Abstract
Objective
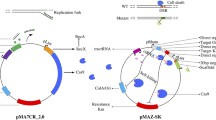
To construct a clustered, regularly interspaced, short palindromic repeats (CRISPR)/cas9 system and use this system to obtain a recombinant Escherichia coli strain possessing the fatty acid metabolism genes from a lipid-rich marine bacterium.
Results
The fatty acid regulatory transcription factor (fadR), delta9 (Δ9 desaturase) and acetyl-CoA carboxylase (acc) genes were cloned from Shewanella frigidimarina. The fatty acid regulatory transcription factor (fadD) and phosphoenolpyruvate carboxylase inactivated strains were used to construct the fadR/delta9 and acc knock-in strains, which are both markerless and “scar”-less, and identified the change in fatty acid composition in the recombinant strains. There was no change in fatty acid composition between the wild-type strain and recombinant strains. All strains had 11:0, 12:0, 13:0, 14:0, 15:0, 16:0, 17:1, 17:0 and 18:0 fatty acids, with 16:0 and 18:0 fatty acids being dominant. The total lipid content of each recombinant strain was higher than the wild-type strain, with a maximum of 13.1 %, nearly 5.3 % higher than wild-type strain.
Conclusion
The CRISPR/cas9 system, in conjunction with λ-Red recombinases, can rapidly and efficiently edit the E. coli genome. The CRISPR/cas9 recombineering machinery can be modified to select biotechnologically-relevant bacteria other than E. coli.


Similar content being viewed by others
References
Bozal N, Montes MJ, Tudela E, Jimenez F, Guinea J (2002) Shewanella frigidimarina and Shewanella livingstonensis sp. nov. isolated from Antarctic coastal areas. Int J Syst Evol Microbiol 52:195–205
Broekman JH, Steenbakkers JF (1973) Growth in high osmotic medium of an unsaturated fatty acid auxotroph of Escherichia coli K-12. J Bacteriol 116:285–289
Campbell JW, Cronan JE Jr (2001) Bacterial fatty acid biosynthesis: targets for antibacterial drug discovery. Annu Rev Microbiol 55:305–332
Deltcheva E et al (2011) CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471:602–607
DiRusso CC, Metzger AK, Heimert TL (1993) Regulation of transcription of genes required for fatty acid transport and unsaturated fatty acid biosynthesis in Escherichia coli by FadR. Mol Microbiol 7:311–322
Hsu PD, Lander ES, Zhang F (2014) Development and applications of CRISPR-Cas9 for genome engineering. Cell 157:1262–1278
Jiang W, Bikard D, Cox D, Zhang F, Marraffini LA (2013) RNA-guided editing of bacterial genomes using CRISPR-Cas systems. Nat Biotechnol 31:233–239
Jiang Y, Chen B, Duan C, Sun B, Yang J, Yang S (2015) Multigene editing in the Escherichia coli genome via the CRISPR-Cas9 system. Appl Environ Microbiol 81:2506–2514
Klein K, Steinberg R, Fiethen B, Overath P (1971) Fatty acid degradation in Escherichia coli. An inducible system for the uptake of fatty acids and further characterization of old mutants. Eur J Biochem 19:442–450
Lennen RM, Pfleger BF (2012) Engineering Escherichia coli to synthesize free fatty acids. Trends Biotechnol 30:659–667
Los DA, Murata N (1998) Structure and expression of fatty acid desaturases. Biochim Biophys Acta 1394:3–15
Magnuson K, Jackowski S, Rock CO, Cronan JE Jr (1993) Regulation of fatty acid biosynthesis in Escherichia coli. Microbiol Rev 57:522–542
Thomason L, Court DL, Bubunenko M, Costantino N, Wilson H, Datta S, Oppenheim A (2007) Recombineering: genetic engineering in bacteria using homologous recombination. Curr Protoc Mol Biol. doi:10.1002/0471142727.mb0116s78
Yu D, Ellis HM, Lee EC, Jenkins NA, Copeland NG, Court DL (2000) An efficient recombination system for chromosome engineering in Escherichia coli. Proc Natl Acad Sci USA 97:5978–5983
Zhang F, Ouellet M, Batth TS, Adams PD, Petzold CJ, Mukhopadhyay A, Keasling JD (2012) Enhancing fatty acid production by the expression of the regulatory transcription factor FadR. Metab Eng 14:653–660
Acknowledgments
This work was supported by the Basic Scientific Fund for National Public Research Institutes of China (Grant No. 2015T05), Marine Renewable Energy Funds Project (Grant No. 201305022 GHME2001SW02) and Taishan Scholar award. We sincerely thank Sheng Yang from Shanghai Institutes for Biological Sciences for kindly providing pTargetF and pCas.
Supporting information
Supplementary protocols—Total lipid extraction and FAME preparation.
Supplementary Table 1—Strains and plasmids used in this study.
Supplementary Table 2—Primers used in this study.
Supplementary Fig. 1—The TIC of FAME Mix and sample using GC–MS.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xia, J., Wang, L., Zhu, Jb. et al. Expression of Shewanella frigidimarina fatty acid metabolic genes in E. coli by CRISPR/cas9-coupled lambda Red recombineering. Biotechnol Lett 38, 117–122 (2016). https://doi.org/10.1007/s10529-015-1956-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-015-1956-4