Abstract
Purpose
Sirtuin7 (SIRT7), as a member of the sirtuin and NAD+-dependent protein-modifying enzyme family, plays an important role in regulating cellular metabolism, stress responses, tumorigenesis, and aging. Ubiquitination and deubiquitination are reversible post-translational modifications that regulate protein stability, enzyme activity, protein–protein interactions, and cellular signaling transduction. However, whether SIRT7 is regulated by deubiquitination signaling is unclear. This study aims to elucidate the molecular mechanism of SIRT7 via deubiquitination signaling.
Methods
USP17L2 or SIRT7-targeting shRNAs were used to deplete USP17L2 or SIRT7. Western blot was applied to assess the effects of USP17L2 or SIRT7 depletion. A co-immunoprecipitation assay was used to detect the interaction relationship. Cell Counting Kit-8 assays were applied to assess the viability of breast cancer cells. An immunohistochemistry assay was employed to detect the protein level in samples from breast cancer patients, and the TCGA database was applied to analyze the survival rate of breast cancer patients. Statistical analyses were performed with the Student’s t test (two-tailed unpaired) and χ2 test.
Results
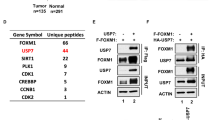
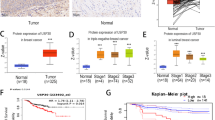
We find that the deubiquitinase USP17L2 interacts with and deubiquitinates SIRT7, thereby increasing SIRT7 protein stability. In addition, USP17L2 regulates DNA damage repair through SIRT7. Furthermore, SIRT7 polyubiquitination is increased by knocking down of USP17L2, which leads to cancer cells sensitizing to chemotherapy. In breast cancer patient samples, high expression of USP17L2 is correlated with increased levels of SIRT7 protein. In conclusion, our study demonstrates that the USP17L2-SIRT7 axis is the new regulator in DNA damage response and chemo-response, suggesting that USP17L2 may be a prognostic factor and a potential therapeutic target in breast cancer.
Conclusion
Our results highlighted that USP17L2 regulates the chemoresistance of breast cancer cells in a SIRT7-dependent manner. Moreover, the role of USP17L2 as a potential therapeutic target in breast cancer and a prognostic factor for patients was elucidated.






Similar content being viewed by others
Data availability
The datasets analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- SIRT7:
-
Sirtuin7
- HR:
-
Homologous recombination repair
- NHEJ:
-
Non-homologous end joining
- ATM:
-
Ataxia-telangiectasia mutated
- 5-FU:
-
5-Fluorouracil
- CDK1:
-
Cyclin-dependent kinase 1
- DUBs:
-
Deubiquitinating enzymes
- UPS:
-
Ubiquitin–proteasome system
- ATCC:
-
American Type Culture Collection
- DMEM:
-
Dulbecco’s modified Eagle’s medium
- FBS:
-
Fetal bovine serum
- IHC:
-
Immunohistochemical
- WT:
-
Wild-type
- CA mutant:
-
Catalytically inactive mutant
- Co-IP:
-
Co-immunoprecipitation
- IP:
-
Immunoprecipitated
- CHX:
-
Cycloheximide
- CCK-8:
-
Cell counting Kit-8
- DSBs:
-
DNA double-strand breaks
- HDAC:
-
Histone deacetylase
- AMPK:
-
Adenosine 5′-monophosphate (AMP)-activated protein kinase
- MALAT1:
-
Metastasis associated in lung adenocarcinoma transcript 1
- NRF2:
-
NF-E2-related factor 2
- BRD4:
-
Bromodomain-containing protein 4
- MCL1:
-
Myeloid cell leukemia 1
- BRCA1:
-
Breast cancer gene 1
- BRCA2:
-
Breast cancer gene 2
References
Wu D, Li Y, Zhu KS, Wang H, Zhu WG (2018) Advances in Cellular Characterization of the Sirtuin Isoform, SIRT7. Front Endocrinol (Lausanne) 9:652
Qi H, Shi X, Yu M, Liu B, Liu M, Song S, Chen S, Zou J, Zhu WG, Luo J (2018) Sirtuin 7-mediated deacetylation of WD repeat domain 77 (WDR77) suppresses cancer cell growth by reducing WDR77/PRMT5 transmethylase complex activity. J Biol Chem 293(46):17769–17779
Barber MF, Michishita-Kioi E, Xi Y, Tasselli L, Kioi M, Moqtaderi Z, Tennen RI, Paredes S, Young NL, Chen K et al (2012) SIRT7 links H3K18 deacetylation to maintenance of oncogenic transformation. Nature 487(7405):114–118
Lu YF, Xu XP, Lu XP, Zhu Q, Liu G, Bao YT, Wen H, Li YL, Gu W, Zhu WG (2020) SIRT7 activates p53 by enhancing PCAF-mediated MDM2 degradation to arrest the cell cycle. Oncogene 39(24):4650–4665
Mohrin M, Shin J, Liu Y, Brown K, Luo H, Xi Y, Haynes CM, Chen D (2015) Stem cell aging. A mitochondrial UPR-mediated metabolic checkpoint regulates hematopoietic stem cell aging. Science 347(6228):1374–1377
Nahalkova J (2015) Novel protein-protein interactions of TPPII, p53, and SIRT7. Mol Cell Biochem 409(1–2):13–22
Ryu D, Jo YS, Lo Sasso G, Stein S, Zhang H, Perino A, Lee JU, Zeviani M, Romand R, Hottiger MO et al (2014) A SIRT7-dependent acetylation switch of GABPbeta1 controls mitochondrial function. Cell Metab 20(5):856–869
Tang M, Li Z, Zhang C, Lu X, Tu B, Cao Z, Li Y, Chen Y, Jiang L, Wang H et al (2019) SIRT7-mediated ATM deacetylation is essential for its deactivation and DNA damage repair. Sci Adv 5(3):eaav1118
Tang X, Shi L, Xie N, Liu Z, Qian M, Meng F, Xu Q, Zhou M, Cao X, Zhu WG et al (2017) SIRT7 antagonizes TGF-beta signaling and inhibits breast cancer metastasis. Nat Commun 8(1):318
Ashraf N, Zino S, Macintyre A, Kingsmore D, Payne AP, George WD, Shiels PG (2006) Altered sirtuin expression is associated with node-positive breast cancer. Br J Cancer 95(8):1056–1061
Kim JK, Noh JH, Jung KH, Eun JW, Bae HJ, Kim MG, Chang YG, Shen Q, Park WS, Lee JY et al (2013) Sirtuin7 oncogenic potential in human hepatocellular carcinoma and its regulation by the tumor suppressors MiR-125a-5p and MiR-125b. Hepatology 57(3):1055–1067
Li L, Bhatia R (2013) The controversial role of Sirtuins in tumorigenesis—SIRT7 joins the debate. Cell Res 23(1):10–12
Malik S, Villanova L, Tanaka S, Aonuma M, Roy N, Berber E, Pollack JR, Michishita-Kioi E, Chua KF (2015) SIRT7 inactivation reverses metastatic phenotypes in epithelial and mesenchymal tumors. Sci Rep 5:9841
Paredes S, Villanova L, Chua KF (2014) Molecular pathways: emerging roles of mammalian Sirtuin SIRT7 in cancer. Clin Cancer Res 20(7):1741–1746
Yu H, Ye W, Wu J, Meng X, Liu RY, Ying X, Zhou Y, Wang H, Pan C, Huang W (2014) Overexpression of sirt7 exhibits oncogenic property and serves as a prognostic factor in colorectal cancer. Clin Cancer Res 20(13):3434–3445
Vazquez BN, Thackray JK, Simonet NG, Kane-Goldsmith N, Martinez-Redondo P, Nguyen T, Bunting S, Vaquero A, Tischfield JA, Serrano L (2016) SIRT7 promotes genome integrity and modulates non-homologous end joining DNA repair. EMBO J 35(14):1488–1503
Tang M, Lu X, Zhang C, Du C, Cao L, Hou T, Li Z, Tu B, Cao Z, Li Y et al (2017) Downregulation of SIRT7 by 5-fluorouracil induces radiosensitivity in human colorectal cancer. Theranostics 7(5):1346–1359
Blank MF, Grummt I (2017) The seven faces of SIRT7. Transcription 8(2):67–74
Grob A, Roussel P, Wright JE, McStay B, Hernandez-Verdun D, Sirri V (2009) Involvement of SIRT7 in resumption of rDNA transcription at the exit from mitosis. J Cell Sci 122(Pt 4):489–498
Sun L, Fan G, Shan P, Qiu X, Dong S, Liao L, Yu C, Wang T, Gu X, Li Q et al (2016) Regulation of energy homeostasis by the ubiquitin-independent REGgamma proteasome. Nat Commun 7:12497
Hershko A, Ciechanover A (1998) The ubiquitin system. Annu Rev Biochem 67:425–479
Nijman SM, Luna-Vargas MP, Velds A, Brummelkamp TR, Dirac AM, Sixma TK, Bernards R (2005) A genomic and functional inventory of deubiquitinating enzymes. Cell 123(5):773–786
Clague MJ, Urbe S, Komander D (2019) Breaking the chains: deubiquitylating enzyme specificity begets function. Nat Rev Mol Cell Biol 20(6):338–352
Yuan J, Luo K, Deng M, Li Y, Yin P, Gao B, Fang Y, Wu P, Liu T, Lou Z (2014) HERC2-USP20 axis regulates DNA damage checkpoint through Claspin. Nucleic Acids Res 42(21):13110–13121
Luo K, Li Y, Yin Y, Li L, Wu C, Chen Y, Nowsheen S, Hu Q, Zhang L, Lou Z et al (2017) USP49 negatively regulates tumorigenesis and chemoresistance through FKBP51-AKT signaling. EMBO J 36(10):1434–1446
Liu T, Yu J, Deng M, Yin Y, Zhang H, Luo K, Qin B, Li Y, Wu C, Ren T et al (2017) CDK4/6-dependent activation of DUB3 regulates cancer metastasis through SNAIL1. Nat Commun 8:13923
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25(4):402–408
Kiran S, Anwar T, Kiran M, Ramakrishna G (2015) Sirtuin 7 in cell proliferation, stress and disease: rise of the Seventh Sirtuin! Cell Signal 27(3):673–682
Michishita E, Park JY, Burneskis JM, Barrett JC, Horikawa I (2005) Evolutionarily conserved and nonconserved cellular localizations and functions of human SIRT proteins. Mol Biol Cell 16(10):4623–4635
Chen S, Seiler J, Santiago-Reichelt M, Felbel K, Grummt I, Voit R (2013) Repression of RNA polymerase I upon stress is caused by inhibition of RNA-dependent deacetylation of PAF53 by SIRT7. Mol Cell 52(3):303–313
Liu GF, Lu JY, Zhang YJ, Zhang LX, Lu GD, Xie ZJ, Cheng MB, Shen YF, Zhang Y (2016) C/EBPalpha negatively regulates SIRT7 expression via recruiting HDAC3 to the upstream-promoter of hepatocellular carcinoma cells. Biochim Biophys Acta 1859(2):348–354
Cioffi M, Vallespinos-Serrano M, Trabulo SM, Fernandez-Marcos PJ, Firment AN, Vazquez BN, Vieira CR, Mulero F, Camara JA, Cronin UP et al (2015) MiR-93 controls adiposity via inhibition of Sirt7 and Tbx3. Cell Rep 12(10):1594–1605
Han Y, Liu Y, Zhang H, Wang T, Diao R, Jiang Z, Gui Y, Cai Z (2013) Hsa-miR-125b suppresses bladder cancer development by down-regulating oncogene SIRT7 and oncogenic long non-coding RNA MALAT1. FEBS Lett 587(23):3875–3882
Shi H, Ji Y, Zhang D, Liu Y, Fang P (2016) MicroRNA-3666-induced suppression of SIRT7 inhibits the growth of non-small cell lung cancer cells. Oncol Rep 36(5):3051–3057
Jiang L, Xiong J, Zhan J, Yuan F, Tang M, Zhang C, Cao Z, Chen Y, Lu X, Li Y et al (2017) Ubiquitin-specific peptidase 7 (USP7)-mediated deubiquitination of the histone deacetylase SIRT7 regulates gluconeogenesis. J Biol Chem 292(32):13296–13311
Burrows JF, McGrattan MJ, Rascle A, Humbert M, Baek KH, Johnston JA (2004) DUB-3, a cytokine-inducible deubiquitinating enzyme that blocks proliferation. J Biol Chem 279(14):13993–14000
Hu B, Deng T, Ma H, Liu Y, Feng P, Wei D, Ling N, Li L, Qiu S, Zhang L et al (2019) Deubiquitinase DUB3 regulates cell cycle progression via stabilizing Cyclin A for proliferation of non-small cell lung cancer cells. Cells 8(4):297
Delgado-Diaz MR, Martin Y, Berg A, Freire R, Smits VA (2014) Dub3 controls DNA damage signalling by direct deubiquitination of H2AX. Mol Oncol 8(5):884–893
Jin X, Yan Y, Wang D, Ding D, Ma T, Ye Z, Jimenez R, Wang L, Wu H, Huang H (2018) DUB3 promotes BET inhibitor resistance and cancer progression by deubiquitinating BRD4. Mol Cell 71(4):592–605
Wu X, Luo Q, Zhao P, Chang W, Wang Y, Shu T, Ding F, Li B, Liu Z (2019) MGMT-activated DUB3 stabilizes MCL1 and drives chemoresistance in ovarian cancer. Proc Natl Acad Sci USA 116(8):2961–2966
Wu Y, Wang Y, Lin Y, Liu Y, Wang Y, Jia J, Singh P, Chi YI, Wang C, Dong C et al (2017) Dub3 inhibition suppresses breast cancer invasion and mestasis by promoting Snail1 degradation. Nat Commun 8:14228
Zhang Q, Zhang ZY, Du H, Li SZ, Tu R, Jia YF, Zheng Z, Song XM, Du RL, Zhang XD (2019) DUB3 deubiquitinates and stabilizes NRF2 in chemotherapy resistance of colorectal cancer. Cell Death Differ 26(11):2300–2313
Helleday T (2010) Homologous recombination in cancer development, treatment and development of drug resistance. Carcinogenesis 31:955–960
Mavaddat N, Peock S, Frost D et al (2013) Cancer risks for BRCA1 and BRCA2 mutation carriers: results from prospective analysis of EMBRACE. J Natl Cancer Inst 105(11):812–822
Venkitaraman AR (2014) Cancer suppression by the chromosome custodians, BRCA1 and BRCA2. Science 343(6178):1470–1475
Bryant HE, Schultz N, Thomas HD et al (2005) Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature 434(7035):913–917
Farmer H, McCabe N, Lord CJ et al (2005) Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434(7035):917–921
Bhattacharyya A, Ear US, Koller BH et al (2000) The breast cancer susceptibility gene BRCA1 is required for subnuclear assembly of Rad51 and survival following treatment with the DNA cross-linking agent cisplatin. J Biol Chem 275(31):23899–23903
Acknowledgements
Our study was supported by the National Natural Science Foundation of China (91749115, 32090032, 32070713) and the Natural Science Foundation of Jiangxi Province (20181ACB20021).
Author information
Authors and Affiliations
Contributions
YS carried out the molecular lab work, participated in data analysis, participated in the design of the study, and drafted the manuscript; CW participated in the design of the study and revised the manuscript; LL, YChen, XW, XJ, YChang, and YL provided experimental technology. BYu guided experimental technology. JY conceived of the study, designed the study, coordinated the study, and helped draft the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Su, Y., Wu, C., Chang, Y. et al. USP17L2-SIRT7 axis regulates DNA damage repair and chemoresistance in breast cancer cells. Breast Cancer Res Treat 196, 31–44 (2022). https://doi.org/10.1007/s10549-022-06711-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-022-06711-3