Abstract
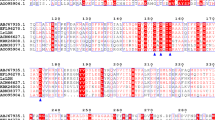
The gene coding for d-lactate dehydrogenase (d-LDH) from Pediococcus acidilactici DSM 20284 was cloned and expressed in E. coli. The recombinant enzyme was purified by nickel-affinity chromatography. It converted phenylpyruvic acid (PPA) to 3-phenyllactic acid maximally at 30°C and pH 5.5 with a specific activity of 140 and 422 U/mg for PPA and pyruvate, respectively. The K m, turnover number (k cat), and catalytic efficiency (k cat/K m) for PPA were 2.9 mM, 305 s−1, and 105 mM−1 s−1, respectively.



Similar content being viewed by others
References
Dieuleveux V, Lemarinier S, Gueguen M (1998a) Antimicrobial spectrum and target site of d-3-phenyllactic acid. Int J Food Microbiol 40:177–183
Dieuleveux V, Van Der Pyl D, Chataud J, Gueguen M (1998b) Purification and characterization of anti-Listeria compounds produced by Geotrichum candidum. Appl Environ Microbiol 64:800–803
Hummel W, Schütte H, Kula M (1983) Large scale production of d-lactate dehydrogenase for the stereospecific reduction of pyruvate and phenyipyruvate. Eur J Appl Microbiol Biotechnol 18:75–85
Jia J, Mu W, Zhang T, Jiang B (2010) Bioconversion of phenylpyruvate to phenyllactate: gene cloning, expression, and enzymatic characterization of d- and L1-lactate dehydrogenases from Lactobacillus plantarum SK002. Appl Biochem Biotechnol 162:242–251
Lavermicocca P, Valerio F, Evidente A, Lazzaroni S, Corsetti A, Gobbetti M (2000) Purification and characterization of novel antifungal compounds from the sourdough Lactobacillus plantarum strain 21B. Appl Environ Microbiol 66:4084–4090
Li X, Jiang B, Pan B, Mu W, Zhang T (2008) Purification and partial characterization of Lactobacillus species SK007 lactate dehydrogenase (LDH) catalyzing phenylpyruvic acid (PPA) conversion into phenyllactic acid (PLA). J Agric Food Chem 56:2392–2399
Ohhira I, Kuwaki S, Morita H, Suzuki T, Tomita S, Hisamatsu S, Sonoki S, Shinoda S (2004) Identification of 3-phenyllactic acid as a possible antibacterial substance produced by Enterococcus faecalis TH10. Biocontrol Sci 9:77–81
Prema P, Smila D, Palavesam A, Immanuel G (2010) Production and characterization of an antifungal compound (3-phenyllactic acid) produced by Lactobacillus plantarum strain. Food Bioprocess Technol 3:379–386
Schnürer J, Magnusson J (2005) Antifungal lactic acid bacteria as biopreservatives. Trends Food Sci Technol 16:70–78
Taguchi H, Ohta T (1991) d-Lactate dehydrogenase is a member of the d-isomer-specific 2-hydroxyacid dehydrogenase family. Cloning, sequencing, and expression in Escherichia coli of the d-lactate dehydrogenase gene of Lactobacillus plantarum. J Biol Chem 266:12588–12594
Thierry A, Maillard MB (2002) Production of cheese flavour compounds derived from amino acid catabolism by Propionibacterium freudenreichii. Lait 82:17–32
Tokuda C, Ishikura Y, Shigematsu M, Mutoh H, Tsuzuki S, Nakahira Y, Tamura Y, Shinoda T, Arai K, Takahashi O, Taguchi H (2003) Conversion of Lactobacillus pentosus d-lactate dehydrogenase to a d-hydroxyisocaproate dehydrogenase through a single amino acid replacement. J Bacteriol 185:5023–5026
Zheng Z, Ma C, Gao C, Li F, Qin J, Zhang H, Wang K, Xu P (2011) Efficient conversion of phenylpyruvic acid to phenyllactic acid by using whole cells of Bacillus coagulans SDM. PLoS ONE. doi:10.1371/journal.pone.0019030
Acknowledgments
This work was supported by the 973 Project (No. 2012CB720802), NSFC Project (No. 31171705 and 20906040), the 863 Project (No. 2011AA100904), the Support Project of Jiangsu Province (No. BE2010678 and BE2010626), and the Open Project Program of State Key Laboratory of Food Science and Technology, Jiangnan University (No. SKLF-KF-201105).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mu, W., Yu, S., Jiang, B. et al. Characterization of d-lactate dehydrogenase from Pediococcus acidilactici that converts phenylpyruvic acid into phenyllactic acid. Biotechnol Lett 34, 907–911 (2012). https://doi.org/10.1007/s10529-012-0847-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-012-0847-1