Abstract
Purpose
miRNAs have been linked to chemosensitivity of breast cancer cells in vitro. In patients, however, there is no clinically validated method for predicting chemotherapy response. The aim of this study was to assess whether (I) a specific pattern of miRNA expression in pretherapeutic biopsies can predict response to neoadjuvant chemotherapy, and (II) differential miRNA expression in residual tumor after completion of chemotherapy allows further prognostic stratification of non-responding patients.
Methods
Sixty-four patients with newly diagnosed large (≥3 cm) or locally advanced primary breast cancers who underwent neoadjuvant anthracycline/taxane-based chemotherapy were included. Relative expression of 10 miRNAs likely to be associated with chemotherapy response (miR-7,-21,-29a,-29b,-34a,-125b,-155,-200c,-340,-451) was determined by quantitative RT-PCR from pretherapeutic biopsies (n = 64) and residual invasive tumor after chemotherapy (n = 42). Pathologic complete response (pCR) defined by absence of invasive tumor served as reference standard. In addition, miRNA expression was compared with disease-free and overall survival.
Results
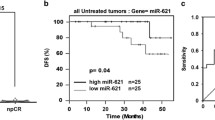
Nine (14%) of 64 patients achieved pCR. High expression of miR-7 and low expression of miR-340 in pretherapeutic biopsies predicted pCR with a negative predictive value of 96 and 97%, respectively (specificity 54 and 57%). The combined profile of miR-7high/miR-340low demonstrated improved specificity of 86% while maintaining a high negative predictive value (96%) to identify non-responders. Pretherapeutic expression of miR-200c and miR-155 showed prognostic information, and low expression was associated with increased overall survival (115 vs. 90 months, p ≤ 0.03). After chemotherapy, the overall survival of patients with residual invasive tumor was better for those demonstrating low miR-7 or high miR-125b (p = 0.01).
Conclusions
Intratumoral expression of miR-7 and miR-340 prior to neoadjuvant chemotherapy could be used to predict pCR and a profile of miR-7low or miR-340high identified patients unlikely to achieve pCR who might benefit from alternative treatment options including earlier surgery. Our study identifies miRNAs as promising predictive biomarkers, which could aid in optimization of breast cancer management and treatment stratification.



Similar content being viewed by others
References
Akalay I, Tan TZ, Kumar P et al (2015) Targeting WNT1-inducible signaling pathway protein 2 alters human breast cancer cell susceptibility to specific lysis through regulation of KLF-4 and miR-7 expression. Oncogene 34:2261–2271
Al-Khanbashi M, Al-Moundhri M (2015) Micro-ribonucleic acid and carcinogenesis: breast cancer as an example. Oncol Rev 9:279
Altman DG, Mcshane LM, Sauerbrei W et al (2012) Reporting recommendations for tumor marker prognostic studies (REMARK): explanation and elaboration. BMC Med 10:51
Betel D, Wilson M, Gabow A et al (2008) The microRNA.org resource: targets and expression. Nucleic Acids Res 36:D149–D153
Blenkiron C, Goldstein LD, Thorne NP et al (2007) MicroRNA expression profiling of human breast cancer identifies new markers of tumour subtype. Genome Biol 8:R214
Bonadonna G, Valagussa P, Brambilla C et al (1998) Primary chemotherapy in operable breast cancer: eight-year experience at the Milan Cancer Institute. J Clin Oncol 16:93–100
Bovy N, Blomme B, Freres P et al (2015) Endothelial exosomes contribute to the antitumor response during breast cancer neoadjuvant chemotherapy via microRNA transfer. Oncotarget 6:10253–10266
Buffa FM, Camps C, Winchester L et al (2011) microRNA-associated progression pathways and potential therapeutic targets identified by integrated mRNA and microRNA expression profiling in breast cancer. Cancer Res 71:5635–5645
Cekaite L, Eide PW, Lind GE et al (2016) MicroRNAs as growth regulators, their function and biomarker status in colorectal cancer. Oncotarget 7:6476–6505
Chen GQ, Zhao ZW, Zhou HY et al (2010) Systematic analysis of microRNA involved in resistance of the MCF-7 human breast cancer cell to doxorubicin. Med Oncol 27:406–415
Cortazar P, Zhang L, Untch M et al (2014) Pathological complete response and long-term clinical benefit in breast cancer: the CTNeoBC pooled analysis. Lancet 384:164–172
Doleshal M, Magotra AA, Choudhury B et al (2008) Evaluation and validation of total RNA extraction methods for microRNA expression analyses in formalin-fixed, paraffin-embedded tissues. J Mol Diagn 10:203–211
Dvinge H, Git A, Graf S et al (2013) The shaping and functional consequences of the microRNA landscape in breast cancer. Nature 497:378–382
Elston CW, Ellis IO (1991) Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. Histopathology 19:403–410
Enerly E, Steinfeld I, Kleivi K et al (2011) miRNA-mRNA integrated analysis reveals roles for miRNAs in primary breast tumors. PLoS One 6:e16915
Fabris L, Ceder Y, Chinnaiyan AM et al (2016) The potential of microRNAs as prostate cancer biomarkers. Eur Urol 70:312–322
Farazi TA, Horlings HM, Ten Hoeve JJ et al (2011) MicroRNA sequence and expression analysis in breast tumors by deep sequencing. Cancer Res 71:4443–4453
Farazi TA, Spitzer JI, Morozov P et al (2011) miRNAs in human cancer. J Pathol 223:102–115
Fisher ER, Wang J, Bryant J et al (2002) Pathobiology of preoperative chemotherapy: findings from the National Surgical Adjuvant Breast and Bowel (NSABP) protocol B-18. Cancer 95:681–695
Freres P, Josse C, Bovy N et al (2015) Neoadjuvant chemotherapy in breast cancer patients induces miR-34a and miR-122 expression. J Cell Physiol 230:473–481
Hasemeier B, Christgen M, Kreipe H et al (2008) Reliable microRNA profiling in routinely processed formalin-fixed paraffin-embedded breast cancer specimens using fluorescence labelled bead technology. BMC Biotechnol 8:90
Higgs G, Slack F (2013) The multiple roles of microRNA-155 in oncogenesis. J Clin Bioinform 3:17
Iorio MV, Ferracin M, Liu CG et al (2005) MicroRNA gene expression deregulation in human breast cancer. Cancer Res 65:7065–7070
Kastl L, Brown I, Schofield AC (2012) miRNA-34a is associated with docetaxel resistance in human breast cancer cells. Breast Cancer Res Treat 131:445–454
Kaufmann M, von Minckwitz G, Mamounas EP et al (2012) Recommendations from an international consensus conference on the current status and future of neoadjuvant systemic therapy in primary breast cancer. Ann Surg Oncol 19:1508–1516
Kim B, Fatayer H, Hanby AM et al (2013) Neoadjuvant chemotherapy induces expression levels of breast cancer resistance protein that predict disease-free survival in breast cancer. PLoS One 8:e62766
Korn EL, Mcshane LM, Troendle JF et al (2002) Identifying pre-post chemotherapy differences in gene expression in breast tumours: a statistical method appropriate for this aim. Br J Cancer 86:1093–1096
Kovalchuk O, Filkowski J, Meservy J et al (2008) Involvement of microRNA-451 in resistance of the MCF-7 breast cancer cells to chemotherapeutic drug doxorubicin. Mol Cancer Ther 7:2152–2159
Krek A, Grun D, Poy MN et al (2005) Combinatorial microRNA target predictions. Nat Genet 37:495–500
Lewis BP, Burge CB, Bartel DP (2005) Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120:15–20
Lu J, Getz G, Miska EA et al (2005) MicroRNA expression profiles classify human cancers. Nature 435:834–838
Mattiske S, Suetani RJ, Neilsen PM et al (2012) The oncogenic role of miR-155 in breast cancer. Cancer Epidemiol Biomark Prev 21:1236–1243
Mei M, Ren Y, Zhou X et al (2010) Downregulation of miR-21 enhances chemotherapeutic effect of taxol in breast carcinoma cells. Technol Cancer Res Treat 9:77–86
Perou CM, Sorlie T, Eisen MB et al (2000) Molecular portraits of human breast tumours. Nature 406:747–752
Pogribny IP, Filkowski JN, Tryndyak VP et al (2010) Alterations of microRNAs and their targets are associated with acquired resistance of MCF-7 breast cancer cells to cisplatin. Int J Cancer 127:1785–1794
Provenzano E, Bossuyt V, Viale G et al (2015) Standardization of pathologic evaluation and reporting of postneoadjuvant specimens in clinical trials of breast cancer: recommendations from an international working group. Modern Pathol 28(9):1185–1201
Raychaudhuri M, Schuster T, Buchner T et al (2012) Intratumoral heterogeneity of microRNA expression in breast cancer. J Mol Diagn 14:376–384
Rodriguez-Gonzalez FG, Sieuwerts AM, Smid M et al (2011) MicroRNA-30c expression level is an independent predictor of clinical benefit of endocrine therapy in advanced estrogen receptor positive breast cancer. Breast Cancer Res Treat 127:43–51
Rudas M, Filipits M, Taucher S et al (2003) Expression of MRP1, LRP and Pgp in breast carcinoma patients treated with preoperative chemotherapy. Breast Cancer Res Treat 81:149–157
Sassen S, Miska EA, Caldas C (2008) MicroRNA: implications for cancer. Virchows Archiv 452:1–10
Schwarz-Dose J, Untch M, Tiling R et al (2009) Monitoring primary systemic therapy of large and locally advanced breast cancer by using sequential positron emission tomography imaging with [18F]fluorodeoxyglucose. J Clin Oncol 27:535–541
Siebolts U, Varnholt H, Drebber U et al (2009) Tissues from routine pathology archives are suitable for microRNA analyses by quantitative PCR. J Clin Pathol 62:84–88
Symmans WF, Peintinger F, Hatzis C et al (2007) Measurement of residual breast cancer burden to predict survival after neoadjuvant chemotherapy. J Clin Oncol 25:4414–4422
Tavazoie SF, Alarcon C, Oskarsson T et al (2008) Endogenous human microRNAs that suppress breast cancer metastasis. Nature 451:147–152
Tuomarila M, Luostari K, Soini Y et al (2014) Overexpression of microRNA-200c predicts poor outcome in patients with PR-negative breast cancer. PLoS One 9:e109508
Untch M, Mobus V, Kuhn W et al (2009) Intensive dose-dense compared with conventionally scheduled preoperative chemotherapy for high-risk primary breast cancer. J Clin Oncol 27:2938–2945
van der Hage JA, van de Velde CJ, Julien JP et al (2001) Preoperative chemotherapy in primary operable breast cancer: results from the European Organization for Research and Treatment of Cancer trial 10902. J Clin Oncol 19:4224–4237
Varamo C, Occelli M, Vivenza D et al (2016) MicroRNAs role as potential biomarkers and key regulators in melanoma. Genes Chromosom Cancer. doi:10.1002/gcc.22402
Webster RJ, Giles KM, Price KJ et al (2009) Regulation of epidermal growth factor receptor signaling in human cancer cells by microRNA-7. J Biol Chem 284:5731–5741
Zheng Y, Li S, Boohaker RJ et al (2015) A microRNA expression signature in taxane-anthracycline-based neoadjuvant chemotherapy response. J Cancer 6:671–677
Acknowledgements
This study was funded by the Deutsche Forschungsgemeinschaft (German Research Foundation, DFG) Grant No SA1698/1-2 awarded to SA. MR was supported by a fellowship of the Technische Universität München. SA is supported by the Clinical and Translational Science Collaborative of Cleveland (KL2TR000440) from the National Center for Advancing Translational Sciences (NCATS) component of the NIH.
Author contributions
S.A. conceived of the study. S.A., M.R., and H.B. analyzed and interpreted data. S.A., M.R. and T.B. carried out experiments. H.B. provided study material and clinical follow-up data. W.W. and M.K. participated in data interpretation. All authors were involved in writing the paper and had final approval of the submitted and published versions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed involving human participants were in accordance with the ethical standards of the institutional and national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Additional information
Mithu Raychaudhuri and Holger Bronger have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Raychaudhuri, M., Bronger, H., Buchner, T. et al. MicroRNAs miR-7 and miR-340 predict response to neoadjuvant chemotherapy in breast cancer. Breast Cancer Res Treat 162, 511–521 (2017). https://doi.org/10.1007/s10549-017-4132-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-017-4132-9