Abstract
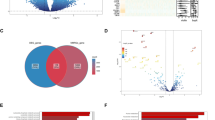
Triple-negative breast cancer is a particularly aggressive and lethal breast cancer subtype that is more likely to be interval-detected rather than screen-detected. The purpose of this study is to discover and initially validate novel early detection biomarkers for triple-negative breast cancer using preclinical samples. Plasma samples collected up to 17 months before diagnosis from 28 triple-negative cases and 28 matched controls from the Women’s Health Initiative Observational Study were equally divided into a training set and a test set and interrogated by a customized antibody array. Data were available on 889 antibodies; in the training set, statistically significant differences in case versus control signals were observed for 93 (10.5 %) antibodies at p < 0.05. Of these 93 candidates, 29 were confirmed in the test set at p < 0.05. Areas under the curve for these candidates ranged from 0.58 to 0.79. With specificity set at 98 %, sensitivity ranged from 4 to 68 % with 20 candidates having a sensitivity ≥20 % and 6 having a sensitivity ≥40 %. In an analysis of KEGG gene sets, the pyrimidine metabolism gene set was upregulated in cases compared to controls (p = 0.004 in the testing set) and the JAK/Stat signaling pathway gene set was downregulated (p = 0.003 in the testing set). Numerous potential early detection biomarkers specific to triple-negative breast cancer in multiple pathways were identified. Further research is required to followup on promising candidates in larger sample sizes and to better understand their potential biologic importance as our understanding of the etiology of triple-negative breast cancer continues to grow.
Similar content being viewed by others
References
Bebenek M, Dus D, Kozlak J (2006) Fas and Fas ligand as prognostic factors in human breast carcinoma. Med Sci Monit 12:CR457–CR461
Bebenek M, Dus D, Kozlak J (2007) Fas/Fas-ligand expressions in primary breast cancer are significant predictors of its skeletal spread. Anticancer Res 27:215–218
Brown M, Tsodikov A, Bauer KR, Parise CA, Caggiano V (2008) The role of human epidermal growth factor receptor 2 in the survival of women with estrogen and progesterone receptor-negative, invasive breast cancer: the California Cancer Registry, 1999–2004. Cancer 112:737–747
Campbell TN, Attwell S, Arcellana-Panlilio M, Robbins SM (2006) Ephrin A5 expression promotes invasion and transformation of murine fibroblasts. Biochem Biophys Res Commun 350:623–628
Carey LA, Perou CM, Livasy CA, Dressler LG, Cowan D, Conway K, Karaca G, Troester MA, Tse CK, Edmiston S, Deming SL, Geradts J, Cheang MC, Nielsen TO, Moorman PG, Earp HS, Millikan RC (2006) Race, breast cancer subtypes, and survival in the Carolina Breast Cancer Study. JAMA 295:2492–2502
Collett K, Stefansson IM, Eide J, Braaten A, Wang H, Eide GE, Thoresen SO, Foulkes WD, Akslen LA (2005) A basal epithelial phenotype is more frequent in interval breast cancers compared with screen detected tumors. Cancer Epidemiol Biomarkers Prev 14:1108–1112
Dunnwald LK, Rossing MA, Li CI (2007) Hormone receptor status, tumor characteristics, and prognosis: a prospective cohort of breast cancer patients. Breast Cancer Res 9:R6
Gaudet MM, Press MF, Haile RW, Lynch CF, Glaser SL, Schildkraut J, Gammon MD, Douglas TW, Bernstein JL (2011) Risk factors by molecular subtypes of breast cancer across a population-based study of women 56 years or younger. Breast Cancer Res Treat 130:587–597
Hays J, Hunt JR, Hubbell FA, Anderson GL, Limacher M, Allen C, Rossouw JE (2003) The Women’s Health Initiative recruitment methods and results. Ann Epidemiol 13:S18–S77
Kalin M, Cima I, Schiess R, Fankhauser N, Powles T, Wild P, Templeton A, Cerny T, Aebersold R, Krek W, Gillessen S (2011) Novel prognostic markers in the serum of patients with castration-resistant prostate cancer derived from quantitative analysis of the pten conditional knockout mouse proteome. Eur Urol 60:1235–1243
Kaplan HG, Malmgren JA (2008) Impact of triple negative phenotype on breast cancer prognosis. Breast J 14:456–463
Kim MJ, Ro JY, Ahn SH, Kim HH, Kim SB, Gong G (2006) Clinicopathologic significance of the basal-like subtype of breast cancer: a comparison with hormone receptor and Her2/neu-overexpressing phenotypes. Hum Pathol 37:1217–1226
Lehmann BD, Bauer JA, Chen X, Sanders ME, Chakravarthy AB, Shyr Y, Pietenpol JA (2011) Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J Clin Invest 121:2750–2767
Loch CP, Ramirez AB, Liu Y, Sather CL, Delrow JJ, Scholler N, Garvik GM, Urban ND, McIntosh MW, Lampe PD (2007) Use of high density antibody arrays to validate and discover cancer serum biomarkers. Mol Oncol 1:313–320
Lund MJ, Trivers KF, Porter PL, Coates RJ, Leyland-Jones B, Brawley OW, Flagg EW, O’Regan RM, Gabram SG, Eley JW (2009) Race and triple negative threats to breast cancer survival: a population-based study in Atlanta, GA. Breast Cancer Res Treat 113:357–370
Ma H, Wang Y, Sullivan-Halley J, Weiss L, Marchbanks PA, Spirtas R, Ursin G, Burkman RT, Simon MS, Malone KE, Strom BL, McDonald JA, Press MF, Bernstein L (2010) Use of four biomarkers to evaluate the risk of breast cancer subtypes in the women’s contraceptive and reproductive experiences study. Cancer Res 70:575–587
Metzger-Filho O, Tutt A, de AE, Saini KS, Viale G, Loi S, Bradbury I, Bliss JM, Azim HA, Jr., Ellis P, Di LA, Baselga J, Sotiriou C, Piccart-Gebhart M (2012) Dissecting the heterogeneity of triple-negative breast cancer. J Clin Oncol (In press)
Nelson HD, Tyne K, Naik A, Bougatsos C, Chan BK, Humphrey L (2009) Screening for breast cancer: an update for the U.S. preventive services task force. Ann Intern Med 151:727–742
Omenn GS (2004) The human proteome organization plasma proteome project pilot phase: reference specimens, technology platform comparisons, and standardized data submissions and analyses. Proteomics 4:1235–1240
Perou CM, Sorlie T, Eisen MB, van de RM, Jeffrey SS, Rees CA, Pollack JR, Ross DT, Johnsen H, Akslen LA, Fluge O, Pergamenschikov A, Williams C, Zhu SX, Lonning PE, Borresen-Dale AL, Brown PO, Botstein D (2000) Molecular portraits of human breast tumours. Nature 406:747–752
Phipps AI, Buist DS, Malone KE, Barlow WE, Porter PL, Kerlikowske K, Li CI (2011) First-degree family history of breast cancer and triple-negative breast cancer risk. Breast Cancer Res Treat 136:671–678
Phipps AI, Chlebowski R, Prentice R, McTiernan A, Wactawski-Wende J, Kuller LH, Adams-Campbell LL, Lane D, Stefanick ML, Vitolins M, Kabat GC, Rohan TE, Li CI (2010) Reproductive history and oral contraceptive use in relation to risk of triple-negative breast cancer. J Natl Cancer Inst (In press)
Phipps AI, Chlebowski RT, Prentice R, McTiernan A, Stefanick ML, Wactawski-Wende J, Kuller LH, Adams-Campbell LL, Lane D, Vitolins M, Kabat GC, Rohan TE, Li CI (2011) Body size, physical activity, and risk of triple-negative and estrogen receptor-positive breast cancer. Cancer Epidemiol Biomarkers Prev 20:454–463
Phipps AI, Malone KE, Porter PL, Daling JR, Li CI (2008) Body size and risk of luminal, HER2-overexpressing, and triple-negative breast cancer in postmenopausal women. Cancer Epidemiol Biomarkers Prev 17:2078–2086
Phipps AI, Malone KE, Porter PL, Daling JR, Li CI (2008) Reproductive and hormonal risk factors for postmenopausal luminal, HER-2-overexpressing, and triple-negative breast cancer. Cancer 113:1521–1526
Porter PL, El-Bastawissi AY, Mandelson MT, Lin MG, Khalid N, Watney EA, Cousens L, White D, Taplin S, White E (1999) Breast tumor characteristics as predictors of mammographic detection: comparison of interval- and screen-detected cancers. J Natl Cancer Inst 91:2020–2028
Rakha EA, Reis-Filho JS, Ellis IO (2008) Basal-like breast cancer: a critical review. J Clin Oncol 26:2568–2581
Ramirez AB, Lampe PD (2010) Discovery and validation of ovarian cancer biomarkers utilizing high density antibody microarrays. Cancer Biomark 8:293–307
Ramirez AB, Loch CM, Zhang Y, Liu Y, Wang X, Wayner EA, Sargent JE, Sibani S, Hainsworth E, Mendoza EA, Eugene R, Labaer J, Urban ND, McIntosh MW, Lampe PD (2010) Use of a single-chain antibody library for ovarian cancer biomarker discovery. Mol Cell Proteomics 9:1449–1460
Scholler N, Gross JA, Garvik B, Wells L, Liu Y, Loch CM, Ramirez AB, McIntosh MW, Lampe PD, Urban N (2008) Use of cancer-specific yeast-secreted in vivo biotinylated recombinant antibodies for serum biomarker discovery. J Transl Med 6:41
Smyth GK, Speed T (2003) Normalization of cDNA microarray data. Methods 31:265–273
Sorlie T, Tibshirani R, Parker J, Hastie T, Marron JS, Nobel A, Deng S, Johnsen H, Pesich R, Geisler S, Demeter J, Perou CM, Lonning PE, Brown PO, Borresen-Dale AL, Botstein D (2003) Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc Natl Acad Sci USA 100:8418–8423
The Women’s Health Initiative Study Group (1998) Design of the Women’s Health Initiative clinical trial and observational study. Control Clin Trials 19:61–109
Thompson IM, Ankerst DP, Chi C, Lucia MS, Goodman PJ, Crowley JJ, Parnes HL, Coltman CA Jr (2005) Operating characteristics of prostate-specific antigen in men with an initial PSA level of 3.0 ng/ml or lower. JAMA 294:66–70
White DE, Kurpios NA, Zuo D, Hassell JA, Blaess S, Mueller U, Muller WJ (2004) Targeted disruption of beta1-integrin in a transgenic mouse model of human breast cancer reveals an essential role in mammary tumor induction. Cancer Cell 6:159–170
Acknowledgments
This work was supported by the National Cancer Institute (grant number U01-CA152637) and the National Heart Lung and Blood Institute (grant number N01-WH-74313).
Conflicts of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, C.I., Mirus, J.E., Zhang, Y. et al. Discovery and preliminary confirmation of novel early detection biomarkers for triple-negative breast cancer using preclinical plasma samples from the Women’s Health Initiative observational study. Breast Cancer Res Treat 135, 611–618 (2012). https://doi.org/10.1007/s10549-012-2204-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-012-2204-4