Abstract
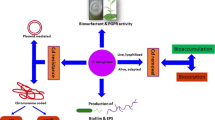
The 16S rRNA sequence and biochemical characteristics revealed the isolated organism as Pseudomonas sp. SU-EBT. This strain showed 97 and 90% decolorization of a recalcitrant dye, Congo red (100 mg l−1) and textile industry effluent with 50% reduction in COD within 12 and 60 h, respectively. The optimum pH and temperature for the decolorization was 8.0 and 40°C, respectively. Pseudomonas sp. SU-EBT was found to tolerate the dye concentration up to 1.0 g l−1. Significant induction in the activity of intracellular laccase suggested its involvement in the decolorization of Congo red. The metabolites formed after decolorization of Congo red, such as p-dihydroxy biphenyl, 8-amino naphthol 3-sulfonic acid and 3-hydroperoxy 8-nitrosonaphthol were characterized using FTIR and GC–MS. Phytotoxicity study revealed nontoxic nature of the degradation metabolites to Sorghum bicolor, Vigna radiata, Lens culinaris and Oryza sativa plants as compared to Congo red and textile industry effluent. Pseudomonas sp. SU-EBT decolorized several individual textile dyes, dye mixtures and textile industry effluent, thus it is a useful strain for the development of effluent treatment methods in textile processing industries.






Similar content being viewed by others
References
Amitaia G, Adania R, Sod-Moriah G, Rabinovitz I, Vincze A, Leader H, Chefetz B, Leibovitz-Persky L, Friesem D, Hear Y (1998) Oxidative biodegradation of phosphorothiolates by fungal lacccase. FEBS Lett 438:195–200. doi:10.1016/S0014-5793(98)01300-3
APHA-AWWA-WEF (2005) Standard methods for the examination of water and wastewater, 21st edn. American Public Health Association, Washington, DC
Blanquez P, Casas N, Gabarell FX, Sarra M, Caminal G, Vincent T (2004) Mechanism of textile metal dye biotransformation by Trametes versicolor. Water Res 38:2166–2172. doi:10.1016/j.watres.2004.01.019
Chen H, Hopper SL, Cerniglia CE (2005) Biochemical and molecular characterization of an azoreductase from Staphylococcus aureus, a tetrameric NADPH-dependent flavoprotein. Microbiology 151:1433–1441. doi:10.1099/mic.0.27805-0
Chivukula M, Renganathan V (1995) Phenolic azo dye oxidation by laccase from Pyricularia oryzae. Appl Environ Microbiol 61:4374–4377
Constapel M, Schellenträger M, Marzinkowski JM, Gab S (2009) Degradation of reactive dyes in wastewater from the textile industry by ozone: analysis of the products by accurate masses. Water Res 43:733–743. doi:10.1016/j.watres.2008.11.006
Daneshvar N, Ayazloo M, Khataee AR, Pourhassan M (2007) Biological decolorization of dye solution containing malachite green by microalgae Cosmarium sp. Bioresour Technol 98:1176–1182. doi:10.1016/j.biortech.2006.05.025
Dhanve RS, Kalyani DC, Phugare SS, Jadhav JP (2009) Coordinate action of exiguobacterial oxidoreductive enzymes in biodegradation of reactive yellow 84A dye. Biodegradation 20:245–255. doi:10.1007/s10532-008-9217-z
Faraco V, Pezzella C, Miele A, Giardina P, Sannia G (2009) Bio-remediation of colored industrial wastewaters by the white-rot fungi Phanerochaete chrysosporium and Pleurotus ostreatus and their enzymes. Biodegradation 20:209–220. doi:10.1007/s10532-008-9214-2
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Figueroa S, Vázqueza L, Alvarez-Gallegos A (2009) Decolorizing textile wastewater with Fenton’s reagent electrogenerated with a solar photovoltaic cell. Water Res 43:283–294. doi:10.1016/j.watres.2008.10.014
Hsueh CC, Chen BY (2007) Comparative study on reaction selectivity of azo dye decolorization by Pseudomonas luteola. J Hazard Mater 141:842–849. doi:10.1016/j.jhazmat.2006.07.056
Jadhav JP, Govindwar SP (2006) Biotransformation of Malachite green by Saccharomyces cerevisiae MTCC 463. Yeast 23:315–323. doi:10.1002/yea.1356
Jang MS, Jung BG, Sung NC, Lee YC (2007) Decolorization of textile plant effluent by Citrobacter sp. strain KCTC 18061P. J Gen Appl Microbiol 53:339–343. doi:10.2323/jgam.53.339
Kalme SD, Parshetti GK, Jadhav SU, Govindwar SP (2007) Biodegradation of benzidine based dye direct blue-6 by Pseudomonas desmolyticum NCIM 2112. Bioresour Technol 98:1405–1410. doi:10.1016/j.biortech.2006.05.023
Kalyani DC, Telke AA, Dhanve RS, Jadhav JP (2009) Ecofriendly biodegradation and detoxification of Reactive Red 2 textile dye by newly isolated Pseudomonas sp. SUK1. J Hazard Mater 163:735–742. doi:10.1016/j.jhazmat.2008.07.020
Kaushik P, Malik A (2009) Fungal dye decolorization: recent advances and future potential. Environ Int 35:127–141. doi:10.1016/j.envint.2008.05.010
Lowry OH, Rosebrough NJ, Farr AL, Randall RL (1951) Protein measurement with the folin phenol reagent. J Biol Chem 193:265–275
Mane UV, Gurav PN, Deshmukh AM, Govindwar SP (2008) Degradation of textile dye reactive navy blue RX (Reactive blue–59) by an isolated Actinomycete Streptomyces krainskii SUK-5. Malays J Microbiol 4:1–5
Mathur N, Bhatnagar P, Bakre P (2005) Assessing mutagenicity of textile dyes from Pali (Rajasthan) using ames bioassay. Appl Ecol Environ Res 4:111–118
Meehan C, Banal IM, McMullan G, Nigam P, Smyth F, Marchant R (2000) Decolorization of Remazol Black-B using a thermotolerant yeast, Kluyveromyces marxianus IMB3. Environ Int 26:75–79. doi:10.1016/S0160-4120(00)00084-2
Miller R, Kuglin J, Gallagher S, Flurkey WH (1997) A spectrophotometric assay for laccase using o-tolidine. J Food Biochem 21:445–459
Parshetti GK, Kalme SD, Saratale GD, Govindwar SP (2006) Biodegradation of malachite green by Kocuria rosea MTCC 1532. Acta Chim Slov 53:492–498
Saitou N, Nei M (1987) The neighbour-joining method: a new method for reconstruction phylogenetic trees. Mol Biol Evol 4:406–425
Salokhe MD, Govindwar SP (1999) Effect of carbon source on the biotransformation enzymes in Serratia marcescens. World J Microbiol Biotechnol 15:229–232. doi:10.1023/A:1008875404889
Sugiura W, Miyashita T, Yokoyama T, Arai M (1999) Isolation of azo dye degrading microorganisms and their application to white discharge printing of fabric. J Biosci Bioeng 88:577–581. doi:10.1016/S1389-1723(00)87680-X
Swamy J, Ramsay JA (1999) The evaluation of white rot fungi in the decolorization of textile dyes. Enzym Microb Technol 24:130–137. doi:0141-0229/99
Takezaki N, Rzhetsky A, Nei M (2004) Phylogenetic test of the molecular clock and linearized trees. Mol Biol Evol 12:823–833
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenies by using the neighbor joining method. PNAS 101:11034–11035. doi:10.1073/pnas.0404206101
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:598–599. doi:10.1093/molbev/msm092
Telke AA, Kalyani DC, Jadhav JP (2008) Govindwar SP. Kinetics and mechanism of reactive red 141 degradation by a bacterial isolate Rhizobium radiobacter MTCC 8161. Acta Chim Slov 55:320–329
Telke AA, Kalyani DC, Dawkar VV, Govindwar SP (2009) Influence of organic and inorganic compounds on oxidoreductive decolorization of sulfonated azo dye C.I. Reactive Orange 16. J Hazard Mater doi:10.1016/j.jhazmat.2009.07.008
Acknowledgments
One of the authors (Mr. Amar A. Telke) is thankful to the Council of Scientific and Industrial Research, New Delhi for financial assistance. Authors also thank the Common Facility Center, Shivaji University, Kolhapur, India for GC–MS facility.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Telke, A.A., Joshi, S.M., Jadhav, S.U. et al. Decolorization and detoxification of Congo red and textile industry effluent by an isolated bacterium Pseudomonas sp. SU-EBT. Biodegradation 21, 283–296 (2010). https://doi.org/10.1007/s10532-009-9300-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-009-9300-0