Abstract
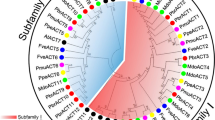
Actin is the most abundant protein in eukaryotic cells and is a key cytoskeletal component controlling cell morphology and motility. In this study, four MiACT genes were isolated from mango by homological cloning and designated as MiACT1, MiACT4, MiACT7, and MiACT9. Sequence alignments and phylogenetic analysis demonstrated that the four MiACT genes of mango were highly similar to each other at the nucleotide and amino acid levels. All of four MiACT proteins showed high similarity to the known actin proteins from other species. Reverse transcription polymerase chain reaction revealed that the four MiACT genes were constitutively and stably expressed in all organs tested. Application of plant growth regulators and four stress treatments had a remarkable effect on the expression of MiACT4, MiACT7 and MiACT9, whereas expression of MiACT1 was unresponsive. In contrast, the expression profiles of the four MiACT genes were not regulated by diurnal rhythms. Moreover, the expression of MiACT1 was not affected by heavy metal treatments and the transcript level of MiACT1 was rather stable in different days during the post-harvest period either under treatment or not. Our results suggest that the four actin genes play important roles throughout the entire life cycle of mango; the constitutively and stably expressed MiACT1 is the best candidate as an internal standard for differential gene expression analysis in mango.
Similar content being viewed by others
Abbreviations
- ABA:
-
abscisic acid
- BA:
-
benzyladenine
- DAF:
-
days after full bloom
- f-flower:
-
full flowering period
- GA3 :
-
gibberellic acid
- IAA:
-
indole-3-acetic acid
- l-flower:
-
late flowering period
- PP:
-
paclobutrazol
- RT-PCR:
-
reverse transcription polymerase chain reaction
References
An, Y.Q., Huang, S., McDowell, J.M., McKinney, E.C., Meagher, R.B.: Conserved expression of the Arabidopsis ACT1 and ACT3 actin subclass in organ primordia nad mature pollen. — Plant Cell 8: 15–30, 1996a.
An, Y.Q., McDowell, J.M., Huang, S., McKinney, E.C., Chambliss, S., Meagher, R.B.: Strong, constitutive expression of the Arabidopsis ACT2/ACT8 actin subclass in vegetative tissues. — Plant J. 10: 107–21, 1996b.
Anwar, R., Malik, A.U.: Hot water treatment affects ripening quality and storage life of mango (Mangifera indica L.). — Pak. J. agr. Sci, 44: 304–311, 2007.
Baird, W.V., Meagher, R.B.: A complex gene superfamily encodes actin in petunia. — EMBO J. 6: 3223–3231, 1987.
Dean, J.D., Goodwin, P.H., Hsiang, T.: Comparison of relative RT-PCR and Northern blot analyses to measure expression of β-l,3-glucanase in Nicotiana benthamiana infected with Colltotrichum destructivum. — Plant mol. Biol. Rep. 20: 347–356, 2002.
Deng, Z., Liu, X.H., Chen, C.L., Tian, W.M., Xia, Z.H., Li, D.J.: Molecular cloning and characterization of an actindepolymerizing factor gene in Hevea brasiliensis. — Afr. J. Biotechnol. 9:7603–7610, 2010.
Drouin, G., Dover, G.A.: Independent gene evolution in the potato actin gene family demonstrated by phylogenetic procedures for resolving gene conversions and the phylogeny of angiosperm actin genes. — J. mo1. Evol. 31: 132–150, 1990.
Huang, S., An, Y.Q., McDowell, J.M., McKinney, E.C., Meagher, R.B.: The Arabidopsis thaliana ACT4/ACT12 actin gene subclass is strongly expressed throughout pollen development. — Plant J. 10: 189–202, 1996.
Kabsch, W., Vandekerckhove, J.: Structure and function of actin. — Annu. Rev. biophys. biomol. Struct. 21: 49–76, 1992.
Kakinuma, M., Courya, D.A., Inagaki, E., Itoha, S., Yoshiura, Y., Amanoa, H.: Isolation and characterization of a single copy actin gene from a sterile mutant of Ulva pertusa (Ulvales, Chlorophyta). — Gene 334: 145–155, 2004.
Kandasamy, M.K., McKinney, E.C., Meagher, R.B.: Functional nonequivalency of actin isovariants in Arabidopsis. — Mol. Biol. Cell 13: 251–261, 2002.
Li, X.B., Fan, X.P., Wang, X.L., Cai, L., Yang, W.C.: The cotton ACTIN1 gene is functionally expressed in fibers nad participates in fiber elongation. — Plant Cell 17: 859–75, 2005.
Licausi, F., Giorgi, F.M., Zenoni, S., Osti, F., Pezzotti, M., Perata, P.: Genomic and transcriptomic analysis of the AP2/ERF superfamily in Vitis vinifera. — BMC Genomics 11: 719–--, 2010.
McDowell, J.M., An, Y.Q., Huang, S., McKinney, E.C., Meagher, R.B.: The Arabidopsis ACT7 actin gene is expressed in rapidly developing tissues and responds to several external stimuli. — Plant Physiol. 111: 699–711, 1996a.
McDowell, J.M., Huang, S., McKinney, E.C., An, Y.Q., Meaghed, R.B.: Structure and evolution of the actin gene family in Arabidopsis thaliana. — Genetics 142: 587–602, 1996b.
McElroy, D., Rothenberg, M., Reece, K.S., Wu, R.: Characterization of the rice (Oryza safiva) actin gene family. — Plant mo1. Biol. l5: 257–268, 1990.
McKinney, E.C., Kandasamy, M.K., Meagher, R.B.: Arabidopsis contains ancient classes of differentially expressed actin-related protein genes. — Plant Physiol. 128: 997–1007, 2002.
McKinney, E.C., Meagher, R.B.: Members of the Arabidopsis actin gene family are widely dispersed in the genome. — Genetics 149: 663–675, 1998.
Meagher, R.B., McLean, B.G.: Diversity of plant actins. — Cell Motil. Cytoskel. 16: 164–166, 1990.
Meagher, R.B., Williamson, R.E.: The plant cytoskeleton. — In Meyerowitz, E., Somerville, C. (ed.): Arabidopsis. Pp. 1049–1084. Cold Spring Harbor Laboratory Press, Cold Spring Harbor 1994.
Nagao, R.T., Shah, D.M., Eckenrode, V.K., Meagher, R.B.: Multigene family of actin-related sequences isolated from soybean genomic library. — DNA 1: 1–9, 1981.
Peng, H., Cheng, H.Y., Yu, X.W., Shi, Q.H., Zhang, H., Li, J.G., Ma, H.: Molecular analysis of an actin gene, CarACT1, from chickpea (Cicer arietinum L.). — Mol. Biol. Rep. 37: 1081–1088, 2010.
Pollard, T.D., Cooper, J.A.: Actin and actin binding proteins. A critical evaluation of mechanisms and functions. — Annu. Rev. Biochem. 55: 987–1035, 1986.
Rubenstein, P.A.: The functional importance of multiple actin isoforms. — Bioessays 12: 309–315, 1990.
Tamura, K., Dudley, J., Nei, M., Kumar, S.: MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. — Mol. Biol. Evol. 24: 1596–1599, 2007.
Thangavelu, M., Belostotsky, D., Bevan, M.W., Flavell, R.B., Rogers, H.J., Lonsdale, D.M.: Partial characterization of the Nicotiana tabacum actin gene family: evidence for pollen specific expression of one of the gene family members. — Mol. gen. Genet. 240: 290–295, 1993.
Thompson, J.D., Gibson, T.J., Plewniak, F., Jeanmougin, F., Higgins, D.G.: The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. — Nucl. Acids Res. 25: 4876–4882, 1997.
Valentijn, K., Valentijn, J.A., Jamieson, J.D.: Role of actin in regulated exocytosis and compensatory membrane retrieval: insights from an old acquaintance. — Biochem. biophys. Res. Commun. 266: 652–661, 1999.
Xiao, J.N., Huang, X.L., Li, Y.L., Huang, X.L., Li, X.J.: RNA extraction from cotyledon of mango with high levels of secondary substances and carbohydrates. — China Biotechnol. 23: 53–56, 2003.
Yang, Y.F., Wu, J., Zhu, K., Liu, L.Q., Chen, F.D., Yu, D.Y.: Identification and characterization of two chrysanthemum (Dendronthema moriforlium) DREB genes, belonging to the AP2/EREBP family. — Mol. biol. Rep. 36: 71–81, 2009.
Zhu, Q., Zhang, J.T., Gao, X.S., Tong, J.H., Xiao, L.T., Li, W.B., Zhang, H.X.: The Arabidopsis AP2/ERF transcription factor RAP2.6 participates in ABA, salt and osmotic stress responses. — Gene 457: 1–12, 2010.
Author information
Authors and Affiliations
Corresponding author
Additional information
Acknowledgments: This research was supported by Guangxi Natural Science Foundation (2011GXNSFA018115); Program for Excellent Talents in Guangxi Higher Education Institutions (201065); Innovation Project of Guangxi Postgraduate Education (105931001011), and the Scientific Research Foundation of Guangxi University (Grant No. 20090042).
Rights and permissions
About this article
Cite this article
Luo, C., He, X.H., Chen, H. et al. Molecular cloning and expression analysis of four actin genes (MiACT) from mango. Biol Plant 57, 238–244 (2013). https://doi.org/10.1007/s10535-012-0278-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10535-012-0278-9