Abstract
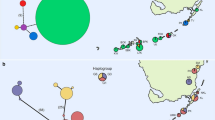
The brown anole, Anolis sagrei, is one of the most widespread and successful colonisers of the diverse Anolis genus, which comprises c. 400 species occurring naturally in Central and South America and the Caribbean. Based on extensive between and within population sampling from a previously published study (334 mitochondrial DNA sequences) and sampling for this study (37 mtDNA sequences), we reconstruct a phylogeny and produce a haplotype network to assign a recently introduced population in St Vincent, Lesser Antilles to its geographic origin. A single haplotype was present in the St Vincent population, which was identical to a haplotype from Tampa, FL. We show that genetic diversity within native range populations, combined with low frequencies of introduced haplotypes in native ranges, may impair attempts to identify source populations, even despite intensive sampling effort. The absence of mtDNA haplotype diversity suggests a significant genetic founder effect within the St Vincent population.



Similar content being viewed by others
References
Allendorf FW, Lundquist LL (2003) Population biology evolution and control of invasive species. Conserv Biol 17:24–30. doi:10.1046/j.1523-1739.2003.02365.x
Avise JC (1994) Molecular markers, natural history and evolution. Chapman and Hall, New York
Ballard JWO, Kreitman M (1995) Is mitochondrial DNA a strictly neutral marker? Trends Ecol Evol 10:485–488. doi:10.1016/S0169-5347(00)89195-8
Calsbeek R, Smith TB (2003) Ocean currents mediate evolution in island lizards. Nature 426:552–555. doi:10.1038/nature02143
Campbell TS (1996) Northern range expansion of the brown anole (Anolis sagrei) in Florida and Georgia. Herpetol Rev 27:155–157
Campbell TS (2003) The introduced brown anole (Anolis sagrei) occurs in every county in peninsular Florida. Herpetol Rev 34:173–174
Campbell TS, Echternacht AC (2003) Introduced species as moving targets: changes in body sizes of introduced lizards following experimental introductions and historical invasions. Biol Invasions 5:193–212. doi:10.1023/A:1026172314139
Carson HL (1990) Increased genetic variance after a population bottleneck. Trends Ecol Evol 5:228–230. doi:10.1016/0169-5347(90)90137-3
Clement M, Posada DA, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1659. doi:10.1046/j.1365-294x.2000.01020.x
Costa-Pierce BA (2003) Rapid evolution of an established feral tilapia (Oreochromis spp): the need to incorporate invasion science into regulatory structures. Biol Invasions 5:71–84. doi:10.1023/A:1024094606326
Durka W, Bossdorf O, Prati D et al (2005) Molecular evidence for multiple introductions of Garlic mustard (Alliaria petiolata, Brassicaceae) to North America. Mol Ecol 14:1697–1706. doi:10.1111/j.1365-294X.2005.02521.x
Felenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evol Int J Org Evol 39:783–791. doi:10.2307/2408678
Frankham R, Ballou JD, Briscoe DA (2002) Introduction to Conservation Genetics. Cambridge University Press, Cambridge
Glor RE, Gifford ME, Larson A et al (2004) Partial island submergence and speciation in an adaptive radiation: a multilocus analysis of the Cuban green anoles. Proc R Soc Lond B Biol Sci 271:2257–2265. doi:10.1098/rspb.2004.2819
Greene BT, Yorks DT, Parmerlee JS et al (2002) Discovery of Anolis sagrei in Grenada with comments on its potential impact on native anoles. Caribb J Sci 38:270–272
Hawley DM, Hanley D, Dhondt AA et al (2006) Molecular evidence for a founder effect in invasive house finch (Carpodacus mexicanus) populations experiencing an emergent disease epidemic. Mol Ecol 15:263–275. doi:10.1111/j.1365-294X.2005.02767.x
Huelsenbeck JP, Ronquist FR (2001) mrbayes: Bayesian inference in phylogeny. Bioinformatics 17:754–755. doi:10.1093/bioinformatics/17.8.754
Kolbe JJ, Glor RE, Rodrıguez Schettino L et al (2004) Genetic variation increases during biological invasion by a Cuban lizard. Nature 431:177–181. doi:10.1038/nature02807
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163. doi:10.1093/bib/5.2.150
Lee JC (1992) Anolis sagrei in Florida: phenetics of a colonising species I. meristic characters. Copeia 1985:182–194. doi:10.2307/1444808
Lee CE (2002) Evolutionary genetics of invasive species. Trends Ecol Evol 17:386–391. doi:10.1016/S0169-5347(02)02554-5
Macey JR, Larson A, Ananjeva NB et al (1997) Evolutionary shifts in three major structural features of the mitochondrial genome among Iguanian lizards. J Mol Evol 44:660–674. doi:10.1007/PL00006190
McDonald JH, Kreitman M (1991) Adaptive protein evolution at the adh locus in Drosophila. Nature 351:652–654. doi:10.1038/351652a0
McKeown S (1996) A field guide to reptiles and amphibians in the Hawaiian islands. Diamond Head Publishing Inc, Los Osos
Muirhead JR, Gray DK, Kelly D et al (2008) Identifying the source of species invasions: sampling intensity vs. genetic diversity. Mol Ecol 17:1020–1035. doi:10.1111/j.1365-294X.2008.03669.x
Norval G, Mao JJ, Chu HP et al (2002) A new record of an introduced species, the brown anole (Anolis sagrei) (Duméril & Bibron, 1837), in Taiwan. Zool Stud 41:332–336
Posada D, Crandall KA (1998) modeltest: testing the model of DNA substitution. Bioinformatics 14:817–818. doi:10.1093/bioinformatics/14.9.817
Puillandre N, Dupas S, Dangles O et al (2008) Genetic bottleneck in invasive species: the potato tuber moth adds to the list. Biol Invasions 10:319–333. doi:10.1007/s10530-007-9132-y
Sakai AK, Allendorf FW, Holt JS et al (2001) The population biology of invasive species. Annu Rev Ecol Syst 32:305–332. doi:10.1146/annurev.ecolsys.32.081501.114037
Schloesser DW, Nakepa TF, Mackie GL (1996) Zebra Mussel infestation of unionid bivalves (Unionidae) in North America. Am Zool 36:300–310
Schoener TW, Gorman GC (1968) Some niche differences among three species of lesser Antillean anoles. Ecology 49:819–830. doi:10.2307/1936533
Shigesada N, Kawasaki K (1997) Biological Invasions: theory and practice. Oxford University Press, Oxford
Stenson AG, Thorpe RS, Malhotra A (2004) Evolutionary differentiation of bimaculatus group anoles based on analyses of mtDNA and microsatellite data. Mol Phylogenet Evol 32:1–10. doi:10.1016/j.ympev.2003.12.008
Suarez AV, Tsutsui ND (2008) The evolutionary consequences of biological invasions. Mol Ecol 17:351–360. doi:101111/j1365-294X200703456x
Swofford DL (1998) Phylogenetic analysis using parsimony (* and other Methods). Sinauer Associates, Sunderland
Templeton AR, Crandall KA, Sing CF (1992) A cladistic analysis of phenotypic associations with haplotypes inferred from restriction endonuclease mapping and DNA sequence data III Cladogram estimation. Genetics 132:619–633
Williams EE (1969) The ecology of colonisation as seen in the zoogeography of anoline lizards on small islands. Q Rev Biol 44:345–389. doi:10.1086/406245
Acknowledgments
We would like to thank the St Vincent Forestry Department, in particular Fitzgerald Providence, for information and granting of permits necessary for field collection of material. Further thanks to Cathy Pook and Yann Surget-Groba for laboratory assistance, and Rob Ogden and an anonymous referee for valuable comments on earlier drafts of the manuscript. This work was funded by a NERC PhD studentship to JE, with CASE partner Wildlife DNA Services.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Eales, J., Thorpe, R.S. Revealing the geographic origin of an invasive lizard: the problem of native population genetic diversity. Biol Invasions 12, 77–86 (2010). https://doi.org/10.1007/s10530-009-9431-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-009-9431-6