Abstract
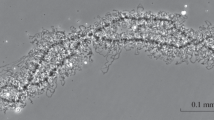
The lampbrush chromosomes present in the nuclei of amphibian oocytes offer unique biological approaches for study of the mechanisms that regulate chromatin structure with high spatial resolution. We discuss fundamental aspects of the remarkable organization and plasticity exhibited by lampbrush chromosomes. We then utilize lampbrush chromosomes to characterize the chromosomal distribution and dynamics of cohesin, the four-protein complex (RAD21/MCD1/SCC1, SMC1, SMC3, SCC3/SA2) responsible for sister chromatid cohesion. We find that endogenous SMC3 and newly expressed hRAD21 co-localize on chromosomal axes, sites where sister chromatids are tightly paired. We present evidence suggesting that hRAD21 recruitment to lampbrush chromosomes is modulated by chromosomal SMC1 and SMC3. Notably, using a technique for de novo chromosome assembly, we demonstrate that both SMC3 and hRAD21 are recruited to single, unreplicated lampbrush chromatids. Finally, we used our novel method of analyzing the oocyte nucleus under oil combined with fluorescence recovery after photobleaching, to provide direct evidence that cohesin is highly dynamic at discrete, condensed chromosomal regions. Collectively, these data demonstrate that lampbrush chromosomes provide a unique and powerful tool for combining biochemical and cytological analyses for dissection of complex chromosomal processes.








Similar content being viewed by others
Abbreviations
- EGFP:
-
enhanced green fluorescent protein
- FRAP:
-
fluorescence recovery after photobleaching
- LBC:
-
lampbrush chromosome
- SMC:
-
structural maintenance of chromosomes
- RNP:
-
ribonucleoprotein
- RNAPII:
-
RNA polymerase II
- YFP:
-
yellow fluorescent protein
References
Alexandru G, Uhlmann F, Mechtler K, Poupart MA, Nasmyth K (2001) Phosphorylation of the cohesin subunit Scc1 by Polo/Cdc5 kinase regulates sister chromatid separation in yeast. Cell 105:459–472
Beenders B, Watrin E, Legagneux V, Kireev I, Bellini M (2003) Distribution of XCAP-E and XCAP-D2 in the Xenopus oocyte nucleus. Chromosome Res 11:549–564
Beenders B, Jones PL, Bellini M (2007) The tripartite motif of nuclear factor 7 is required for its association with transcriptional units. Mol Cell Biol 27:2615–2624
Bellini M (2000) Coilin, more than a molecular marker of the cajal (coiled) body. Bioessays 22:861–867
Brar GA et al (2006) Rec8 phosphorylation and recombination promote the step-wise loss of cohesins in meiosis. Nature 441:532–536
Callan HG (1986) Lampbrush Chromosomes. Springer-Verlag, Berlin, Heidelberg, New York
Chiu A, Revenkova E, Jessberger R (2004) DNA interaction and dimerization of eukaryotic SMC hinge domains. J Biol Chem 279:26233–26242
Ciosk R et al (1998) An ESP1/PDS1 complex regulates loss of sister chromatid cohesion at the metaphase to anaphase transition in yeast. Cell 93:1067–1076
Cohen-Fix O, Peters JM, Kirschner MW, Koshland D (1996) Anaphase initiation in Saccharomyces cerevisiae is controlled by the APC-dependent degradation of the anaphase inhibitor Pds1p. Genes Dev 10:3081–3093
Dorsett D (2007) Roles of the sister chromatid cohesion apparatus in gene expression, development, and human syndromes. Chromosoma 116:1–13
Fischer D, Hock R, Scheer U (1993) DNA topoisomerase II is not detectable on lampbrush chromosomes but enriched in the amplified nucleoli of Xenopus oocytes. Exp Cell Res 209:255–260
Flemming W (1882) Zellsubstanz, Kern und Zelltheilung. F. C. W. Vogel, Leipzig
Funabiki H, Kumada K, Yanagida M (1996) Fission yeast Cut1 and Cut2 are essential for sister chromatid separation, concentrate along the metaphase spindle and form large complexes. EMBO J 15:6617–6628
Gall JG (1956) On the submicroscopic structure of chromosomes. In Mutation 8:17–32
Gall JG, Murphy C (1998) Assembly of lampbrush chromosomes from sperm chromatin. Mol Biol Cell 9:733–747
Gall JG, Bellini M, Wu Z, Murphy C (1999) Assembly of the nuclear transcription and processing machinery: Cajal bodies (coiled bodies) and transcriptosomes. Mol Biol Cell 10:4385–4402
Gerlich D, Koch B, Dupeux F, Peters JM, Ellenberg J (2006) Live-cell imaging reveals a stable cohesin–chromatin interaction after but not before DNA replication. Curr Biol 16:1571–1578
Glynn EF et al (2004) Genome-wide mapping of the cohesin complex in the yeast Saccharomyces cerevisiae. PLoS Biol 2:E259
Gruber S, Haering CH, Nasmyth K (2003) Chromosomal cohesin forms a ring. Cell 112:765–777
Guacci V (2007) Sister chromatid cohesion: the cohesin cleavage model does not ring true. Genes Cells 12:693–708
Guacci V, Hogan E, Koshland D (1994) Chromosome condensation and sister chromatid pairing in budding yeast. J Cell Biol 125:517–530
Guacci V, Koshland D, Strunnikov A (1997) A direct link between sister chromatid cohesion and chromosome condensation revealed through the analysis of MCD1 in S. cerevisiae. Cell 91:47–57
Gurdon JB, De Robertis EM, Partington G (1976) Injected nuclei in frog oocytes provide a living cell system for the study of transcriptional control. Nature 260:116–120
Haering CH et al (2004) Structure and stability of cohesin’s Smc1-kleisin interaction. Mol Cell 15:951–64
Haering CH, Lowe J, Hochwagen A, Nasmyth K (2002) Molecular architecture of SMC proteins and the yeast cohesin complex. Mol Cell 9:773–788
Handwerger KE, Murphy C, Gall JG (2003) Steady-state dynamics of Cajal body components in the Xenopus germinal vesicle. J Cell Biol 160:495–504
Hirano T, Kobayashi R, Hirano M (1997) Condensins, chromosome condensation protein complexes containing XCAP-C, XCAP-E and a Xenopus homolog of the Drosophila Barren protein. Cell 89:511–521
Hock R, Moorman A, Fischer D, Scheer U (1993) Absence of somatic histone H1 in oocytes and preblastula embryos of Xenopus laevis. Dev Biol 158:510–522
Jessberger R (2002) The many functions of SMC proteins in chromosome dynamics. Nat Rev Mol Cell Biol 3:767–778
Jessberger R (2003) SMC proteins at the crossroads of diverse chromosomal processes. IUBMB Life 55:643–652
Kateneva AV, Konovchenko AA, Guacci V, Dresser ME (2005) Recombination protein Tid1p controls resolution of cohesin-dependent linkages in meiosis in Saccharomyces cerevisiae. J Cell Biol 171:241–253
Klein F et al (1999) A central role for cohesins in sister chromatid cohesion, formation of axial elements, and recombination during yeast meiosis. Cell 98:91–103
Krasikova A, Barbero JL, Gaginskaya E (2005) Cohesion proteins are present in centromere protein bodies associated with avian lampbrush chromosomes. Chromosome Res 13:675–685
Krasikova A et al (2006) On the positions of centromeres in chicken lampbrush chromosomes. Chromosome Res 14:777–789
Kumada K et al (1998) Cut1 is loaded onto the spindle by binding to Cut2 and promotes anaphase spindle movement upon Cut2 proteolysis. Curr Biol 8:633–641
Leon P, Kezer J (1990) Loop size in newt lampbrush chromosomes. Chromosoma 99:83–86
Losada A, Hirano M, Hirano T (1998) Identification of Xenopus SMC protein complexes required for sister chromatid cohesion. Genes Dev 12:1986–1997
Losada A, Yokochi T, Kobayashi R, Hirano T (2000) Identification and characterization of SA/Scc3p subunits in the Xenopus and human cohesin complexes. J Cell Biol 150:405–416
Lund E, Paine PL (1990) Nonaqueous isolation of transcriptionally active nuclei from Xenopus oocytes. Methods Enzymol 181:36–43
McKay MJ et al (1996) Sequence conservation of the rad21 Schizosaccharomyces pombe DNA double-strand break repair gene in human and mouse. Genomics 36:305–315
McNairn and Gerton, this issue
Michaelis C, Ciosk R, Nasmyth K (1997) Cohesins: chromosomal proteins that prevent premature separation of sister chromatids. Cell 91:35–45
Morgan GT (2002) Lampbrush chromosomes and associated bodies: new insights into principles of nuclear structure and function. chromosome Res 10:177–200
Murphy C, Wang Z, Roeder RG, Gall JG (2002) RNA polymerase III in Cajal bodies and lampbrush chromosomes of the Xenopus oocyte nucleus. Mol Biol Cell 13:3466–3476
Newmeyer DD, Wilson KL (1991) Egg extracts for nuclear import and nuclear assembly reactions. In: Kay BK, Peng HB (eds) Xenopus laevis: Practical Uses in Cell and Molecular Biology, Vol. 36. Academic Press, San Diego, pp 607–634
Onn I, Heidinger-Pauli JM, Guacci V, Unal E, Koshland DE (2008) Sister chromatid cohesion: a simple concept with a complex reality. Annu Rev Cell Dev Biol
Paine PL, Johnson ME, Lau Y-T, Tluczek LJM, Miller DS (1992) The oocyte nucleus isolated in oil retains in vivo structure and functions. BioTechniques 13:238–245
Parisi S et al (1999) Rec8p, a meiotic recombination and sister chromatid cohesion phosphoprotein of the Rad21p family conserved from fission yeast to humans. Mol Cell Biol 19:3515–3528
Parra MT et al (2004) Involvement of the cohesin Rad21 and SCP3 in monopolar attachment of sister kinetochores during mouse meiosis I. J Cell Sci 117:1221–1234
Patel SB, Novikova N, Bellini M (2007) Splicing-independent recruitment of spliceosomal small nuclear RNPs to nascent RNA polymerase II transcripts. J Cell Biol 178:937–949
Patel S, Novikova N, Beenders B, Austin C, Bellini M (2008) Live images of RNA polymerase II transcription units. Chromosome Res 16:223–232
Prieto I et al (2004) Cohesin component dynamics during meiotic prophase I in mammalian oocytes. Chromosome Res 12:197–213
Revenkova E, Jessberger R (2005) Keeping sister chromatids together: cohesins in meiosis. Reproduction 130:783–790
Rückert J (1892) Zur Entwickelungsgeschichte des Ovarialeies bei Selachiern. Anatomische Anzeiger 7:107–158
Sommerville J, Baird J, Turner BM (1993) Histone H4 acetylation and transcription in amphibian chromatin. J Cell Biol 120:277–290
Stewart MD, Sommerville J, Wong J (2006) Dynamic regulation of histone modifications in Xenopus oocytes through histone exchange. Mol Cell Biol 26:6890–6901
Stoop-Myer C, Amon A (1999) Meiosis: Rec8 is the reason for cohesion. Nat Cell Biol 1:E125–127
Sumara I, Vorlaufer E, Gieffers C, Peters BH, Peters JM (2000) Characterization of vertebrate cohesin complexes and their regulation in prophase. J Cell Biol 151:749–762
Toth A et al (1999) Yeast cohesin complex requires a conserved protein, Eco1p(Ctf7), to establish cohesion between sister chromatids during DNA replication. Genes Dev 13:320–333
Tuma R, Stolk JA, Roth MB (1993) Identification and characterization of a sphere organelle protein. J Cell Biol 122:767–773
Uhlmann F, Lottspeich F, Nasmyth K (1999) Sister-chromatid separation at anaphase onset is promoted by cleavage of the cohesin subunit Scc1. Nature 400:37–42
Uhlmann F, Wernic D, Poupart MA, Koonin EV, Nasmyth K (2000) Cleavage of cohesin by the CD clan protease separin triggers anaphase in yeast. Cell 103:375–386
Waizenegger IC, Hauf S, Meinke A, Peters JM (2000) Two distinct pathways remove mammalian cohesin from chromosome arms in prophase and from centromeres in anaphase. Cell 103:399–410
Wallace RA, Jared DW, Dumont JN, Sega MW (1973) Protein incorporation by isolated amphibian oocytes: III. Optimum incubation conditions. J Exp Zool 184:321–333
Xu H et al (2004) A new role for the mitotic RAD21/SCC1 cohesin in meiotic chromosome cohesion and segregation in the mouse. EMBO Rep 5:378–384
Xu H, Beasley MD, Warren WD, van der Horst GT, McKay MJ (2005) Absence of mouse REC8 cohesin promotes synapsis of sister chromatids in meiosis. Dev Cell 8:949–61
Acknowledgements
We thank Dr. T. Hirano (Cold Spring Harbor Laboratory) for his generous gift of the antibody directed against the amphibian SMC3. We are greatly indebted to Dr. J.G. Gall (Carnegie Institution of Washington) for providing the amazing images of LBC loops where the distribution patterns of histone H4 and RNAPII are compared. We would also like to thank him, as well as Dr. S.B. Patel, for very insightful discussions, and Dr. B. Beenders (UIUC) for the initial anti-cohesin antibody screen. We thank Dr. P.L. Jones (UIUC) for providing Xenopus sperm heads. Finally, we would like to thank our managing editor, Dr. Christian Haering, and the two anonymous reviewers for their very insightful and constructive critique of our manuscript. This work was a supported by a CAREER award from the National Science Foundation. Vincent Guacci is supported by the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Dr. Christian Haering.
Rights and permissions
About this article
Cite this article
Austin, C., Novikova, N., Guacci, V. et al. Lampbrush chromosomes enable study of cohesin dynamics. Chromosome Res 17, 165–184 (2009). https://doi.org/10.1007/s10577-008-9015-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-008-9015-9