Abstract
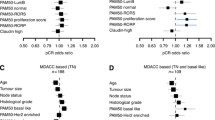
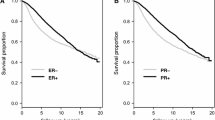
The goal of this study was to assess the prognostic value of a 3-gene (TOP2A, FOXM1, and MKI67) proliferation score and use it to risk stratify grade-2, estrogen receptor (ER)-positive breast cancers into low- and high-risk groups. We used 4 different breast cancer gene expression datasets including two cohorts of patients who received no systemic adjuvant therapy (Mainz: n = 206, TRANSBIG: n = 134) and two other cohorts that received adjuvant tamoxifen (JBI: n = 227, MDACC/SET: n = 192). We compared individual and combined expression values of the 3 genes between grade 1, 2, and 3 tumors and plotted distant metastasis-free survival (DMFS) curves by the 3-gene score for grade-2 cancers. We compared the prognostic value of the 3-gene score to the Genomic Grade Index (GGI). The individual and combined expression of TOP2A, FOXM1, and MKI67 were significantly different between the 3 histological grade groups with the highest expression in grade-3 and the lowest in grade-1 cancers. Expression levels were variable in grade-2 cancers. Grade-2 tumors with high expression of the 3 genes (>median) showed significantly worse DMFS in one prognostic and one tamoxifen-treated set and showed a similar but non-significant trend for worse survival in the remaining two datasets. The 3-gene score performed equally well in risk stratification as the GGI. A 3-gene proliferation score shows similar prognostic value as the GGI in ER-positive, grade-2 cancers and may serve as basis for a PCR-based assay that could aid prognostic prediction for clinically intermediate-risk cancers.



Similar content being viewed by others
References
Galea MH, Blamey RW, Elston CE, Ellis IO (1992) The Nottingham prognostic index in primary breast cancer. Breast Cancer Res Treat 22(3):207–219
Elston CW, Ellis IO (1991) Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. Histopathology 19(5):403–410
Simpson JF, Gray R, Dressler LG et al (2000) Prognostic value of histologic grade and proliferative activity in axillary node-positive breast cancer: results from the Eastern Cooperative Oncology Group Companion Study, EST 4189. J Clin Oncol 18:2059–2069
Frierson HF, Wolber RA, Berean KW et al (1995) Interobserver reproducibility of the Nottingham modification of the Bloom and Richardson histologic grading scheme for infiltrating ductal carcinoma. Am J Clin Pathol 103:195–198
Dalton LW, Page DL, Dupont WD (1994) Histologic grading of breast carcinoma: a reproducibility study. Cancer 73:2765–2770
Robbins P, Pinder S, de Klerk N et al (1995) Histologic grading of breast carcinomas: a study of interobserver agreement. Hum Pathol 26:873–879
Sotiriou C, Wirapati P, Loi S, Harris A, Fox S, Smeds J, Nordgren H, Farmer P, Praz V, Haibe-Kains B, Desmedt C, Larsimont D, Cardoso F, Peterse H, Nuyten D, Buyse M, Van de Vijver MJ, Bergh J, Piccart M, Delorenzi M (2006) Gene expression profiling in breast cancer: understanding the molecular basis of histologic grade to improve prognosis. J Natl Cancer Inst 98(4):262–272
Wirapati P, Sotiriou C, Kunkel S, Farmer P, Pradervand S, Haibe-Kains B, Desmedt C, Ignatiadis M, Sengstag T, Schutz F, Goldstein DR, Piccart M, Delorenzi M (2008) Meta-analysis of gene expression profiles in breast cancer: toward a unified understanding of breast cancer subtyping and prognosis signatures. Breast Cancer Res 10(4):R65
Toussaint J, Sieuwerts AM, Haibe-Kains B, Desmedt C, Rouas G, Harris AL, Larsimont D, Piccart M, Foekens JA, Durbecq V, Sotiriou C (2009) Improvement of the clinical applicability of the genomic grade index through a qrt-pcr test performed on frozen and formalin-fixed paraffin-embedded tissues. BMC Genomics 10:424. doi:10.1186/1471-2164-10-424
Ma XJ, Salunga R, Dahiya S, Wang W, Carney E, Durbecq V, Harris A, Goss P, Sotiriou C, Erlander M, Sgroi D (2008) A five-gene molecular grade index and hoxb13:Il17br are complementary prognostic factors in early stage breast cancer. Clin Cancer Res 14(9):2601–2608
Aleskandarany MA, Rakha EA, Macmillan RD, Powe DG, Ellis IO, Green AR (2010) Mib1/ki-67 labelling index can classify grade 2 breast cancer into two clinically distinct subgroups. Breast Cancer Res Treat 127(3):591–599. doi:10.1007/s10549-010-1028-3
Wang JC (2002) Cellular roles of DNA topoisomerases: a molecular perspective. Nat Rev Mol Cell Biol 3:430–440
Myatt SS, Lam EW (2007) The emerging roles of forkhead box (fox) proteins in cancer. Nat Rev Cancer 7(11):847–859
Wierstra I, Alves J (2007) Foxm1, a typical proliferation-associated transcription factor. Biol Chem 388(12):1257–1274
Duchrow M, Gerdes J, Schlüter C (1994) The proliferation-associated Ki-67 protein: definition in molecular terms. Cell Prolif 27:235–242
Bueno-de-Mesquita JM, van Harten WH, Retel VP, van’t Veer LJ, van Dam FS, Karsenberg K, Douma KF, van Tinteren H, Peterse JL, Wesseling J, Wu TS, Atsma D, Rutgers EJ, Brink G, Floore AN, Glas AM, Roumen RM, Bellot FE, van Krimpen C, Rodenhuis S, van de Vijver MJ, Linn SC (2007) Use of 70-gene signature to predict prognosis of patients with node-negative breast cancer: a prospective community-based feasibility study (RASTER). Lancet Oncol 8(12):1079–1087
Goldstein LJ, Gray R, Badve S, Childs BH, Yoshizawa C, Rowley S, Shak S, Baehner FL, Ravdin PM, Davidson NE, Sledge GW Jr, Perez EA, Shulman LN, Martino S, Sparano JA (2008) Prognostic utility of the 21-gene assay in hormone receptor-positive operable breast cancer compared with classical clinicopathologic features. J Clin Oncol 26(25):4063–4071
Schmidt M, Bohm D, von Torne C, Steiner E, Puhl A, Pilch H, Lehr HA, Hengstler JG, Kolbl H, Gehrmann M (2008) The humoral immune system has a key prognostic impact in node-negative breast cancer. Cancer Res 68(13):5405–5413
Desmedt C, Piette F, Loi S, Wang Y, Lallemand F, Haibe-Kains B, Viale G, Delorenzi M, Zhang Y, d’Assignies MS, Bergh J, Lidereau R, Ellis P, Harris AL, Klijn JG, Foekens JA, Cardoso F, Piccart MJ, Buyse M, Sotiriou C (2007) Strong time dependence of the 76-gene prognostic signature for node-negative breast cancer patients in the transbig multicenter independent validation series. Clin Cancer Res 13(11):3207–3214
Loi S, Haibe-Kains B, Desmedt C, Lallemand F, Tutt AM, Gillet C, Ellis P, Harris A, Bergh J, Foekens JA, Klijn JG, Larsimont D, Buyse M, Bontempi G, Delorenzi M, Piccart MJ, Sotiriou C (2007) Definition of clinically distinct molecular subtypes in estrogen receptor-positive breast carcinomas through genomic grade. J Clin Oncol 25(10):1239–1246
Symmans WF, Hatzis C, Sotiriou C, Andre F, Peintinger F, Regitnig P, Daxenbichler G, Desmedt C, Domont J, Marth C, Delaloge S, Bauernhofer T, Valero V, Booser DJ, Hortobagyi GN, Pusztai L (2010) Genomic index of sensitivity to endocrine therapy for breast cancer. J Clin Oncol 28:4273
Gong Y, Yan K, Lin F, Anderson K, Sotiriou C, Andre F, Holmes FA, Valero V, Booser D, Pippen JE Jr, Vukelja S, Gomez H, Mejia J, Barajas LJ, Hess KR, Sneige N, Hortobagyi GN, Pusztai L, Symmans WF (2007) Determination of oestrogen-receptor status and erbb2 status of breast carcinoma: a gene-expression profiling study. Lancet Oncol 8(3):203–211
Haibe-Kains B, Desmedt C, Piette F, Buyse M, Cardoso F, Van’t Veer L et al (2008) Comparison of prognostic gene expression signatures for breast cancer. BMC Genomics 9:394
Goldhirsch A, Wood WC, Gelber RD et al (2007) Progress and promise: highlights of the international expert consensus on the primary therapy of early breast cancer 2007. Ann Oncol 18:1133–1144
Paik S, Shak S, Tang G, Kim C, Baker J, Cronin M, Baehner FL, Walker MG, Watson D, Park T, Hiller W, Fisher ER, Wickerham DL, Bryant J, Wolmark N (2004) A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med 351(27):2817–2826
Sotiriou C, Pusztai L (2009) Gene expression signatures in breast cancer. N Engl J Med 360(8):790–800
Szasz AM, Li Q, Eklund AC, Sztupinszki Z, Rowan A, Tokes AM, Szekely B, Kiss A, Szendroi M, Gyorffy B, Szallasi Z, Swanton C, Kulka J (2013) The CIN4 chromosomal instability qPCR classifier defines tumor aneuploidy and stratifies outcome in grade 2 breast cancer. PLoS One 8(2):e56707
Acknowledgments
Grants from the Breast Cancer Research Foundation (LP and CS). TÁMOP-4.2.2/B-10/1-2010-0013 and 2012TKI643 (KJ).
Conflict of interest
The authors have no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Borbala Szekely and Takayuki Iwamoto contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Szekely, B., Iwamoto, T., Szasz, A.M. et al. A 3-gene proliferation score (TOP-FOX-67) can re-classify histological grade-2, ER-positive breast cancers into low- and high-risk prognostic categories. Breast Cancer Res Treat 138, 691–698 (2013). https://doi.org/10.1007/s10549-013-2475-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-013-2475-4