Abstract
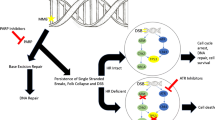
Pancreas ductal adenocarcinoma (PDAC) is the third most common cause of cancer death in the USA. While other cancers with historically poor prognoses have benefited from new immunotherapies and targeted agents, the 5-year survival rate for PDAC patients has remained static. The accessibility to genomic testing has improved in recent years, and it is now clear that PDAC is a heterogenous disease, with a subset of patients harboring actionable mutations. There are several targeted therapies approved by the Food and Drug administration (FDA) in PDAC: EGFR inhibitor erlotinib (combined with gemcitabine) in unselected patients, TRK inhibitors larotrectinib and entrectinib for patients with NTRK fusion mutation, the PD-1 inhibitor pembrolizumab for mismatch repair-deficient patients, and the poly-ADP-ribose polymerase (PARP) inhibitor olaparib in patients with germline BRCA mutation as a maintenance therapy. DNA damage repair (DDR) is paramount to genomic integrity and cell survival. The defective repair of DNA damage is one of the hallmarks of cancer, and abnormalities in DDR pathways are closely linked with the development of malignancies and upregulation of these pathways linked with resistance to treatment. The prevalence of somatic and germline mutations in DDR pathways in metastatic PDAC is reported to be approximately 15–25%. Patients with DDR gene alterations benefit from a personalized approach to treatment. Recently, the POLO trial demonstrated a progression-free survival (PFS) benefit in metastatic PDAC patients with a germline BRCA1/2 mutation treated with maintenance olaparib following platinum-based induction chemotherapy. This was the first phase 3 randomized trial to establish a biomarker-driven approach in the treatment of PDAC and establishes a precedent for maintenance therapy in PDAC. The review herein aims to outline the current treatment landscape for PDAC patients with DDR gene-mutated tumors, highlight novel therapeutic approaches focused on surmounting tumor resistance, and explore new strategies which may lead to an expansion in the number of patients who benefit from these targeted treatments.

Similar content being viewed by others
References
Siegel, R. L., Miller, K. D., Fuchs, H. E., & Jemal, A. (2021). Cancer statistics, 2021. CA: A Cancer Journal for Clinicians, 71(1), 7–33. https://doi.org/10.3322/caac.21654
Conroy, T., Desseigne, F., Ychou, M., Bouché, O., Guimbaud, R., Bécouarn, Y., et al. (2011). FOLFIRINOX versus gemcitabine for metastatic pancreatic cancer. New England Journal of Medicine, 364(19), 1817–1825. https://doi.org/10.1056/NEJMoa1011923
Von Hoff, D. D., Ervin, T., Arena, F. P., Chiorean, E. G., Infante, J., Moore, M., et al. (2013). Increased survival in pancreatic cancer with nab-paclitaxel plus gemcitabine. New England Journal of Medicine, 369(18), 1691–1703. https://doi.org/10.1056/NEJMoa1304369
Prawira, A., Pugh, T. J., Stockley, T. L., & Siu, L. L. (2017). Data resources for the identification and interpretation of actionable mutations by clinicians. Annals of Oncology, 28(5), 946–957. https://doi.org/10.1093/annonc/mdx023
Pishvaian, M. J., Blais, E. M., Brody, J. R., Lyons, E., DeArbeloa, P., Hendifar, A., et al. (2020). Overall survival in patients with pancreatic cancer receiving matched therapies following molecular profiling: A retrospective analysis of the Know Your Tumor registry trial. The Lancet Oncology, 21(4), 508–518. https://doi.org/10.1016/S1470-2045(20)30074-7
Singhi, A. D., George, B., Greenbowe, J. R., Chung, J., Suh, J., Maitra, A., et al. (2019). Real-time targeted genome profile analysis of pancreatic ductal adenocarcinomas identifies genetic alterations that might be targeted with existing drugs or used as biomarkers. Gastroenterology, 156(8), 2242-2253.e2244. https://doi.org/10.1053/j.gastro.2019.02.037
Heestand, G. M., & Kurzrock, R. (2015). Molecular landscape of pancreatic cancer: Implications for current clinical trials. Oncotarget, 6(7), 4553–4561. https://doi.org/10.18632/oncotarget.2972
Lowery, M. A., Wong, W., Jordan, E. J., Lee, J. W., Kemel, Y., Vijai, J., et al. (2018). Prospective evaluation of germline alterations in patients with exocrine pancreatic neoplasms. Journal of the National Cancer Institute, 110(10), 1067–1074. https://doi.org/10.1093/jnci/djy024
Salo-Mullen, E. E., O’Reilly, E. M., Kelsen, D. P., Ashraf, A. M., Lowery, M. A., Yu, K. H., et al. (2015). Identification of germline genetic mutations in patients with pancreatic cancer. Cancer, 121(24), 4382–4388. https://doi.org/10.1002/cncr.29664
Das, S., & Cardin, D. (2020). Targeting DNA damage repair pathways in pancreatic adenocarcinoma. Current Treatment Options in Oncology, 21(8), 62. https://doi.org/10.1007/s11864-020-00763-7
Dreyer, S., Paulus-Hock, V., Upstill-Goddard, R., Lampraki, E., Jamieson, N., Cooke, S., et al. (2019). Defining DNA damage repair deficiency and replication stress in pancreatic cancer. Journal of Clinical Oncology, 37(4_suppl), 285–285. https://doi.org/10.1200/JCO.2019.37.4_suppl.285
Heeke, A. L., Pishvaian, M. J., Lynce, F., Xiu, J., Brody, J. R., Chen, W.-J., et al. (2018). Prevalence of homologous recombination–related gene mutations across multiple cancer types. JCO Precision Oncology(2), 1–13. https://doi.org/10.1200/PO.17.00286
Lindahl, T., & Wood, R. D. (1999). Quality control by DNA repair. Science, 286(5446), 1897–1905. https://doi.org/10.1126/science.286.5446.1897
Curtin, N. J. (2012). DNA repair dysregulation from cancer driver to therapeutic target. Nature Reviews Cancer, 12(12), 801–817. https://doi.org/10.1038/nrc3399
Chae, Y. K., Anker, J. F., Carneiro, B. A., Chandra, S., Kaplan, J., Kalyan, A., et al. (2016). Genomic landscape of DNA repair genes in cancer. Oncotarget, 7(17), 23312–23321. https://doi.org/10.18632/oncotarget.8196
Umar, A., Boland, C. R., Terdiman, J. P., Syngal, S., de la Chapelle, A., Rüschoff, J., et al. (2004). Revised Bethesda Guidelines for hereditary nonpolyposis colorectal cancer (Lynch syndrome) and microsatellite instability. Journal of the National Cancer Institute, 96(4), 261–268. https://doi.org/10.1093/jnci/djh034
Le, D. T., Durham, J. N., Smith, K. N., Wang, H., Bartlett, B. R., Aulakh, L. K., et al. (2017). Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science, 357(6349), 409. https://doi.org/10.1126/science.aan6733
Marabelle, A., Le, D. T., Ascierto, P. A., Di Giacomo, A. M., De Jesus-Acosta, A., Delord, J. P., et al. (2020). Efficacy of pembrolizumab in patients with noncolorectal high microsatellite instability/mismatch repair-deficient cancer: Results from the phase II KEYNOTE-158 study. Journal of Clinical Oncology, 38(1), 1–10. https://doi.org/10.1200/jco.19.02105
Beatty, G. L., Eghbali, S., & Kim, R. (2017). Deploying immunotherapy in pancreatic cancer: Defining mechanisms of response and resistance. American Society of Clinical Oncology Educational Book(37), 267–278. https://doi.org/10.1200/EDBK_175232
Balachandran, V. P., Beatty, G. L., & Dougan, S. K. (2019). Broadening the impact of immunotherapy to pancreatic cancer: Challenges and opportunities. Gastroenterology, 156(7), 2056–2072. https://doi.org/10.1053/j.gastro.2018.12.038
Sarantis, P., Koustas, E., Papadimitropoulou, A., Papavassiliou, A. G., & Karamouzis, M. V. (2020). Pancreatic ductal adenocarcinoma: Treatment hurdles, tumor microenvironment and immunotherapy. World journal of gastrointestinal oncology, 12(2), 173–181. https://doi.org/10.4251/wjgo.v12.i2.173
Sahin, I. H., Askan, G., Hu, Z. I., & O’Reilly, E. M. (2017). Immunotherapy in pancreatic ductal adenocarcinoma: An emerging entity? Annals of Oncology, 28(12), 2950–2961. https://doi.org/10.1093/annonc/mdx503
Yimit, A., Adebali, O., Sancar, A., & Jiang, Y. (2019). Differential damage and repair of DNA-adducts induced by anti-cancer drug cisplatin across mouse organs. Nature Communications, 10(1), 309. https://doi.org/10.1038/s41467-019-08290-2
Plo, I., Liao, Z. Y., Barceló, J. M., Kohlhagen, G., Caldecott, K. W., Weinfeld, M., et al. (2003). Association of XRCC1 and tyrosyl DNA phosphodiesterase (Tdp1) for the repair of topoisomerase I-mediated DNA lesions. DNA Repair (Amst), 2(10), 1087–1100. https://doi.org/10.1016/s1568-7864(03)00116-2
Hoeijmakers, J. H. J. (2001). Genome maintenance mechanisms for preventing cancer. Nature, 411(6835), 366–374. https://doi.org/10.1038/35077232
Sishc, B. J., & Davis, A. J. (2017). The role of the core non-homologous end joining factors in carcinogenesis and cancer. Cancers, 9(7), 81. https://doi.org/10.3390/cancers9070081
Chapman, J. R., Taylor, M. R. G., & Boulton, S. J. (2012). Playing the end game: DNA double-strand break repair pathway choice. Molecular Cell, 47(4), 497–510. https://doi.org/10.1016/j.molcel.2012.07.029
Perkhofer, L., Gout, J., Roger, E., Kude de Almeida, F., Baptista Simões, C., Wiesmüller, L., et al. (2021). DNA damage repair as a target in pancreatic cancer: State-of-the-art and future perspectives. Gut, 70(3), 606–617. https://doi.org/10.1136/gutjnl-2019-319984
Brown, J. S., O’Carrigan, B., Jackson, S. P., & Yap, T. A. (2017). Targeting DNA repair in cancer: Beyond PARP inhibitors. Cancer Discovery, 7(1), 20–37. https://doi.org/10.1158/2159-8290.CD-16-0860
Li, Z., Pearlman, A. H., & Hsieh, P. (2016). DNA mismatch repair and the DNA damage response. DNA Repair, 38, 94–101. https://doi.org/10.1016/j.dnarep.2015.11.019
Cimprich, K. A., & Cortez, D. (2008). ATR: An essential regulator of genome integrity. Nature Reviews Molecular Cell Biology, 9(8), 616–627. https://doi.org/10.1038/nrm2450
Krejci, L., Altmannova, V., Spirek, M., & Zhao, X. (2012). Homologous recombination and its regulation. Nucleic Acids Research, 40(13), 5795–5818. https://doi.org/10.1093/nar/gks270
Dietlein, F., Thelen, L., & Reinhardt, H. C. (2014). Cancer-specific defects in DNA repair pathways as targets for personalized therapeutic approaches. Trends in Genetics, 30(8), 326–339. https://doi.org/10.1016/j.tig.2014.06.003
Knijnenburg, T. A., Wang, L., Zimmermann, M. T., Chambwe, N., Gao, G. F., Cherniack, A. D., et al. (2018). Genomic and molecular landscape of DNA damage repair deficiency across The Cancer Genome Atlas. Cell Reports, 23(1), 239-254.e236. https://doi.org/10.1016/j.celrep.2018.03.076
Shindo, K., Yu, J., Suenaga, M., Fesharakizadeh, S., Cho, C., Macgregor-Das, A., et al. (2017). Deleterious germline mutations in patients with apparently sporadic pancreatic adenocarcinoma. Journal of clinical oncology : Official journal of the American Society of Clinical Oncology, 35(30), 3382–3390. https://doi.org/10.1200/JCO.2017.72.3502
Margaret, A. T. (2019). NCCN guidelines updates: Pancreatic cancer. Journal of the National Comprehensive Cancer Network J Natl Compr Canc Netw, 17(5.5), 603–605. https://doi.org/10.6004/jnccn.2019.5007
Goggins, M., Schutte, M., Lu, J., Moskaluk, C. A., Weinstein, C. L., Petersen, G. M., et al. (1996). Germline BRCA2 gene mutations in patients with apparently sporadic pancreatic carcinomas. Cancer Research, 56(23), 5360–5364.
Roberts, N. J., Norris, A. L., Petersen, G. M., Bondy, M. L., Brand, R., Gallinger, S., et al. (2016). Whole genome sequencing defines the genetic heterogeneity of familial pancreatic cancer. Cancer Discovery, 6(2), 166–175. https://doi.org/10.1158/2159-8290.CD-15-0402
Waddell, N., Pajic, M., Patch, A. M., Chang, D. K., Kassahn, K. S., Bailey, P., et al. (2015). Whole genomes redefine the mutational landscape of pancreatic cancer. Nature, 518(7540), 495–501. https://doi.org/10.1038/nature14169
Knudsen, E. S., O’Reilly, E. M., Brody, J. R., & Witkiewicz, A. K. (2016). Genetic diversity of pancreatic ductal adenocarcinoma and opportunities for precision medicine. Gastroenterology, 150(1), 48–63. https://doi.org/10.1053/j.gastro.2015.08.056
Lucas, A. L., Shakya, R., Lipsyc, M. D., Mitchel, E. B., Kumar, S., Hwang, C., et al. (2013). High prevalence of BRCA1 and BRCA2 germline mutations with loss of heterozygosity in a series of resected pancreatic adenocarcinoma and other neoplastic lesions. Clinical cancer research : An official journal of the American Association for Cancer Research, 19(13), 3396–3403. https://doi.org/10.1158/1078-0432.CCR-12-3020
Krantz, B. A., Yu, K. H., & O’Reilly, E. M. (2017). Pancreas adenocarcinoma: Novel therapeutics. Chinese Clinical Oncology, 6(3), 30. https://doi.org/10.21037/cco.2017.06.14
Lee, K., Yoo, C., Kim, K. P., Park, K. J., Chang, H. M., Kim, T. W., et al. (2018). Germline BRCA mutations in Asian patients with pancreatic adenocarcinoma: A prospective study evaluating risk category for genetic testing. Investigational New Drugs, 36(1), 163–169. https://doi.org/10.1007/s10637-017-0497-1
Yadav, S., Kasi, P. M., Bamlet, W. R., Ho, T. P., Polley, E. C., Hu, C., et al. (2020). Effect of germline mutations in homologous recombination repair genes on overall survival of patients with pancreatic adenocarcinoma. Clinical Cancer Research, 26(24), 6505. https://doi.org/10.1158/1078-0432.CCR-20-1788
Ozçelik, H., Schmocker, B., Di Nicola, N., Shi, X. H., Langer, B., Moore, M., et al. (1997). Germline BRCA2 6174delT mutations in Ashkenazi Jewish pancreatic cancer patients. Nature Genetics, 16(1), 17–18. https://doi.org/10.1038/ng0597-17
Lal, G., Liu, G., Schmocker, B., Kaurah, P., Ozcelik, H., Narod, S. A., et al. (2000). Inherited predisposition to pancreatic adenocarcinoma: Role of family history and germ-line p16, BRCA1, and BRCA2 mutations. Cancer Research, 60(2), 409–416.
Ferrone, C. R., Levine, D. A., Tang, L. H., Allen, P. J., Jarnagin, W., Brennan, M. F., et al. (2009). BRCA germline mutations in Jewish patients with pancreatic adenocarcinoma. Journal of Clinical Oncology, 27(3), 433–438. https://doi.org/10.1200/jco.2008.18.5546
Stadler, Z. K., Salo-Mullen, E., Patil, S. M., Pietanza, M. C., Vijai, J., Saloustros, E., et al. (2012). Prevalence of BRCA1 and BRCA2 mutations in Ashkenazi Jewish families with breast and pancreatic cancer. Cancer, 118(2), 493–499. https://doi.org/10.1002/cncr.26191
Chaffee, K. G., Oberg, A. L., McWilliams, R. R., Majithia, N., Allen, B. A., Kidd, J., et al. (2018). Prevalence of germ-line mutations in cancer genes among pancreatic cancer patients with a positive family history. Genetics in Medicine, 20(1), 119–127. https://doi.org/10.1038/gim.2017.85
Grant, R. C., Selander, I., Connor, A. A., Selvarajah, S., Borgida, A., Briollais, L., et al. (2015). Prevalence of germline mutations in cancer predisposition genes in patients with pancreatic cancer. Gastroenterology, 148(3), 556–564. https://doi.org/10.1053/j.gastro.2014.11.042
Hu, Z. I., Shia, J., Stadler, Z. K., Varghese, A. M., Capanu, M., Salo-Mullen, E., et al. (2018). Evaluating mismatch repair deficiency in pancreatic adenocarcinoma: Challenges and recommendations. Clinical Cancer Research, 24(6), 1326. https://doi.org/10.1158/1078-0432.CCR-17-3099
Golan, T., Kindler, H. L., Park, J. O., Reni, M., Macarulla, T., Hammel, P., et al. (2020). Geographic and ethnic heterogeneity of germline BRCA1 or BRCA2 mutation prevalence among patients with metastatic pancreatic cancer screened for entry into the POLO trial. Journal of Clinical Oncology, 38(13), 1442–1454. https://doi.org/10.1200/JCO.19.01890
Golan, T., & Brody, J. R. (2020). Targeting homologous recombination addicted tumors: Challenges and opportunities. Annals of Pancreatic Cancer, 3, 6.
Lord, C. J., & Ashworth, A. (2016). BRCAness revisited. Nature Reviews Cancer, 16(2), 110–120. https://doi.org/10.1038/nrc.2015.21
Kondo, T., Kanai, M., Kou, T., Sakuma, T., Mochizuki, H., Kamada, M., et al. (2018). Association between homologous recombination repair gene mutations and response to oxaliplatin in pancreatic cancer. Oncotarget, 9(28), 19817–19825. https://doi.org/10.18632/oncotarget.24865
Gulhan, D. C., Lee, J. J., Melloni, G. E. M., Cortés-Ciriano, I., & Park, P. J. (2019). Detecting the mutational signature of homologous recombination deficiency in clinical samples. Nature Genetics, 51(5), 912–919. https://doi.org/10.1038/s41588-019-0390-2
Davies, H., Glodzik, D., Morganella, S., Yates, L. R., Staaf, J., Zou, X., et al. (2017). HRDetect is a predictor of BRCA1 and BRCA2 deficiency based on mutational signatures. Nature Medicine, 23(4), 517–525. https://doi.org/10.1038/nm.4292
Telli, M. L., Timms, K. M., Reid, J., Hennessy, B., Mills, G. B., Jensen, K. C., et al. (2016). Homologous recombination deficiency (HRD) score predicts response to platinum-containing neoadjuvant chemotherapy in patients with triple-negative breast cancer. Clinical Cancer Research, 22(15), 3764–3773. https://doi.org/10.1158/1078-0432.Ccr-15-2477
Takaya, H., Nakai, H., Takamatsu, S., Mandai, M., & Matsumura, N. (2020). Homologous recombination deficiency status-based classification of high-grade serous ovarian carcinoma. Scientific Reports, 10(1), 2757. https://doi.org/10.1038/s41598-020-59671-3
Abkevich, V., Timms, K. M., Hennessy, B. T., Potter, J., Carey, M. S., Meyer, L. A., et al. (2012). Patterns of genomic loss of heterozygosity predict homologous recombination repair defects in epithelial ovarian cancer. British journal of cancer, 107(10), 1776–1782. https://doi.org/10.1038/bjc.2012.451
Popova, T., Manié, E., Rieunier, G., Caux-Moncoutier, V., Tirapo, C., Dubois, T., et al. (2012). Ploidy and large-scale genomic instability consistently identify basal-like breast carcinomas with BRCA1/2 inactivation. Cancer Research, 72(21), 5454. https://doi.org/10.1158/0008-5472.CAN-12-1470
Birkbak, N. J., Wang, Z. C., Kim, J.-Y., Eklund, A. C., Li, Q., Tian, R., et al. (2012). Telomeric allelic imbalance indicates defective DNA repair and sensitivity to DNA-damaging agents. Cancer Discovery, 2(4), 366. https://doi.org/10.1158/2159-8290.CD-11-0206
Golan, T., O’Kane, G. M., Denroche, R. E., Raitses-Gurevich, M., Grant, R. C., Holter, S., et al. (2021). Genomic features and classification of homologous recombination deficient pancreatic ductal adenocarcinoma. Gastroenterology. https://doi.org/10.1053/j.gastro.2021.01.220
O’Kane, G. M., Denroche, R., Picardo, S. L., Zhang, A., Holter, S., Grant, R. C., et al. (2020). Homologous recombination deficiency (HRD) scoring in pancreatic ductal adenocarcinoma (PDAC) and response to chemotherapy. Journal of Clinical Oncology, 38(4_suppl), 741–741. https://doi.org/10.1200/JCO.2020.38.4_suppl.741
Park, W., Chen, J., Chou, J. F., Varghese, A. M., Yu, K. H., Wong, W., et al. (2020). Genomic methods identify homologous recombination deficiency in pancreas adenocarcinoma and optimize treatment selection. Clinical cancer research : An official journal of the American Association for Cancer Research, 26(13), 3239–3247. https://doi.org/10.1158/1078-0432.CCR-20-0418
Golan, T., Hammel, P., Reni, M., Van Cutsem, E., Macarulla, T., Hall, M. J., et al. (2021). Overall survival from the phase 3 POLO trial: Maintenance olaparib for germline BRCA-mutated metastatic pancreatic cancer. Journal of Clinical Oncology, 39(3_suppl), 378–378. https://doi.org/10.1200/JCO.2021.39.3_suppl.378
Yurgelun, M. B., Chittenden, A. B., Morales-Oyarvide, V., Rubinson, D. A., Dunne, R. F., Kozak, M. M., et al. (2019). Germline cancer susceptibility gene variants, somatic second hits, and survival outcomes in patients with resected pancreatic cancer. Genetics in Medicine, 21(1), 213–223. https://doi.org/10.1038/s41436-018-0009-5
Javle, M., Shacham-Shmueli, E., Xiao, L., Varadhachary, G., Halpern, N., Fogelman, D., et al. (2021). Olaparib monotherapy for previously treated pancreatic cancer with DNA damage repair genetic alterations other than germline BRCA variants: Findings from 2 phase 2 nonrandomized clinical trials. JAMA oncology. https://doi.org/10.1001/jamaoncol.2021.0006
Dasari, S., & Tchounwou, P. B. (2014). Cisplatin in cancer therapy: Molecular mechanisms of action. European Journal of Pharmacology, 740, 364–378. https://doi.org/10.1016/j.ejphar.2014.07.025
Farmer, H., McCabe, N., Lord, C. J., Tutt, A. N., Johnson, D. A., Richardson, T. B., et al. (2005). Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature, 434(7035), 917–921. https://doi.org/10.1038/nature03445
Pokataev, I., Fedyanin, M., Polyanskaya, E., Popova, A., Agafonova, J., Menshikova, S., et al. (2020). Efficacy of platinum-based chemotherapy and prognosis of patients with pancreatic cancer with homologous recombination deficiency: Comparative analysis of published clinical studies. ESMO Open, 5(1), e000578. https://doi.org/10.1136/esmoopen-2019-000578
Lowery, M. A., Kelsen, D. P., Stadler, Z. K., Yu, K. H., Janjigian, Y. Y., Ludwig, E., et al. (2011). An emerging entity: Pancreatic adenocarcinoma associated with a known BRCA mutation: Clinical descriptors, treatment implications, and future directions. The Oncologist, 16(10), 1397–1402. https://doi.org/10.1634/theoncologist.2011-0185
Golan, T., Kanji, Z. S., Epelbaum, R., Devaud, N., Dagan, E., Holter, S., et al. (2014). Overall survival and clinical characteristics of pancreatic cancer in BRCA mutation carriers. British journal of cancer, 111(6), 1132–1138. https://doi.org/10.1038/bjc.2014.418
Lowery, M. A., Jordan, E. J., Basturk, O., Ptashkin, R. N., Zehir, A., Berger, M. F., et al. (2017). Real-time genomic profiling of pancreatic ductal adenocarcinoma: Potential actionability and correlation with clinical phenotype. Clinical Cancer Research, 23(20), 6094–6100. https://doi.org/10.1158/1078-0432.Ccr-17-0899
Mateo, J., Lord, C. J., Serra, V., Tutt, A., Balmaña, J., Castroviejo-Bermejo, M., et al. (2019). A decade of clinical development of PARP inhibitors in perspective. Annals of Oncology, 30(9), 1437–1447. https://doi.org/10.1093/annonc/mdz192
Ashworth, A. (2008). A synthetic lethal therapeutic approach: Poly(ADP) ribose polymerase inhibitors for the treatment of cancers deficient in DNA double-strand break repair. Journal of Clinical Oncology, 26(22), 3785–3790. https://doi.org/10.1200/JCO.2008.16.0812
Moore, K., Colombo, N., Scambia, G., Kim, B.-G., Oaknin, A., Friedlander, M., et al. (2018). Maintenance olaparib in patients with newly diagnosed advanced ovarian cancer. New England Journal of Medicine, 379(26), 2495–2505. https://doi.org/10.1056/NEJMoa1810858
Golan, T., Hammel, P., Reni, M., Van Cutsem, E., Macarulla, T., Hall, M. J., et al. (2019). Maintenance olaparib for germline BRCA-mutated metastatic pancreatic cancer. New England Journal of Medicine, 381(4), 317–327. https://doi.org/10.1056/NEJMoa1903387
Hall, M. J., Golan, T., Hammel, P., Reni, M., Van Cutsem, E., Macarulla, T., et al. (2020). Pancreatic cancer (PaC)-specific health-related quality of life (HRQoL) with maintenance olaparib (O) in patients (pts) with metastatic (m) PaC and a germline BRCA mutation (gBRCAm): Phase III POLO trial. Journal of Clinical Oncology, 38(4_suppl), 648–648. https://doi.org/10.1200/JCO.2020.38.4_suppl.648
Mirza, M. R., Monk, B. J., Herrstedt, J., Oza, A. M., Mahner, S., Redondo, A., et al. (2016). Niraparib maintenance therapy in platinum-sensitive, recurrent ovarian cancer. New England Journal of Medicine, 375(22), 2154–2164. https://doi.org/10.1056/NEJMoa1611310
Binder, K. A. R., Mick, R., Hara, M., Teitelbaum, U., Karasic, T., Schneider, C., et al. (2019). Abstract CT234: A Phase II, single arm study of maintenance rucaparib in patients with platinum-sensitive advanced pancreatic cancer and a pathogenic germline or somatic mutation in <em>BRCA1, BRCA2</em> or <em>PALB2</em>. Cancer Research, 79(13 Supplement), CT234. https://doi.org/10.1158/1538-7445.AM2019-CT234
Kaufman, B., Shapira-Frommer, R., Schmutzler, R. K., Audeh, M. W., Friedlander, M., Balmaña, J., et al. (2015). Olaparib monotherapy in patients with advanced cancer and a germline BRCA1/2 mutation. Journal of Clinical Oncology, 33(3), 244–250. https://doi.org/10.1200/jco.2014.56.2728
Shroff, R. T., Hendifar, A., McWilliams, R. R., Geva, R., Epelbaum, R., Rolfe, L., et al. (2018). Rucaparib monotherapy in patients with pancreatic cancer and a known deleterious BRCA mutation. JCO Precision Oncology, 2018.https://doi.org/10.1200/po.17.00316
Lowery, M. A., Kelsen, D. P., Capanu, M., Smith, S. C., Lee, J. W., Stadler, Z. K., et al. (2018). Phase II trial of veliparib in patients with previously treated BRCA-mutated pancreas ductal adenocarcinoma. European Journal of Cancer, 89, 19–26. https://doi.org/10.1016/j.ejca.2017.11.004
O’Reilly, E. M., Lowery, M. A., Segal, M. F., Smith, S. C., Moore, M. J., Kindler, H. L., et al. (2014). Phase IB trial of cisplatin (C), gemcitabine (G), and veliparib (V) in patients with known or potential BRCA or PALB2-mutated pancreas adenocarcinoma (PC). Journal of Clinical Oncology, 32(15_suppl), 4023–4023. https://doi.org/10.1200/jco.2014.32.15_suppl.4023
Yarchoan, M., Myzak, M. C., Johnson, B. A., 3rd., De Jesus-Acosta, A., Le, D. T., Jaffee, E. M., et al. (2017). Olaparib in combination with irinotecan, cisplatin, and mitomycin C in patients with advanced pancreatic cancer. Oncotarget, 8(27), 44073–44081. https://doi.org/10.18632/oncotarget.17237
Chiorean, E. G., Guthrie, K. A., Philip, P. A., Swisher, E. M., Jalikis, F., Pishvaian, M. J., et al. (2019). Randomized phase II study of second-line modified FOLFIRI with PARP inhibitor ABT-888 (Veliparib) (NSC-737664) versus FOLFIRI in metastatic pancreatic cancer (mPC): SWOG S1513. Journal of Clinical Oncology, 37(15_suppl), 4014–4014. https://doi.org/10.1200/JCO.2019.37.15_suppl.4014
Moffat, G. T., & O’Reilly, E. M. (2020). The role of PARP inhibitors in germline BRCA-associated pancreatic ductal adenocarcinoma. Clinical Advances in Hematology & Oncology, 18(3), 168–179.
Pishvaian, M. J., Wang, H., He, A. R., Hwang, J. J., Smaglo, B. G., Kim, S. S., et al. (2020). A phase I/II study of veliparib (ABT-888) in combination with 5-fluorouracil and oxaliplatin in patients with metastatic pancreatic cancer. Clinical Cancer Research, 26(19), 5092. https://doi.org/10.1158/1078-0432.CCR-20-1301
O’Reilly, E. M., Lee, J. W., Zalupski, M., Capanu, M., Park, J., Golan, T., et al. (2020). Randomized, multicenter, phase II trial of gemcitabine and cisplatin with or without veliparib in patients with pancreas adenocarcinoma and a germline BRCA/PALB2 mutation. Journal of Clinical Oncology, 38(13), 1378–1388. https://doi.org/10.1200/jco.19.02931
Tuli, R., Shiao, S. L., Nissen, N., Tighiouart, M., Kim, S., Osipov, A., et al. (2019). A phase 1 study of veliparib, a PARP-1/2 inhibitor, with gemcitabine and radiotherapy in locally advanced pancreatic cancer. eBioMedicine, 40, 375–381. https://doi.org/10.1016/j.ebiom.2018.12.060
Li, H., Liu, Z.-Y., Wu, N., Chen, Y.-C., Cheng, Q., & Wang, J. (2020). PARP inhibitor resistance: The underlying mechanisms and clinical implications. Molecular Cancer, 19(1), 107–107. https://doi.org/10.1186/s12943-020-01227-0
D’Andrea, A. D. (2018). Mechanisms of PARP inhibitor sensitivity and resistance. DNA Repair, 71, 172–176. https://doi.org/10.1016/j.dnarep.2018.08.021
Gupta, R., Somyajit, K., Narita, T., Maskey, E., Stanlie, A., Kremer, M., et al. (2018). DNA repair network analysis reveals shieldin as a key regulator of NHEJ and PARP inhibitor sensitivity. Cell, 173(4), 972-988.e923. https://doi.org/10.1016/j.cell.2018.03.050
Kaplan, A. R., Gueble, S. E., Liu, Y., Oeck, S., Kim, H., Yun, Z., et al. (2019). Cediranib suppresses homology-directed DNA repair through down-regulation of BRCA1/2 and RAD51. Sci Transl Med, 11(492). https://doi.org/10.1126/scitranslmed.aav4508
Bindra, R. S., Gibson, S. L., Meng, A., Westermark, U., Jasin, M., Pierce, A. J., et al. (2005). Hypoxia-induced down-regulation of <em>BRCA1</em> expression by E2Fs. Cancer Research, 65(24), 11597. https://doi.org/10.1158/0008-5472.CAN-05-2119
Hegan, D. C., Lu, Y., Stachelek, G. C., Crosby, M. E., Bindra, R. S., & Glazer, P. M. (2010). Inhibition of poly(ADP-ribose) polymerase down-regulates BRCA1 and RAD51 in a pathway mediated by E2F4 and p130. Proceedings of the National Academy of Sciences, 107(5), 2201. https://doi.org/10.1073/pnas.0904783107
Ray-Coquard, I. L., Pautier, P., Pignata, S., Pérol, D., González-Martín, A., Sevelda, P., et al. (2019). LBA2_PR - Phase III PAOLA-1/ENGOT-ov25 trial: Olaparib plus bevacizumab (bev) as maintenance therapy in patients (pts) with newly diagnosed, advanced ovarian cancer (OC) treated with platinum-based chemotherapy (PCh) plus bev. Annals of Oncology, 30, v894–v895. https://doi.org/10.1093/annonc/mdz394.053
Lheureux, S., Oaknin, A., Garg, S., Bruce, J. P., Madariaga, A., Dhani, N. C., et al. (2020). EVOLVE: A multicenter open-label single-arm clinical and translational phase II trial of cediranib plus olaparib for ovarian cancer after PARP inhibition progression. Clinical Cancer Research, 26(16), 4206. https://doi.org/10.1158/1078-0432.CCR-19-4121
Liu, J. F., Brady, M. F., Matulonis, U. A., Miller, A., Kohn, E. C., Swisher, E. M., et al. (2020). A phase III study comparing single-agent olaparib or the combination of cediranib and olaparib to standard platinum-based chemotherapy in recurrent platinum-sensitive ovarian cancer. Journal of Clinical Oncology, 38(15_suppl), 6003–6003. https://doi.org/10.1200/JCO.2020.38.15_suppl.6003
Vénéreau, E., Ceriotti, C., & Bianchi, M. E. (2015). DAMPs from cell death to new life. Frontiers in Immunology, 6, 422–422. https://doi.org/10.3389/fimmu.2015.00422
Gay, C. M., & Byers, L. A. (2019). PARP inhibition combined with immune checkpoint blockade in SCLC: Oasis in an immune desert or mirage? Journal of Thoracic Oncology, 14(8), 1323–1326. https://doi.org/10.1016/j.jtho.2019.05.004
Vikas, P., Borcherding, N., Chennamadhavuni, A., & Garje, R. (2020). Therapeutic potential of combining PARP inhibitor and immunotherapy in solid tumors. Frontiers in Oncology, 10, 570–570. https://doi.org/10.3389/fonc.2020.00570
Ho, S. S., Zhang, W. Y., Tan, N. Y., Khatoo, M., Suter, M. A., Tripathi, S., et al. (2016). The DNA structure-specific endonuclease MUS81 mediates DNA sensor STING-dependent host rejection of prostate cancer cells. Immunity, 44(5), 1177–1189. https://doi.org/10.1016/j.immuni.2016.04.010
Mouw, K. W., Goldberg, M. S., Konstantinopoulos, P. A., & D’Andrea, A. D. (2017). DNA Damage and repair biomarkers of immunotherapy response. Cancer Discovery, 7(7), 675–693. https://doi.org/10.1158/2159-8290.CD-17-0226
Jiao, S., Xia, W., Yamaguchi, H., Wei, Y., Chen, M. K., Hsu, J. M., et al. (2017). PARP inhibitor upregulates PD-L1 expression and enhances cancer-associated immunosuppression. Clinical Cancer Research, 23(14), 3711–3720. https://doi.org/10.1158/1078-0432.Ccr-16-3215
Sato, H., Niimi, A., Yasuhara, T., Permata, T. B. M., Hagiwara, Y., Isono, M., et al. (2017). DNA double-strand break repair pathway regulates PD-L1 expression in cancer cells. Nature Communications, 8(1), 1751. https://doi.org/10.1038/s41467-017-01883-9
Domchek, S. M., Postel-Vinay, S., Im, S. A., Park, Y. H., Delord, J. P., Italiano, A., et al. (2020). Olaparib and durvalumab in patients with germline BRCA-mutated metastatic breast cancer (MEDIOLA): An open-label, multicentre, phase 1/2, basket study. The Lancet Oncology, 21(9), 1155–1164. https://doi.org/10.1016/s1470-2045(20)30324-7
Drew, Y., Kaufman, B., Banerjee, S., Lortholary, A., Hong, S. H., Park, Y. H., et al. (2019). 1190PD—Phase II study of olaparib + durvalumab (MEDIOLA): Updated results in germline BRCA-mutated platinum-sensitive relapsed (PSR) ovarian cancer (OC). Annals of Oncology, 30, v485–v486. https://doi.org/10.1093/annonc/mdz253.016
Kim, H., Xu, H., George, E., Hallberg, D., Kumar, S., Jagannathan, V., et al. (2020). Combining PARP with ATR inhibition overcomes PARP inhibitor and platinum resistance in ovarian cancer models. Nature Communications, 11(1), 3726. https://doi.org/10.1038/s41467-020-17127-2
Sun, C., Fang, Y., Yin, J., Chen, J., Ju, Z., Zhang, D., et al. (2017). Rational combination therapy with PARP and MEK inhibitors capitalizes on therapeutic liabilities in RAS mutant cancers. Science translational medicine, 9(392), eaal5148. https://doi.org/10.1126/scitranslmed.aal5148
Johnson, S. F., Cruz, C., Greifenberg, A. K., Dust, S., Stover, D. G., Chi, D., et al. (2016). CDK12 inhibition reverses de novo and acquired PARP inhibitor resistance in BRCA wild-type and mutated models of triple-negative breast cancer. Cell Reports, 17(9), 2367–2381. https://doi.org/10.1016/j.celrep.2016.10.077
Rundle, S., Bradbury, A., Drew, Y., & Curtin, N. J. (2017). Targeting the ATR-CHK1 axis in cancer therapy. Cancers, 9(5). https://doi.org/10.3390/cancers9050041
Fokas, E., Prevo, R., Hammond, E. M., Brunner, T. B., McKenna, W. G., & Muschel, R. J. (2014). Targeting ATR in DNA damage response and cancer therapeutics. Cancer Treatment Reviews, 40(1), 109–117. https://doi.org/10.1016/j.ctrv.2013.03.002
Karnitz, L. M., Flatten, K. S., Wagner, J. M., Loegering, D., Hackbarth, J. S., Arlander, S. J. H., et al. (2005). Gemcitabine-induced activation of checkpoint signaling pathways that affect tumor cell survival. Molecular Pharmacology, 68(6), 1636. https://doi.org/10.1124/mol.105.012716
Parsels, L. A., Morgan, M. A., Tanska, D. M., Parsels, J. D., Palmer, B. D., Booth, R. J., et al. (2009). Gemcitabine sensitization by checkpoint kinase 1 inhibition correlates with inhibition of a Rad51 DNA damage response in pancreatic cancer cells. Molecular Cancer Therapeutics, 8(1), 45. https://doi.org/10.1158/1535-7163.MCT-08-0662
Ha, D.-H., Min, A., Kim, S., Jang, H., Kim, S. H., Kim, H.-J., et al. (2020). Antitumor effect of a WEE1 inhibitor and potentiation of olaparib sensitivity by DNA damage response modulation in triple-negative breast cancer. Scientific Reports, 10(1), 9930. https://doi.org/10.1038/s41598-020-66018-5
Cuneo, K. C., Morgan, M. A., Sahai, V., Schipper, M. J., Parsels, L. A., Parsels, J. D., et al. (2019). Dose escalation trial of the Wee1 inhibitor adavosertib (AZD1775) in combination with gemcitabine and radiation for patients with locally advanced pancreatic cancer. Journal of Clinical Oncology, 37(29), 2643–2650. https://doi.org/10.1200/JCO.19.00730
Guney Eskiler, G. (2019). The interaction of PI3K inhibition with homologous recombination repair in triple negative breast cancer cells. Journal of Pharmacy & Pharmaceutical Sciences, 22(1), 599–611. https://doi.org/10.18433/jpps30684
Mo, W., Liu, Q., Lin, C. C., Dai, H., Peng, Y., Liang, Y., et al. (2016). mTOR inhibitors suppress homologous recombination repair and synergize with PARP inhibitors via regulating SUV39H1 in BRCA-proficient triple-negative breast cancer. Clinical Cancer Research, 22(7), 1699–1712. https://doi.org/10.1158/1078-0432.Ccr-15-1772
De, P., Sun, Y., Carlson, J. H., Friedman, L. S., Leyland-Jones, B. R., & Dey, N. (2014). Doubling down on the PI3K-AKT-mTOR pathway enhances the antitumor efficacy of PARP inhibitor in triple negative breast cancer model beyond BRCA-ness. Neoplasia, 16(1), 43–72. https://doi.org/10.1593/neo.131694
Funding
Cancer Center Support Grant P30 CA0008748, David M. Rubinstein Center for Pancreas Research, Paul Calabresi Career Development Award K12 CA184746, Parker Institute for Immunotherapy Pilot Grant, and Society of Immunotherapy for Cancer (SITC)—TimIOs.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
FC: No conflicts of interest to declare.
WP: Research funding to institution: Merck, Astellas, Gossamerbio.
EOR: Research funding to MSK: Genentech/Roche, Celgene/BMS, BioNTech, BioAtla, AstraZeneca, Arcus, Elicio. Consulting Role: Cytomx Therapeutics (DSMB), Rafael Therapeutics (DSMB), Sobi, Silenseed, Molecular Templates, Boehringer Ingelheim, BioNTech, Ipsen, Polaris, Tyme, Seagen, Merck, AstraZeneca, Noxxon, BioSapien, Bayer (spouse), Genentech-Roche (spouse), Celgene-BMS (spouse), Eisai (spouse).
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Crowley, F., Park, W. & O’Reilly, E.M. Targeting DNA damage repair pathways in pancreas cancer. Cancer Metastasis Rev 40, 891–908 (2021). https://doi.org/10.1007/s10555-021-09983-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10555-021-09983-1