Ago-1 in transgene induced silencing. a The eye colour of Adh-w/Y transgenic flies carrying one copy of reciprocal w-Adh constructs that are wild type (+/+) and heteroallelic (Ago-1 e-28 /Ago-1 e-37) for Ago-1 mutation were shown. All flies have w minus background. The transgene copy number was noted in the parenthesis. b The adult eye pigment level of the Adh-w/Y; w-Adh/+ flies having zero, one and two copies of Ago-1 mutation were estimated from three independent sets of experiments. The relative ratios were presented in a bar diagram. The values marked with asterisks are significantly different from +/+ control values (P < 0.05). c, d Autoradiogram showing northern blot hybridization probed with w and β-tub antisense RNA (left) and rehybridized with Adh and β-tub antisense RNAs. The graph depicts the mean values of results obtained from three independent blots. The copy number of each construct or Ago-1 + alleles is noted at the bottom. All flies have w minus and Adh fn6 background. The values marked with asterisks are significantly different from the control values (P < 0.05)
Further, the w transcript levels of the same genotypes were also measured by Northern blot hybridization. A combination of Adh-w/w-Adh transgenes produced 50% of the normal Adh-w mRNA level (Pal-Bhadra et al. 1999, 2002). The reduced w mRNA from the same transgene combination (Adh-w/w-Adh) was also restored close to the normal expression level in the presence of same heteroallelic Ago-1 mutations (Fig. 3c). As expected Ago-1 mutation shows no effect on the Adh-w mRNA alone. Data revealed that similar to eye colour, Ago-1 mutations reduce Adh-w/w-Adh silencing considerably at the transcriptional level.
We further reprobed the same RNA blots with radiolabelled Adh antisense RNA to determine the effect on the w-Adh/Adh silencing (Fig. 3d). As shown earlier, the amount of Adh transcripts was proportionately reduced to the w-Adh transgene dosage when one or two copies of w-Adh (Adh-w/Y; w-Adh/+ or Adh-w/Y; w-Adh/w-Adh) were added to the same genotype (Fig. 3d; Pal-Bhadra et al. 1997). However, a marginal increase of Adh mRNA level was found in Adh-w/Y files in the presence of heteroallelic Ago-1 combinations. It suggests that difference in genetic background in the parental chromosomes 2 carrying each Ago-1 allele might provide these changes in Adh transcripts.
Role of Ago1 on tandem mini-w repeats and position effect variegation
A distinct accumulation of HP1 protein and H3me2K9 methylation at the tandem mini-w array (BX2 and DX1) and their subsequent disruption by the HP1 mutation suggests that mini-w repeats transform into a heterochromatin like structure (Dorer and Henikoff 1994). To analyse whether Ago-1 has any role on heterochromatin formation, variegated expression of the white eye colour in the mini-w repeats (BX2 and DX1) was examined in two heterozygous Ago-1/+ (Ago-1 e-28 /+ and Ago-1 e-37 /+) backgrounds. However, we failed to detect the level of variegation in heteroallelic combination because mini-w arrays and Ago-1 mutations located on the chromosome 2. A two- to three folds increase in the eye pigment was observed in different heterozygous Ago-1 mutant combinations (Fig. 4a) compared to wild type. The amount of eye pigment produced by single mini-w insert in the same location remained unchanged by the same Ago-1 mutant combinations (Fig. 4a). These results suggest that Ago-1 plays a definitive role in repeat induced mini-w silencing.
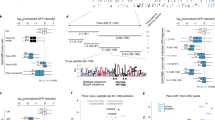
Ago-1 is required for repeat sensitive silencing and position effect variegation (PEV). a The eye colour of the Ago-1 heterozygous males that carry one copy (mini-w 6-2), seven tandem copies (BX2 mini-w) or seven copies with one inverted (DX1 mini-w) white genes were compared. The number of the mini-w copies in the transgenic construct was noted in the parentheses. Mean values (bar) of the eye pigment level from triplicate assays of each genotype were reported by a bar diagram with a standard error. b Ago-1 is a weak suppressor of position effect variegation. The eye colour of the adult males carrying one copy of large X chromosomal inversion [In(1)w m4h] was tested with wild-type or Ago-1 heterozygous mutations. The level of average eye pigment in the adult eyes, displayed by a histogram was marginally high (28–30%) relative to In(1)w m4h flies
Similarly, to analyse the role of Ago-1 on the variegated Drosophila w gene expression or PEV, we combined one copy of Ago-1 mutation with In(1)w m4h chromosome, in which the w gene is juxtaposed close to the heterochromatin. The Ago-1 mutation increases the red pigment in the variegated white eye (w m4h) considerably (28–30% only) compared to the siblings carrying wild-type chromosomes (+/+) (Fig. 4b). This demonstrated that unlike repeat associated silencing, Ago-1 does not show any dramatic change in the pericentric heterochromatic silencing. Thus, the effect of Ago-1 mutations on the two different types of silencing indicates that Ago-1-dependent silencing machinery is involved primarily in heterochromatin like structure formation.
The effect of Ago-1 mutations was further analysed on the variegated allele of the y (y 3p) gene. The y gene (y 3p) displays PEV when placed next to heterochromatin by an inversion In(1)y 3p. In addition to w m4h suppression, Ago-1 also reduced the frequency of variegation of the yellow bristles along the anterior margin of the wing blades. The number of wild type and yellow bristles of ten wing blades was counted from Ago-1 mutant and wild-type flies generated from the same cross. The Ago-1 mutations suppress yellow bristle variegation (22–27%) above the comparable control level (16%) (Table 1). Though the number of yellow bristle varies from wing to wing, but a consistent increase in the variegation of 6–11% greater in the Ago-1 flies over the control was found. A consistent but reasonable inhibition of y and w variegation indicates that Ago–1 is a weak PEV modifier. Therefore, Ago-1 affects heterochromatin formation more intensely at the repeat elements rather than at the centromere.
Drosophila Dicer-1 has no role on w mRNA
Ago-1 is a critical factor for microRNA biogenesis (Diederichs and Haber 2007). Thus, repression of w gene silencing in mini-w repeats and Adh-w transgene in reciprocal constructs (Adh-w/w-Adh) by the Ago-1 mutations may either be operated through miRNA-mediated control or by a miRNA independent pathway. To investigate whether w is an immediate target for any Drosophila microRNAs which are processed by Dicer-1 and AGO-1 proteins, we tried to predict the targeted microRNAs that bind to the 3′ UTR of the w gene using three commonly used miRGen, Targetscan and Pictar predicted tools. Unfortunately, all three predicted tools fail to pick any common miRNA coupled at the 3′ end of Drosophila w gene. It predicts that w is not a direct target for known Drosophila Dicer-1 processed microRNA.
Further, RNAi effector enzymes Dicer-1 (dcr-1) is required for cleavage of pre-microRNA to mature microRNA. Therefore dcr-1 mutation fails to process functional microRNAs. If w gene was controlled by the dcr-1 processed microRNA, an upregulation of w mRNA is expected in the absence of these microRNAs. To test, a loss-of-function dcr-1 mutation (dcr-1Q 1147X) was used (Lee et al. 2004; Grimaud et al. 2006). The mini-w repeats and reciprocal transgenes (Adh-w/w-Adh) were combined with one copy of dcr-1Q 1147X /+ mutation. The presence of dcr-1Q 1147X mutation does not alter eye pigment level of the flies carrying mini-w array or silenced transgene combination (Fig. 5). The amount of eye pigment was assayed in each genotype. An equal expression of eye pigment level in each genotype suggests that dcr-1 has no role on either w gene or Adh-w transgene expression. It shows that the mutational effect of miRNA processor (dcr-1Q 1147X /+) has no influence on w or Adh-w RNA. Therefore, w gene expression might not be regulated directly through Dicer-1-dependent microRNA pathways, though we cannot rule out the possibility of any indirect microRNA influence on the w gene expression.
The effect of dcr-1 mutation on w expression. a The eye colour of the dcr-1Q 1147X /+ heterozygous males that carry one copy (mini-w 6-2), seven tandem copies (BX2 mini-w) or seven copies with one inverted (DX1 mini-w) w genes were compared. Mean values (bar) of the eye pigment level from triplicate assays of each genotype were reported by a bar diagram with a standard error. b The eye colour of the adult males carrying one copy of large X chromosomal inversion [In(1)w m4h] was tested with wild-type or dcr-1Q 1147X /+ heterozygous mutations. The level of average eye pigment in the adult eyes was presented by a histogram
Ago-1 reduces accumulation of H3me2K9 and H3me3K27 methylation
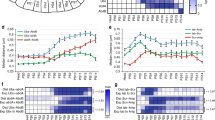
To analyse the contribution of Ago-1 on heterochromatin we have immunostained polytene chromosomes of third instar larvae carrying wild type and heteroallelic Ago-1 mutant alleles using H3me2K9 and H3me3K27 antibodies. In wild-type larvae, an intense accumulation of H3me2K9 and H3me3K27 antibodies was found in the chromocenter, telomere and nearly (40–50) methylation sensitive sites at the chromosomal arms as shown earlier (Pal-Bhadra et al. 2004; Fig. 6a). The binding was consistently disrupted from the majority of the chromosomal sites by the loss of AGO-1 protein. Overall, the reduction of H3me2K9 from the euchromatin sites by the Ago-1 mutation is more pronounced than the loss of H3me3K27 binding (Fig. 6a). At the same time, we also estimated the level of H3me2K9 and H3me3K27 methylation by Western blot analysis. A significant reduction in the H3me2K9 and H3me3K27 levels was found in the Ago-1 heteroallelic mutations (Fig. 6b). These results suggest that the expression and chromosomal distribution of H3me2K9 and H3me3K27 methylation is Ago-1 dependent, thereby indicating that Ago-1 modulates chromosomal organization by largely modifying histone H3 methylation at lysine residues.
Loss of H3me2K9 and H3me3K27 chromosomal binding by the Ago-1 mutations. a Polytene chromosomes were stained with the same above-mentioned antibodies in Canton S and Ago-1 heteroallelic mutant (Ago-1a/Ago-1b). The effect of Ago-1 on H3me2K9 binding on the polytene chromosomes was more intense than the H3me3K27. The H3me2K9 binding was almost eliminated from the euchromatic region but has minimal effect on the intense accumulation at the chromocentre. Scale, 5 μm. b Total histone protein was isolated from the control (Canton S) and heteroallelic Ago-1 mutant (Ago-1a/Ago-1b) flies and Western blot analysis was carried out using H3me2K9 and H3me3K27 antibodies. A substantial decrease in the level of the H3me2K9 and H3me3K27 in the presence of Ago-1 heteroallelic mutations was presented by a histogram depicted from three separate assay
Ago-1 delocalizes histone methylation at the mini-w promoters
We further analysed the binding of histone antibodies on the mini-w insertion sites of BX2 transgenic larvae in a wild-type and/or heteroallelic Ago-1 mutant background. Two histone methylated (H3me2K9 and H3me3K27) antibodies failed to bind at the insertion site of single copy mini-w 6-2 transgene, while a consistent binding of the same antibodies was found at the mini-w arrays of BX2 larvae. However, there was no accumulation of H3me2K9 and H3me3K27 at the same mini-w sites in the Ago-1 heteroallelic larvae indicating a loss of histone methylation at the insertion site of the mini-w array (Fig. 7a). Thus, the Ago-1 is required for proper targeting of histone H3 tail methylation at the Lysine 9 and Lysine 27 residues at the mini-w repeat induced gene silencing.
The Ago-1 mutation diminishes H3me2K9 and H3me3K27 binding in the mini w transgenes. a Merged chromosomal segments showing localization of the H3me2K9 and H3me3K27 antibodies at the 50 C2 site. A strong accumulation was noticed in the seven copy mini-w array (BX2) insertion site in a wild-type background. The binding is abolished by the loss of two copies of Ago-1 alleles at the same site. The grey colour images are pseudo-coloured. Arrows indicate the exact location of mini-w insertion sites as detected by the in situ hybridization. Scale 10 μm. b Chromatin immunoprecipitation was carried out to compare the accumulation of H3me2K9 and H3me3K27 on the w promoter and w second exon of the wild-type and Ago-1a /Ago-1b larvae. The enrichment of proteins from each amplicon of the w locus was measured relative to the w second exon and normalized to wild type. The relative ratios from three independent experiments were depicted as bar diagram. The sequences and location of each primer set of the w gene were summarized in Table S1 (Supplementary Materials)
Next, we performed X Chip analysis using soluble immunoprecipitated chromatins from wild-type and heteroallelic Ago-1 mutant larvae. The immunoprecipitated DNA was amplified from the promoter or second exon of the w gene. The X-Chip assay from the isolated chromatin of the BX2 larvae showed an intense accumulation of H3me2K9 methylation on the promoter site relative to the second exon in wild-type nuclei, whereas accumulation of H3me3K27 at the same promoter site was marginally reduced. However, loss of Ago-1 showed a sharp reduction in the level of H3me2K9 (Fig. 7b) with a nominal reduction in H3me3K27 protein. A similar degree of reduction was also found in relative quantitative RT-PCR assay. These results suggest that Ago-1 is critical for deactivation of mini-w silencing (Fig. 7b).
AGO-1 colocalizes with Pc-G proteins
As noted earlier, non-homologous silencing produced by reciprocal Adh-w and w-Adh transgenes was repressed by the recruitment of the Pc-G proteins in the Adh-w insertion sites (Pal-Bhadra et al. 1999). To reinvestigate, a direct association of Pc bodies and AGO-1 proteins in the embryonic nuclei, we analysed the colocalization of AGO-1 (Kataoka et al. 2001) and Pc-G (Grimaud et al. 2006) proteins by immunostaining (Fig. 8). In wild-type embryos, AGO-1 proteins form uniformly distributed nuclear bodies with 10–15 discrete foci (Fig. S3; stage 5). In contrast, Pc-G proteins, as reported earlier, form a large number of similar nuclear bodies in certain sub-nuclear regions (Fig. 8). In double immunolabelling experiments in the nuclei, 20–25% of the AGO-1 foci were colocalized with Pc-G nuclear bodies, which are apparently 200–500 nm in size. The colocalization between Pc-G and AGO-1 foci suggest that AGO-1 is associated with a subset of Pc-G bodies although Ago-1 is not an integral part of the PRC1 complex.
Colocalization of AGO1 and Polycomb proteins in Drosophila embryos. Embryos (stage 5) were stained with AGO1 (green) and POLYCOMB (Pc) (red) antibodies and propidium iodide (DNA, blue). Images were taken with 100X objective. Projection of merge image of a single nucleus with higher magnification from each panel showed colocalization of AGO-1 and PC proteins. Arrows indicate the overlapping sites (yellow) in the nuclei
Since AGO-1 protein overlaps with Pc-G bodies in embryos, we were interested to examine whether AGO-1 and Pc-G proteins show an overlapping binding on the transgene insertion sites in the larval salivary gland chromosomes. Unfortunately, with repeated attempts, AGO-1 antibody failed to bind on the polytene chromosomes even at a higher concentration. Therefore, only Pc-G bindings at the Adh-w insertion site were tested in different genetic backgrounds. We compared Pc, Ez and H3me3K27 bindings on the Adh-w insertion site (16B) of larvae carrying two copies of Adh-w transgenes showing normal Adh-w expression (Adh-w/Adh-w) or larvae having two copies of the reciprocal Adh-w and w-Adh construct exhibiting cosuppressed state (Adh-w/Adh-w; w-Adh/w-Adh). The Adh-w silencing was strongly correlated with the binding of Pc, Ez and H3me3K27 at the 16B site (Fig. 9). We further analysed the antibody staining of the polytene chromosome from the Ago-1 heteroallelic mutant larvae carrying two copies of each reciprocal transgenes (Adh-w/Adh-w; w-Adh/w-Adh). A significant disruption of binding of Pc, Ez and H3me3K27 on the Adh-w site was noticed indicating repression of transgene cosuppression. However, overall pattern of Pc binding throughout polytene nuclei in wild-type and heteroallelic Ago-1 larvae were identical. This clearly indicates that wild-type AGO-1 protein is required for the proper recruitment of the epigenetic factors that are prerequisite for the Pc-G binding at the silenced insertion sites.
Localization of the PC, Ez and H3K27 (H3me3K27) proteins on the Adh-w insertions sites was determined by the immunostaining of the polytene chromosomes. The location of the Adh-w gene (16B) was determined by in situ hybridization. a The PC binding of the polytene chromosomes in the wild-type and Ago-1a/Ago-1b heteroallelic larvae was determined by immunostaining. Scale 10 μm. b The enlarged view of the chromosomal segment showing recruitment of the PC, Ez and H3me3K27 methylation in the Adh-w sites (16B). The immunostained X chromosomal segment of the 16B region from the Adh-w/Adh-w larvae were compared with homozygous Adh-w larvae carrying two copies of the reciprocal w-Adh transgenes in the wild-type and heteroallelic Ago-1 (Ago-1a/Ago-1b) mutant background. Inclusion of two copies w-Adh transgene recruits the proteins at the same site. In contrast, the presence of the heteroalleleic Ago-1 mutation (Ago-1a/Ago-1b) disrupts the association of all three proteins from the Adh-w insertion site
Discussion
In the present study, Drosophila Ago-1 plays an important role in two types of transcriptional silencing. The Ago-1 mutation impair w-Adh/Adh/Adh-w silencing disrupting Pc-G, Ez and H3meK27 proteins that forms a silencing complex to change the histone tail modification in the localized chromatin structure. On the other hand, loss of H3me2K9 methylation by the same mutation suggests a definitive role in H3me2K9 dependent heterochromatin like structure formation in the mini-w arrays. A similar result has been reported for the piwi mutation (Pal-Bhadra et al. 2002, 2004). In transcriptional silencing, Ago-1 acts as a factor for guiding the Pc-G silencing complex to transgene sites as well as methylation of the lysine residues in the histone tails during heterochromatin assembly. These results suggest that Ago-1 interconnects two types of transcriptional silencing; repeat-induced silencing or heterochromatin formation and transgene cosuppression apart from its role in the microRNA processing.
Two members of the piwi subfamily, aubergine and piwi contribute to transgene silencing, chromatin structure and genome stability. They have been shown to be critical for delocalization of histone lysine 9 methylation and heterochromatin formation (Pal-Bhadra et al. 2004). Here, the loss of AGO-1 protein reduces the chromosomal distribution of H3me2K9 and H3me3K27 methylation. These findings strongly suggest that Ago-1 is an RNAi effector molecule required for chromatin structure, transcriptional repression of the multiple regulatory target sites per se (Grimaud et al. 2006). It probably functions in parallel to piwi action specially by recruiting the silencing complex at the Adh-w sites (Pal-Bhadra et al. 1999). Therefore, members of the same Ago-1 family offer a distinct contribution for rendering TGS at various steps (Meyer et al. 2006).
Depletion of PIWI protein abolishes transgene silencing equally by eliminating Pc-G binding at PRE and non-PRE target sites (Pal-Bhadra et al. 2002; Grimaud et al. 2006). In contrast, Ago-1 only shows a consistent disruption of Pc-G accumulation from the cosuppressed non-PRE transgenes but not from PRE containing elements (Grimaud et al. 2006). The differential effects of Ago-1 in PRE and non-PRE transgenes also suggests that Ago-1 and piwi act in a partially redundant manner to control key recruitment of Pc proteins in the transgene insertion sites and piwi may perform a major function in TGS irrespective of its involvement with a PRE and non-PRE transgenes compared to Ago-1.
Moreover, a connection between Pc-G proteins and endogenous siRNA in TGS has recently been established (Zhao et al. 2008). The Polycomb proteins are targeted by a novel species of noncoding RNA generated from the repetitive sequence of the Xist locus in mammals (Zhao et al. 2008). In Drosophila clustering of PRE targets by the Pc-G proteins on the regulatory Fab-7 element of Abdominal b gene was assisted by the small species of non-coding RNAs (Grimaud et al. 2006). It is reasonable to argue that endogenous siRNAs are processed by Ago-1 but may play a partial instructive role in regulating Polycomb dependent transcriptional silencing and modulation of chromatin structure. Therefore, Ago-1 mutations dislodge Pc-G silencing complex by eliminating specific siRNA.
Argonautes are involved in distinct steps of small RNA maturation and small RNA-mediated gene expression by interacting with diverse protein complexes (Ghildiyal and Zamore 2009). Ago-1 in Arabidopsis serves as an RNAi processor that recruits tasiRNA, rasiRNA processed from repetitive sequences (Agarwal et al. 2007). Recent studies showed that Ago-1 and/or Ago-2 complexes in Drosophila were required for dicing of intermediate siRNA and pre-microRNA based on the selection of their structure (Diederichs and Haber 2007). It was our interest to determine whether Ago-1 might be operating in selecting siRNA and miRNA in the TGS silencing pathways. However, in some cases, Ago-1 is not the only component for cosuppression and heterochromatin to occur, but some specific endogenous siRNA or miRNA-mediated function are required that directly control the immediate components of chromatin bound silencing complex. It is unlikely that miRNA plays any role in transgene silencing though studies establish that miRNA have some specific function for the transcriptional silencing (Kim et al. 2006). We also, however, note that until further study is performed, it remains a formal possibility that mature miRNA might control Polycomb binding and transcriptional silencing in animals as reported in Arabidopsis (Schubert et al. 2005).
Previously, it has been shown that three major components of nuclear RNAi pathway (piwi, spindle E, aubergine) together with heterochromatin-specific proteins are required for the normal organization of the mini-w repeats (Pal-Bhadra et al. 2004). Ago-1 is one of the recent additions in this family. ChIP and immunofluorescence data clearly indicate that H3me2K9 and H3me3K27 levels in the chromatin are reduced in the Ago-1 mutant flies. These findings demonstrate an important role of Ago-1 in establishing the pattern of histone tail modifications that directly regulate nuclear organization by affecting chromatin structure and concomitant gene silencing in condensed track of repetitive sequences.
We also observed that Ago-1 regulates only H3me2K9 and H3me3K27 level significantly. It is reasonable to believe that Ago-1 and other RNAi components may direct H3K9 methylation through siRNA pathways in Drosophila at least in mini-w array. Conversely, the stability of other repeats was reported to be regulated by combining Su(var)3-9 and H3K9 methylation that are independent of the RNAi or siRNA pathways (Peng and Karpen 2007). Therefore, overall distribution of post-translation histone modifications in Drosophila polytene nuclei were also strikingly different from the Su(var)3-9 and Ago-1 mutants. Only H3me2K9 levels were reduced to a considerable level in the Su(var)3-9 mutant nuclei, although visible amounts were retained in the heterochromatin, fourth chromosome and telomeric region (Schotta et al. 2002). In contrast, reduction in the levels of H3me2K9 and H3me3K27 proteins are less in the Ago-1 mutants in the diploid cells but mislocalized from the normal chromosomal position. These findings in Drosophila are surprising as H3me2K9 is found only at a trace amount in the RNAi mutants in Schizosaccharomyces pombe .
The functional diversity of the AGO-1 protein in TGS might shed some light on how Ago-1 functions in small RNA-dependent and RNA-independent transcriptional gene silencing pathways. In conclusion, these studies identify specific functions of AGO-1 to understand their combinatorial role in chromatin modification and heterochromatin formation and its capacity to interact with different species of small regulatory RNAs.
Abbreviations
- Adh:
-
Alcohol dehydrogenase
- AGO:
-
Argonaute
- CS:
-
Canton S
- ChIP:
-
Chromatin-Immunoprecipitation
- CYO:
-
Curly
- FITC:
-
Fluorescein isothiocyanate
- FISH:
-
Flourescent Insitu Hybridisation
- EGTA:
-
Ethylene glycol-bis(2-aminoethylether)-N,N,N′,N′-tetraacetic acid
- H3K9:
-
Histone H3 lysine 9
- HP1:
-
Heterochromatin Protein 1
- miRNA:
-
MicroRNA
- mRNA:
-
messenger RNA
- mw:
-
Miniwhite
- NCBI-BLAST:
-
NCBI-Basic Local Alignment Search Tool
- OD:
-
Optical density
- PBT:
-
PBS +0.1% tween 20
- PBS:
-
Phosphate buffered Saline
- Pc-G:
-
Polycomb group
- PCR:
-
Polymerase Chain Reaction
- P-element:
-
P transposon element
- piRNA:
-
Piwi interacting RNA
- PIWI:
-
P-element induced wimpy testis
- PTGS:
-
Posttranscriptional gene silencing
- PRE:
-
Polycomb response element
- RNAi:
-
RNA interference
- rRNA:
-
Ribosomal RNA
- RT:
-
Reverse Transcription
- SiRNA:
-
Small Interfering RNA
- SWI6:
-
SWItch 6
- TGS:
-
Transcriptional gene silencing
References
Agarwal SK, Gao S, Smith AV, Jin H (2007) A novel class of bacteria-induced small RNAs in Arabidopsis. Genes Dev 21:3123–3134
Aravin AA, Klenov MS, Vagin VV, Bantignies F, Cavalli G, Gvozdev VA (2004) Dissection of a natural RNA silencing process in the Drosophila melanogaster germ line. Mol Cell Biol 24:6742–6750
Bernstein E, Allis CD (2005) RNA meets chromatin. Genes Dev 19:1635–1655
Bhadra U, Pal-Bhadra M, Birchler JA (1997) A sex-influenced modifier in Drosophila that affects a broad spectrum of target loci including the histone repeats. Genetics 146:903–917
Boivin A, Gally C, Netter S, Anxolabéhère D, Ronsseray S (2003) Telomeric associated sequences of Drosophila recruit Polycomb-group proteins in vivo and can induce pairing-sensitive repression. Genetics 164:195–208
Bradford MM (1976) A dye binding assay for protein. Anal Biochem 72:248–254
Brower-Toland B, Findley SD, Jiang L, Liu L, Yin H, Dus M, Zhou P, Elgin SCR, Lin H (2007) Drosophila PIWI associates with chromatin and interacts directly with HP1a. Genes Dev 21:2300–2311
Buhler M, Moazed D (2007) Transcription and RNAi in heterochromatic gene silencing. Nat Struct Mol Biol 14:1041–1048
Cavalli G, Orlando V, Paro R (1999) Mapping DNA target sites of chromatin-associated proteins by formaldehyde cross-linking in Drosophila embryos. In: Chromosome structural analysis: a practical approach. Oxford University Press, UK, pp20–37
Diederichs S, Haber DA (2007) Dual role Argonautes in microRNA processing and post-transcriptional regulation of microRNA expression. Cell 131:1097–1110
Dorer DR, Henikoff S (1994) Expansions of transgene repeats cause heterochromatin formation and gene silencing in Drosophila. Cell 77:993–1002
Ephrussi B, Herold JL (1944) Studies of eye pigments of Drosophila. Methods of extractions and quantitative estimation of the pigment components. Genetics 29:148–175
Fanti L, Dorer DR, Berloco M, Henikoff S, Pimpinelli S (1998) Heterochromatin protein 1 binds transgene arrays. Chromosoma 107:286–292
Ghildiyal M, Zamore PD (2009) Small silencing RNAs: an expanding universe. Nat Rev Genet 10:94–108
Grimaud C, Bantignies F, Pal-Bhadra M, Ghana P, Bhadra U, Cavalli G (2006) RNAi components are required for nuclear clustering of polycomb group response elements. Cell 124:957–971
Hall IM, Shankaranarayana GD, Noma K, Ayoub N, Cohen A, Grewal SI (2002) Establishment and maintenance of a heterochromatin domain. Science 297:2232–2237
Hanon GJ (2002) RNA interference. Nature 418:244–251
Huang AM, Rehm Jay E, Rubin GM (2000) In: Sullivan A et al (eds) Quick preparation of genomic DNA from Drosophila adapted from Drosophila Protocols. CSHL Press, Cold Spring Harbor, NY, USA
Kataoka Y, Takeichi M, Uemura T (2001) Developmental roles and molecular characterization of a Drosophila homologue of Arabidopsis Argonaute1, the founder of a novel gene superfamily. Genes Cells 6:313–325
Kim DH, Villeneuve LM, Morris KV, Rossi JJ (2006) Argonaute-1 directs siRNA-mediated transcriptional gene silencing in human cells. Nat Struct Mol Biol 13:793–797
Lee YS, Nakahara K, Pham JW, Kim K, He Z, Sontheimer EJ, Carthew RW (2004) Distinct roles for Drosophila Dicer-1 and Dicer-2 in the siRNA/miRNA silencing pathways. Cell 117:69–81
Mazke MA, Birchler JA (2005) RNAi-mediated pathways in the nucleus. Nat Rev Genet 6:24–35
Meyer WJ, Schreiber S, Guo Y, Volkmann T, Welte MA, Muller HAJ (2006) Overlapping function of argonaute proteins in patterning and morphogenesis of Drosophila embryos. PLoS Genet 2(8):e134
Moshkovich N, Lei EP (2010) HP1 recruitment in the absence of Argonaute proteins in Drosophila. PLoS Genet 12:e1000880
Okamura K, Ishizika A, Siomi H, Siomi MC (2004) Distinct roles of Argonaute proteins in small RNA-Directed RNA cleavage pathways. Genes Dev 18:1655–1666
Pal-Bhadra M, Bhadra U, Birchler JA (1997) Cosuppression in Drosophila: gene silencing of Alcohol Dehydrogenase by w-Adh transgenes is polycomb dependent. Cell 90:479–490
Pal-Bhadra M, Bhadra U, Birchler JA (1999) Cosuppression in nonhomologous transgenes involves mutually related endogenous sequences. Cell 99:35–46
Pal-Bhadra M, Bhadra U, Birchler JA (2002) RNAi related mechanisms affect both transcriptional and post-transcriptional transgene silencing in Drosophila. Mol Cell 9:315–327
Pal-Bhadra M, Leibovitch BA, Gandhi SG, Rao M, Bhadra U, Birchler JA, Elgin SCR (2004) Heterochromatin silencing and HP1 localization in Drosophila are dependent on the RNAi machinery. Science 303:669–672
Pal-Bhadra M, Bhadra U, Jackson DE, Mamatha L, Park P, Shukla SD (2007) Distinct methylation patterns in histone H3 at Lys-4 and Lys-9 correlate with up- and down-regulation of genes by ethanol in hepatocytes. Life Sci 81:979–987
Peng JC, Karpen GH (2007) H3K9 methylation and RNA interference regulate nucleolar organization and repeated DNA stability. Nat Cell Biol 9:25–35
Schotta G, Ebert A, Krauss V, Fischer A, Hoffmann J, Rea S, Jenuwein T, Dorn R, Reuter G (2002) Central role of Drosophila SU(VAR)3-9 in histone H3-K9 methylation and heterochromatic gene silencing. EMBO J 21:1121–1131
Schubert D, Clarenz O, Goodrich J (2005) Epigenetic control of plant development by polycomb-group proteins. Curr Opinion Plant Biol 8:553–561
Verdel A, Jia S, Gerber S, Sugiyama T, Gygi S, Grewal SI, Moazed D (2004) RNAi-mediated targeting of heterochromatin by the RITS complex. Science 303:672–676
Williams RW, Rubin GM (2002) ARGONAUTE 1 is required for efficient RNA interference in Drosophila embryos PNAS. USA 99:6889–6894
Yin H, Lin H (2007) An epigenetic activation role of piwi and Piwi-associated piRNA in Drosophila melanogaster. Nature 450:304–309
Zhao J, Sun BK, Erwin JA, Song JJ, Lee JT (2008) Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 322:750–756
Acknowledgments
We are grateful to G. Cavalli for providing dcr-1Q 1147X mutant flies and Polycomb and Enhacer of Zeste antibodies and T. Uemura for AGO-1 antibody. We thank J.A. Birchler and lab members of the Bhadra and Pal-Bhadra Group for critical reading of the manuscripts and for the technical help. The work was supported by Senior Wellcome Trust fellowship to UB (GAP0015), MPB (GAP0158) and CSIR Network Project (NPW-34) and DBT Funding (GAP0039) to MPB.
Open Access
This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Dean A. Jackson.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOC 622 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Pushpavalli, S.N.C.V.L., Bag, I., Pal-Bhadra, M. et al. Drosophila Argonaute-1 is critical for transcriptional cosuppression and heterochromatin formation. Chromosome Res 20, 333–351 (2012). https://doi.org/10.1007/s10577-012-9279-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-012-9279-y