Abstract
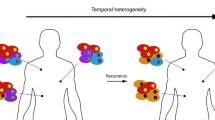
Conflicting theories of epithelial carcinogenesis disagree on the clonal composition of primary tumors and on the time at which metastases occur. In order to study the spatial distribution of disparate clonal populations within breast carcinomas and the extent of the genetic relationship between primary tumors and regional metastases, we have analyzed by comparative genomic hybridization 122 tissue samples from altogether 60 breast cancer patients, including 34 tumor samples obtained from different quadrants of 9 breast carcinomas, as well as paired primary-metastatic samples from 12 patients. The median intratumor genetic heterogeneity score (HS) was 17.4% and unsupervised hierarchical clustering analysis comparing the genetic features to those of an independent series of 41 breast carcinomas confirmed intratumor clonal divergence in a high proportion of cases. The median HS between paired primary breast tumors and lymph node metastases was 33.3%, but the number of genomic imbalances did not differ significantly. Clustering analysis confirmed extensive clonal divergence between primary carcinomas and lymph node metastases in several cases. In the independent series of 41 breast carcinomas, the number of genomic imbalances in primary tumors was significantly higher in patients presenting lymph node metastases (median = 15.5) than in the group with no evidence of disease spreading at diagnosis (median = 5.0). We conclude that primary breast carcinomas may be composed of several genetically heterogeneous and spatially separated cell populations and that paired primary breast tumors and lymph node metastases often present divergent clonal evolution, indicating that metastases may occur relatively early during breast carcinogenesis.
Similar content being viewed by others
References
Nowell PC (1976) The clonal evolution of tumor cell populations. Science 194:23–28
Heim S, Mandahl N, Mitelman F (1988) Genetic convergence and divergence in tumor progression. Cancer Res 48:5911–5916
Weigelt B, Peterse JL, van’t Veer LJ (2005) Breast cancer metastasis: markers and models. Nat Rev Cancer 5:591–602
Pantel K, Brakenhoff RH (2004) Dissecting the metastatic cascade. Nat Rev Cancer 4:448–456
Teixeira MR, Pandis N, Heim S (2002) Cytogenetic clues to breast carcinogenesis. Genes Chromosomes Cancer 33:1–16
Teixeira MR, Pandis N, Bardi G et al (1995) Clonal heterogeneity in breast cancer: karyotypic comparisons of multiple intra- and extra-tumorous samples from 3 patients. Int J Cancer 63:63–68
Teixeira MR, Pandis N, Bardi G et al (1996) Karyotypic comparisons of multiple tumorous and macroscopically normal surrounding tissue samples from patients with breast cancer. Cancer Res 56:855–859
Pandis N, Teixeira MR, Adeyinka A et al (1998) Cytogenetic comparison of primary tumors and lymph node metastases in breast cancer patients. Genes Chromosomes Cancer 22:122–129
Nishizaki T, DeVries S, Chew K et al (1997) Genetic alterations in primary breast cancers and their metastases: direct comparison using modified comparative genomic hybridization. Genes Chromosomes Cancer 19:267–272
Kuukasjarvi T, Karhu R, Tanner M et al (1997) Genetic heterogeneity and clonal evolution underlying development of asynchronous metastasis in human breast cancer. Cancer Res 57:1597–1604
Sobin LH (1981) Histological typing of breast tumors, 2nd edn. World Health Organization, Geneva
Kallioniemi OP, Kallioniemi A, Piper J et al (1994) Optimizing comparative genomic hybridization for analysis of DNA sequence copy number changes in solid tumors. Genes Chromosomes Cancer 10:231–243
Teixeira MR, Ribeiro FR, Torres L et al (2004) Assessment of clonal relationships in ipsilateral and bilateral multiple breast carcinomas by comparative genomic hybridisation and hierarchical clustering analysis. Br J Cancer 91:775–782
Kirchhoff M, Gerdes T, Rose H et al (1998) Detection of chromosomal gains and losses in comparative genomic hybridization analysis based on standard reference intervals. Cytometry 31:163–173
ISCN (1995) An international system for human cytogenetic nomenclature. S. Karger, Basel
Waldman FM, DeVries S, Chew KL et al (2000) Chromosomal alterations in ductal carcinomas in situ and their in situ recurrences. J Natl Cancer Inst 92:313–320
Chen LC, Kurisu W, Ljung BM et al (1992) Heterogeneity for allelic loss in human breast cancer. J Natl Cancer Inst 84:506–510
Aubele M, Mattis A, Zitzelsberger H et al (1999) Intratumoral heterogeneity in breast carcinoma revealed by laser-microdissection and comparative genomic hybridization. Cancer Genet Cytogenet 110:94–102
Beerman H, Smit VT, Kluin PM et al (1991) Flow cytometric analysis of DNA stemline heterogeneity in primary and metastatic breast cancer. Cytometry 12:147–154
Barranco SC, Perry RR, Durm ME et al (1994) Intratumor variability in prognostic indicators may be the cause of conflicting estimates of patient survival and response to therapy. Cancer Res 54:5351–5356
Arnerlov C, Emdin SO, Cajander S et al (2001) Intratumoral variations in DNA ploidy and s-phase fraction in human breast cancer. Anal Cell Pathol 23:21–28
Braakhuis BJ, Tabor MP, Kummer JA et al (2003) A genetic explanation of Slaughter’s concept of field cancerization: evidence and clinical implications. Cancer Res 63:1727–1730
Tomlinson IP (2001) Mutations in normal breast tissue and breast tumours. Breast Cancer Res 3:299–303
Steinarsdottir M, Jonasson JG, Vidarsson H et al (2004) Cytogenetic changes in nonmalignant breast tissue. Genes Chromosomes Cancer 41:47–55
Deng G, Lu Y, Zlotnikov G, Thor AD et al (1996) Loss of heterozygosity in normal tissue adjacent to breast carcinomas. Science 274:2057–2059
Lakhani SR, Chaggar R, Davies S et al (1999) Genetic alterations in ‘normal’ luminal and myoepithelial cells of the breast. J Pathol 189:496–503
Li Z, Moore DH, Meng ZH et al (2002) Increased risk of local recurrence is associated with allelic loss in normal lobules of breast cancer patients. Cancer Res 62:1000–1003
Braakhuis BJ, Leemans CR, Brakenhoff RH (2004) Using tissue adjacent to carcinoma as a normal control: an obvious but questionable practice. J Pathol 203:620–621
Aubele M, Auer G, Braselmann H et al (2002) Chromosomal imbalances are associated with metastasis-free survival in breast cancer patients. Anal Cell Pathol 24:77–87
Hislop RG, Pratt N, Stocks SC et al (2002) Karyotypic aberrations of chromosomes 16 and 17 are related to survival in patients with breast cancer. Br J Surg 89:1581–1586
Zudaire I, Odero MD, Caballero C et al (2002) Genomic imbalances detected by comparative genomic hybridization are prognostic markers in invasive ductal breast carcinomas. Histopathology 40:547–555
Isola JJ, Kallioniemi OP, Chu LW et al (1995) Genetic aberrations detected by comparative genomic hybridization predict outcome in node-negative breast cancer. Am J Pathol 147:905–911
Hermsen MA, Baak JP, Meijer GA et al (1998) Genetic analysis of 53 lymph node-negative breast carcinomas by CGH and relation to clinical, pathological, morphometric, and DNA cytometric prognostic factors. J Pathol 186:356–362
Dellas A, Torhorst J, Schultheiss E et al (2002) DNA sequence losses on chromosomes 11p and 18q are associated with clinical outcome in lymph node-negative ductal breast cancer. Clin Cancer Res 8:1210–1216
Teixeira MR, Tsarouha H, Kraggerud SM et al (2001) Evaluation of breast cancer polyclonality by combined chromosome banding and comparative genomic hybridization analysis. Neoplasia 3:204–214
Kleivi K, Lothe RA, Heim S et al (2002) Genome profiling of breast cancer cells selected against in vitro shows copy number changes. Genes Chromosomes Cancer 33:304–309
Acknowledgments
F.R.R. is a fellow of Fundação para a Ciência e a Tecnologia. L.T. is a fellow of Liga Portuguesa Contra o Cancro, Centro Regional do Norte. The financial support of Liga Portuguesa Contra o Cancro, Centro Regional do Norte, and the Norwegian Cancer Society is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Torres, L., Ribeiro, F.R., Pandis, N. et al. Intratumor genomic heterogeneity in breast cancer with clonal divergence between primary carcinomas and lymph node metastases. Breast Cancer Res Treat 102, 143–155 (2007). https://doi.org/10.1007/s10549-006-9317-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-006-9317-6