Abstract
Objective
To inspect the feasibility of recombinant stable HEK293 cell lines development for biopharmaceuticals production using CRISPR/Cas9-mediated site-specific integration.
Results
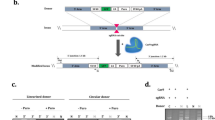
Using EGFP as a model protein, we first confirmed that the ‘safe harbor’ AAVS1 locus could be successfully targeted and the exogenous genes could be integrated through homology-directed repair induced by CRISPR/Cas9 technology. Then we constructed a donor plasmid harboring CTLA4Ig gene with an upstream CMV promoter and a downstream puromycin N-acetyltransferase gene to accelerate the efficient integration and selection of CTLA4Ig expression clones. After puromycin enrichment, the transfected pool was diluted for single clone selection, and 12 recombinant clones with CTLA4Ig expression were finally selected with a targeting efficiency of 25.8%. Productivity assay demonstrated that a frequency of 83.3% of selected clone were of consistent productivities, thus illustrating the high efficiency and success rate of this strategy.
Conclusions
CRISPR/Cas9 mediated site-specific integration is an efficient and reliable tool to establishment recombinant stable HEK293 cell lines for both academic and industrial applications.





Similar content being viewed by others
References
Barnes LM, Bentley CM, Dickson AJ (2003) Stability of protein production from recombinant mammalian cells. Biotechnol Bioeng 81:631–639. https://doi.org/10.1002/bit.10517
Barrangou R et al (2007) CRISPR provides acquired resistance against viruses in prokaryotes. Science 315:1709–1712
Bluestone JA, Clair EW, La SCE, Turka LA (2006) CTLA4Ig: bridging the basic immunology with clinical application. Immunity 24:233–238
Castano J et al (2017) Generation and characterization of a human iPSC cell line expressing inducible Cas9 in the “safe harbor” AAVS1 locus. Stem Cell Res 21:137–140. https://doi.org/10.1016/j.scr.2017.04.011
Charpentier M et al (2018) CtIP fusion to Cas9 enhances transgene integration by homology-dependent repair. Nat Commun 9:1133. https://doi.org/10.1038/s41467-018-03475-7
Cheng JK, Lewis AM, Kim DS, Dyess T, Alper HS (2016) Identifying and retargeting transcriptional hot spots in the human genome. Biotechnol J. https://doi.org/10.1002/biot.201600015
Cong L et al (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339:819–823. https://doi.org/10.1126/science.1231143
Cristea S, Freyvert Y, Santiago Y, Holmes MC, Urnov FD, Gregory PD, Cost GJ (2013) In vivo cleavage of transgene donors promotes nuclease-mediated targeted integration. Biotechnol Bioeng 110:871–880. https://doi.org/10.1002/bit.24733
Doench JG et al (2014) Rational design of highly active sgRNAs for CRISPR-Cas9-mediated gene inactivation. Nat Biotechnol 32:1262–1267. https://doi.org/10.1038/nbt.3026
Doench JG et al (2016) Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat Biotechnol 34:184–191. https://doi.org/10.1038/nbt.3437
Gaj T, Gersbach CA, Barbas CF 3rd (2013) ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol 31:397–405. https://doi.org/10.1016/j.tibtech.2013.04.004
Gratz SJ, Ukken FP, Rubinstein CD, Thiede G, Donohue LK, Cummings AM, O’Connor-Giles KM (2014) Highly specific and efficient CRISPR/Cas9-catalyzed homology-directed repair in drosophila. Genetics 196:961–971. https://doi.org/10.1534/genetics.113.160713
Hacker DL, Balasubramanian S (2016) Recombinant protein production from stable mammalian cell lines and pools. Curr Opin Struct Biol 38:129–136. https://doi.org/10.1016/j.sbi.2016.06.005
Hockemeyer D et al (2009) Efficient targeting of expressed and silent genes in human ESCs and iPSCs using zinc-finger nucleases. Nat Biotechnol 27:851–857. https://doi.org/10.1038/nbt.1562
Horvath P, Barrangou R (2010) CRISPR/Cas, the immune system of bacteria and archaea. Science 327:167–170. https://doi.org/10.1126/science.1179555
Irion U, Krauss J, Nusslein-Volhard C (2014) Precise and efficient genome editing in zebrafish using the CRISPR/Cas9 system. Development 141:4827–4830. https://doi.org/10.1242/dev.115584
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337:816–821. https://doi.org/10.1126/science.1225829
Joung JK, Sander JD (2012) TALENs: a widely applicable technology for targeted genome editing. Nat Rev Mol Cell Biol 14:49–55. https://doi.org/10.1038/nrm3486
Lee JS, Kallehauge TB, Pedersen LE, Kildegaard HF (2015) Site-specific integration in CHO cells mediated by CRISPR/Cas9 and homology-directed DNA repair pathway. Sci Rep 5:8572. https://doi.org/10.1038/srep08572
Liang XQ, Potter J, Kumar S, Ravinder N, Chesnut JD (2017) Enhanced CRISPR/Cas9-mediated precise genome editing by improved design and delivery of gRNA, Cas9 nuclease, and donor DNA. J Biotechnol 241:136–146. https://doi.org/10.1016/j.jbiotec.2016.11.011
Mali P et al (2013) RNA-guided human genome engineering via Cas9. Science 339:823–826. https://doi.org/10.1126/science.1232033
Moreno-Mateos MA, Vejnar CE, Beaudoin JD, Fernandez JP, Mis EK, Khokha MK, Giraldez AJ (2015) CRISPRscan: designing highly efficient sgRNAs for CRISPR-Cas9 targeting in vivo. Nat Methods 12:982–988. https://doi.org/10.1038/nmeth.3543
Noh SM, Sathyamurthy M, Lee GM (2013) Development of recombinant Chinese hamster ovary cell lines for therapeutic protein production. Curr Opin Chem Eng 2:391–397. https://doi.org/10.1016/j.coche.2013.08.002
Papapetrou EP et al (2011) Genomic safe harbors permit high beta-globin transgene expression in thalassemia induced pluripotent stem cells. Nat Biotechnol 29:73–78. https://doi.org/10.1038/nbt.1717
Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F (2013) Genome engineering using the CRISPR-Cas9 system. Nat Protoc 8:2281–2308. https://doi.org/10.1038/nprot.2013.143
Ronda C, Pedersen LE, Hansen HG, Kallehauge TB, Betenbaugh MJ, Nielsen AT, Kildegaard HF (2014) Accelerating genome editing in CHO cells using CRISPR Cas9 and CRISPy, a web-based target finding tool. Biotechnol Bioeng 111:1604–1616. https://doi.org/10.1002/bit.25233
Sakurai K et al (2010) Efficient integration of transgenes into a defined locus in human embryonic stem cells. Nucleic Acids Res 38:e96. https://doi.org/10.1093/nar/gkp1234
Smith JR et al (2008) Robust, persistent transgene expression in human embryonic stem cells is achieved with AAVS1-targeted integration. Stem Cells 26:496–504. https://doi.org/10.1634/stemcells.2007-0039
Sun T et al (2015) Functional knockout of FUT8 in Chinese hamster ovary cells using CRISPR/Cas9 to produce a defucosylated antibody. Eng Life Sci 15:660–666. https://doi.org/10.1002/elsc.201400218
Vincenti F, Dritselis A, Kirkpatrick P (2011) Belatacept. Nat Rev Drug Discov 10:655–656. https://doi.org/10.1038/nrd3536
Walsh RM, Hochedlinger K (2013) A variant CRISPR-Cas9 system adds versatility to genome engineering. Proc Natl Acad Sci USA 110:15514–15515. https://doi.org/10.1073/pnas.1314697110
Wang D, Quan Y, Yan Q, Morales JE, Wetsel RA (2015) Targeted disruption of the beta2-microglobulin gene minimizes the immunogenicity of human embryonic stem cells. Stem Cells Transl Med 4:1234–1245. https://doi.org/10.5966/sctm.2015-0049
Wilson C, Bellen HJ, Gehring WJ (1990) Position effects on eukaryotic gene expression. Annu Rev Cell Biol 6:679–714. https://doi.org/10.1146/annurev.cb.06.110190.003335
Wurm FM (2004) Production of recombinant protein therapeutics in cultivated mammalian cells. Nat Biotechnol 22:1393–1398. https://doi.org/10.1038/nbt1026
Wurm FM, Petropoulos CJ (1994) Plasmid integration, amplification and cytogenetics in CHO cells: questions and comments. Biologicals 22:95–102. https://doi.org/10.1006/biol.1994.1015
Zhang JP et al (2017) Efficient precise knockin with a double cut HDR donor after CRISPR/Cas9-mediated double-stranded DNA cleavage. Genome Biol 18:35. https://doi.org/10.1186/s13059-017-1164-8
Zhao M et al (2018) Rapid development of stable transgene CHO cell lines by CRISPR/Cas9-mediated site-specific integration into C12orf35. Appl Microbiol Biotechnol 102:6105–6117. https://doi.org/10.1007/s00253-018-9021-6
Zhou H, Liu ZG, Sun ZW, Huang Y, Yu WY (2010) Generation of stable cell lines by site-specific integration of transgenes into engineered Chinese hamster ovary strains using an FLP-FRT system. J Biotechnol 147:122–129. https://doi.org/10.1016/j.jbiotec.2010.03.020
Zhu J (2012) Mammalian cell protein expression for biopharmaceutical production. Biotechnol Adv 30:1158–1170. https://doi.org/10.1016/j.biotechadv.2011.08.022
Acknowledgements
This work was supported in part by the Science & Technology Commission of Shanghai Municipality (Grant Nos. 17ZR1413700 and 17431904500) and Medical and Engineering Cross Research Foundation of Shanghai Jiao Tong University (Grant No. YG2016QN27).
Supporting information
Supplementary Table S1—PCR primers sequences.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Authors declare no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals experiments.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yang, H., Wang, J., Zhao, M. et al. Feasible development of stable HEK293 clones by CRISPR/Cas9-mediated site-specific integration for biopharmaceuticals production. Biotechnol Lett 41, 941–950 (2019). https://doi.org/10.1007/s10529-019-02702-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-019-02702-5