Abstract
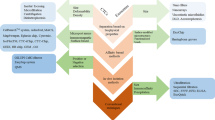
Cancer cells release extracellular vesicles known as extracellular vesicles (EVs), containing tumor-derived DNA, RNA and proteins within their cargo, into the circulation. Circulating tumor-derived extracellular vesicles (TEV) can be used in the context of serial “liquid biopsies” for early detection of cancer, for monitoring disease burden in patients, and for assessing recurrence in the post-resection setting. Nonetheless, isolating sufficient TEV by ultracentrifugation-based approaches, in order to enable molecular assessment of EVs cargo, can be an arduous, time-consuming process and is inconsistent in the context of yield and purity among institutions. Herein, we describe a microfluidic platform, which we have named MITEV (Microfluidic Isolation of Tumor-derived Extracellular Vesicles) for the rapid isolation of TEV from the plasma of pancreatic cancer patients. The device, which has ~100,000 pillars placed in a zigzag pattern and is coated with antibodies against generic EV surface proteins (anti-CD63, -CD9, and -CD81 antibodies) or the TEV specific anti-Epithelial Cell Adhesion Molecule (EpCAM) antibody, is capable of high-throughput EVs isolation and yields sufficient DNA (total of ~2–14 ng from 2-ml plasma) for downstream genomic analysis. Using two independent quantitative platforms, droplet digital polymerase chain reaction (ddPCR) and molecular barcoding using nanoString nCounter® technology, we can reliably identify KRAS mutations within isolated TEV of treatment-naïve metastatic pancreatic cancer patients. Our study suggests that the MITEV device can be used for point-of-care applications, such as in the context of monitoring residual or recurrent tumor presence in pancreatic cancer patients undergoing therapy.





Similar content being viewed by others
Abbreviations
- TEV:
-
Tumor associated Extracellular Vesicles
- NTEV:
-
Non-Tumor associated Extracellular Vesicles
- MITEV:
-
Microfluidic Isolation of Tumor-derived EVs
- PDAC:
-
Pancreatic Ductal Adenocarcinoma
- EV:
-
Extracellular Vesicles
- EpCAM:
-
Epithelial Cell Adhesion Molecule
- ddPCR:
-
Droplet digital Polymerase Chain Reaction
References
K. Allenson, J. Castillo, F.A. San Lucas, G. Scelo, D.U. Kim, V. Bernard, G. Davis, T. Kumar, M. Katz, M.J. Overman, L. Foretova, E. Fabianova, I. Holcatova, V. Janout, F. Meric-Bernstam, P. Gascoyne, I. Wistuba, G. Varadhachary, P. Brennan, S. Hanash, D. Li, A. Maitra, H. Alvarez, High prevalence of mutant KRAS in circulating exosome-derived DNA from early-stage pancreatic cancer patients. Ann. Oncol. 28, 741–747 (2017)
P. Bailey et al., Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 531, 47–52 (2016)
V. Bernard, D.U. Kim, F.A. San Lucas, J. Castillo, K. Allenson, F.C. Mulu, B.M. Stephens, J. Huang, A. Semaan, P.A. Guerrero, N. Kamyabi, J. Zhao, M.W. Hurd, E.J. Koay, C.M. Taniguchi, J.M. Herman, M. Javle, R. Wolff, M. Katz, G. Varadhachary, A. Maitra, H.A. Alvarez, Circulating nucleic acids are associated with outcomes of patients with pancreatic cancer. Gastroenterology 156, 108–118 e104 (2019)
A.V. Biankin et al., Pancreatic cancer genomes reveal aberrations in axon guidance pathway genes. Nature 491, 399–405 (2012)
J. Castillo et al., Surfaceome profiling enables isolation of cancer-specific exosomal cargo in liquid biopsies from pancreatic cancer patients. Ann. Oncol. 29, 223–229 (2018)
A. Chan, I. Prassas, A. Dimitromanolakis, R.E. Brand, S. Serra, E.P. Diamandis, I.M. Blasutig, Validation of biomarkers that complement CA19.9 in detecting early pancreatic cancer. Clin. Cancer Res. 20, 5787–5795 (2014)
C. Chen, J. Skog, C.H. Hsu, R.T. Lessard, L. Balaj, T. Wurdinger, B.S. Carter, X.O. Breakefield, M. Toner, D. Irimia, Microfluidic isolation and transcriptome analysis of serum microvesicles. Lab. Chip. 10, 505–511 (2010)
J.D. Cohen et al., Detection and localization of surgically resectable cancers with a multi-analyte blood test. Science 359, 926–930 (2018)
E. Crowley, F. Di Nicolantonio, F. Loupakis, A. Bardelli, Liquid biopsy: monitoring cancer-genetics in the blood. Nat. Rev. Clin. Oncol. 10, 472–484 (2013)
S. Gillen, T. Schuster, C. Meyer Zum Buschenfelde, H. Friess, J. Kleeff, Preoperative/neoadjuvant therapy in pancreatic cancer: a systematic review and meta-analysis of response and resection percentages. PLoS Med 7, e1000267 (2010)
R. Grant, E. Ansa-Addo, D. Stratton, S. Antwi-Baffour, S. Jorfi, S. Kholia, L. Krige, S. Lange, J. Inal, A filtration-based protocol to isolate human plasma membrane-derived vesicles and exosomes from blood plasma. J. Immunol. Methods 371, 143–151 (2011)
C.L. Hisey, K.D.P. Dorayappan, D.E. Cohn, K. Selvendiran, D.J. Hansford, Microfluidic affinity separation chip for selective capture and release of label-free ovarian cancer exosomes. Lab Chip 18, 3144–3153 (2018)
C. Ionescu-Tirgoviste, P.A. Gagniuc, E. Gubceac, L. Mardare, I. Popescu, S. Dima, M. Militaru, A 3D map of the islet routes throughout the healthy human pancreas. Sci. Rep. 5, 14634 (2015)
D.K. Jeppesen et al., Reassessment of exosome composition. Cell 177, 428–445 e418 (2019)
T. Kamisawa, L.D. Wood, T. Itoi, K. Takaori, Pancreatic cancer. Lancet 388, 73–85 (2016)
N. Kamyabi, V. Bernard, A. Maitra, Liquid biopsies in pancreatic cancer. Expert. Rev. Anticancer. Ther. 19, 869–878 (2019)
N. Kamyabi, J. Huang, J.J. Lee, V. Bernard, A. Semaan, B. Stephens, M.W. Hurd, S.A. Vanapalli, A. Maitra, P.A. Guerrero, A microfluidic device for label-free isolation of tumor cell clusters from unprocessed blood samples. Biomicrofluidics 13, 044111 (2019)
S.S. Kanwar, C.J. Dunlay, D.M. Simeone, S. Nagrath, Microfluidic device (ExoChip) for on-chip isolation, quantification and characterization of circulating exosomes. Lab Chip 14, 1891–1900 (2014)
J. Kleeff et al., Pancreatic cancer. Nat Rev Dis Primers 2, 16022 (2016)
J. Ko et al., Combining machine learning and nanofluidic technology to diagnose pancreatic cancer using exosomes. ACS Nano 11, 11182–11193 (2017)
J.M. Lewis et al., Integrated analysis of Exosomal protein biomarkers on alternating current electrokinetic chips enables rapid detection of pancreatic cancer in patient blood. ACS Nano 12, 3311–3320 (2018)
F. Liu, O. Vermesh, V. Mani, T.J. Ge, S.J. Madsen, A. Sabour, E.C. Hsu, G. Gowrishankar, M. Kanada, J.V. Jokerst, R.G. Sierra, E. Chang, K. Lau, K. Sridhar, A. Bermudez, S.J. Pitteri, T. Stoyanova, R. Sinclair, VS Nair, S.S. Gambhir, U. Demirci, The exosome Total isolation chip. ACS Nano 11, 10712–10723 (2017)
E. Pariset, V. Agache, A. Millet, Extracellular vesicles: isolation methods. Adv. Biosyst. 1, 1700040 (2017)
E. Reátegui, K.E. van der Vos, C.P. Lai, Zeinali, N.A. Atai, B. Aldikacti, F.P. Floyd Jr, A. H Khankhel, V. Thapar, F.H. Hochberg, L.V. Sequist, B.V. Nahed, B. S Carter, M. Toner, L. Balaj, D. T Ting, X.O. Breakefield, S.L. Stott, Engineered nanointerfaces for microfluidic isolation and molecular profiling of tumor-specific extracellular vesicles. Nat. Commun. 9, 175 (2018)
M. Sausen et al., Clinical implications of genomic alterations in the tumour and circulation of pancreatic cancer patients. Nat. Commun. 6, 7686 (2015)
S. Tang et al., Usefulness of 18F-FDG PET, combined FDG-PET/CT and EUS in diagnosing primary pancreatic carcinoma: a meta-analysis. Eur. J. Radiol. 78, 142–150 (2011)
K. Tjensvoll et al., Clinical relevance of circulating KRAS mutated DNA in plasma from patients with advanced pancreatic cancer. Mol. Oncol. 10, 635–643 (2016)
H.K. Woo, V. Sunkara, J. Park, T.H. Kim, J.R. Han, C.J. Kim, H.I. Choi, Y.K. Kim, Y.K. Cho, Exodisc for rapid, size-selective, and efficient isolation and analysis of nanoscale extracellular vesicles from biological samples. ACS Nano 11, 1360–1370 (2017)
M. Wu, Y. Ouyang, Z. Wang, R. Zhang, P.H. Huang, C. Chen, H. Li, P. Li, D. Quinn, M. Dao, S. Suresh, Y. Sadovsky, T.J. Huang, Isolation of exosomes from whole blood by integrating acoustics and microfluidics. Proc. Natl. Acad. Sci. U. S. A. 114, 10584–10589 (2017)
Y. Xia, G.M. Whitesides, Soft lithography. Annu. Rev. Mater. Sci. 28, 153–184 (1998)
R. Xu et al., Extracellular vesicles in cancer - implications for future improvements in cancer care. Nat. Rev. Clin. Oncol. 15, 617–638. (2018)
L. Zhang et al., Microenvironment-induced PTEN loss by exosomal microRNA primes brain metastasis outgrowth. Nature 527, 100–104 (2015)
Acknowledgements
Scanning electron microscopy was performed at the High-Resolution Electron Microscopy Facility, while nanoString analysis was performed at the Sequencing and Microarray facility (SMF) in MD Anderson Cancer Center (Houston, TX), both of which are supported by the NCI Cancer Center Support Grant (P30CA016672). N.K. was supported by the CPRIT Research Training Program (RP170067).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
A.M. discloses receiving royalties from Hangzhou Guangkeande (Cosmos) Biotechnology Company LTD for being a co-inventor on a license related to pancreatic cancer early detection, and this financial relationship is managed by the MD Anderson Conflict of Interest Committee. There are no other conflicts of interests to declare.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 281 kb)
Rights and permissions
About this article
Cite this article
Kamyabi, N., Abbasgholizadeh, R., Maitra, A. et al. Isolation and mutational assessment of pancreatic cancer extracellular vesicles using a microfluidic platform. Biomed Microdevices 22, 23 (2020). https://doi.org/10.1007/s10544-020-00483-7
Published:
DOI: https://doi.org/10.1007/s10544-020-00483-7