Abstract
Objectives
To determine furfural biotransformation capabilities of Acinetobacter baylyi ADP1 and Acinetobacter schindleri ACE.
Results
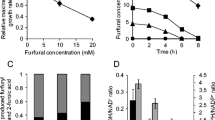
Acinetobacter baylyi ADP1 and A. schindleri ACE could not use furfural as sole carbon source but when acetate was used as substrate, ADP1 and ACE biotransformed 1 g furfural/l in 5 and 9 h, respectively. In both cases, the product of this biotransformation was difurfuryl-ether as shown by FT-IR and 1H and 13C NMR spectroscopy. The presence of furfural decreased the specific growth rate in acetate by 27% in ADP1 and 53% in ACE. For both strains, the MIC of furfural was 1.25 g/l. Nonetheless, ADP1 biotransformed 2 g furfural/l at a rate of 1 g/l/h in the stationary phase of growth. A transcriptional analysis of possible dehydrogenases involved in this biotransformation, identified that the areB and frmA genes were highly overexpressed after the exposure of ADP1 to furfural. The products of these genes are a benzyl-alcohol dehydrogenase and an alcohol dehydrogenase.
Conclusions
Acinetobacter baylyi ADP1 is a candidate for the biological detoxification of furfural, a fermentation inhibitor present in lignocellulosic hydrolysates, with the possible direct involvement of the AreB and FrmA enzymes in the process.
Access this article
We’re sorry, something doesn't seem to be working properly.
Please try refreshing the page. If that doesn't work, please contact support so we can address the problem.





Similar content being viewed by others
References
Bustin SA, Benes V, Garson JA et al (2009) The MIQE guidelines: minimum information for publication of quantitative Real-Time PCR experiments. Clin Chem 55:611–622. https://doi.org/10.1373/clinchem.2008.112797
Friedemann TE, Keegan PK, Witt NF (1964) Determination of furan aldehydes. Reaction with aniline in acetic and hydrochloric acid solutions. Anal Biochem 8:300–311. https://doi.org/10.1016/0003-2697(64)90061-2
Jones RM, Williams PA (2001) areCBA is an operon in Acinetobacter sp. strain ADP1 and is controlled by AreR, a 54-dependent regulator. J Bacteriol 183:405–409. https://doi.org/10.1128/JB.183.1.405-409.2001
Kannisto MS, Mangayil RK, Shrivastava-Bhattacharya A et al (2015) Metabolic engineering of Acinetobacter baylyi ADP1 for removal of Clostridium butyricum growth inhibitors produced from lignocellulosic hydrolysates. Biotechnol Biofuels 8:198. https://doi.org/10.1186/s13068-015-0389-6
Li Q, Metthew Lam LK, Xun L (2011) Cupriavidus necator JMP134 rapidly reduces furfural with a Zn-dependent alcohol dehydrogenase. Biodegradation 22:1215–1225. https://doi.org/10.1007/s10532-011-9476-y
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Miller EN, Jarboe LR, Turner PC et al (2009a) Furfural inhibits growth by limiting sulfur assimilation in ethanologenic Escherichia coli strain LY180. Appl Environ Microbiol 75:6132–6141. https://doi.org/10.1128/AEM.01187-09
Miller EN, Jarboe LR, Yomano LP et al (2009b) Silencing of NADPH-dependent oxidoreductase genes (yqhD and dkgA) in furfural-resistant ethanologenic Escherichia coli. Appl Environ Microbiol 75:4315–4323. https://doi.org/10.1128/AEM.00567-09
Miller EN, Turner PC, Jarboe LR, Ingram LO (2010) Genetic changes that increase 5-hydroxymethyl furfural resistance in ethanol-producing Escherichia coli LY180. Biotechnol Lett 32:661–667. https://doi.org/10.1007/s10529-010-0209-9
Mills TY, Sandoval NR, Gill RT (2009) Cellulosic hydrolysate toxicity and tolerance mechanisms in Escherichia coli. Biotechnol Biofuels 2:26. https://doi.org/10.1186/1754-6834-2-26
Sauer U, Canonaco F, Heri S et al (2004) The soluble and membrane-bound transhydrogenases UdhA and PntAB have divergent functions in NADPH metabolism of Escherichia coli. J Biol Chem 279:6613–6619. https://doi.org/10.1074/jbc.M311657200
Sigala JC, Quiroz L, Arteaga E et al (2019) Physiological and transcriptional comparison of acetate catabolism between Acinetobacter schindleri ACE and Escherichia coli JM101. FEMS Microbiol Lett 366:1–9. https://doi.org/10.1093/femsle/fnz151
Sigala JC, Suárez BP, Lara AR et al (2017) Genomic and physiological characterization of a laboratory-isolated Acinetobacter schindleri ACE strain that quickly and efficiently catabolizes acetate. Microbiology 163:1052–1064. https://doi.org/10.1099/mic.0.000488
Singh A, Bedore SR, Sharma NK et al (2019) Removal of aromatic inhibitors produced from lignocellulosic hydrolysates by Acinetobacter baylyi ADP1 with formation of ethanol by Kluyveromyces marxianus. Biotechnol Biofuels 12:91. https://doi.org/10.1186/s13068-019-1434-7
Tsuge Y, Kudou M, Kawaguchi H et al (2016) FudC, a protein primarily responsible for furfural detoxification in Corynebacterium glutamicum. Appl Microbiol Biotechnol 100:2685–2692. https://doi.org/10.1007/s00253-015-7115-y
Wang S, Cheng G, Joshua C et al (2016) Furfural tolerance and detoxification mechanism in Candida tropicalis. Biotechnol Biofuels 9:1–11. https://doi.org/10.1186/s13068-016-0668-x
Wierckx N, Koopman F, Ruijssenaars HJ, de Winde JH (2011) Microbial degradation of furanic compounds: biochemistry, genetics, and impact. Appl Microbiol Biotechnol 92:1095–1105. https://doi.org/10.1007/s00253-011-3632-5
Wolfe AJ (2005) The acetate switch. Microbiol Mol Biol Rev 69:12–50. https://doi.org/10.1128/MMBR.69.1.12-50.2005
Zhang Y, Ujor V, Wick M, Ezeji TC (2015) Identification, purification and characterization of furfural transforming enzymes from Clostridium beijerinckii NCIMB 8052. Anaerobe 33:124–131. https://doi.org/10.1016/j.anaerobe.2015.03.005
Acknowledgements
We thank technical support by Dr. Gabriel Vigueras Ramírez and Ernesto García García (Universidad Autónoma Metropolitana, México).
Funding
This study was funded by the Consejo Nacional de Ciencia y Tecnología (CONACyT) [CB-2012–01/183 813], and Programa Especial de Apoyo a la Investigación Básica 2019 Rectoría General Universidad Autónoma Metropolitana, to Sigala JC.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Arteaga, J.E., Cerros, K., Rivera-Becerril, E. et al. Furfural biotransformation in Acinetobacter baylyi ADP1 and Acinetobacter schindleri ACE. Biotechnol Lett 43, 1043–1050 (2021). https://doi.org/10.1007/s10529-021-03094-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-021-03094-1