Abstract
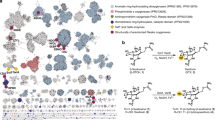
The epoxide hydrolase (EH) of a marine fish, Mugil cephalus, was engineered to improve the catalytic activity based on comparative homology modeling. The 3-D crystal structure of the EH from Aspergillus niger was used as a template. A triple point mutant, F193Y for spatial orientation of the nucleophile (D199), W200L for removing electron density overlap between W200 and Y348, and E378D for good charge relay in the active site, was developed. The initial hydrolysis rate, the reaction time to reach 98 %ee, and yield were enhanced up to 35-fold, 26-fold and 32%, respectively, by homology modeling-inspired site-directed mutagenesis of M. cephalus EH.





Similar content being viewed by others
References
Arand M, Muller F, Mecky A, Hinz W, Urban P, Pompon D, Kellner R, Oesch F (1999) Catalytic triad of microsomal epoxide hydrolase: replacement of Glu404 with Asp leads to a strongly increased turnover rate. Biochem J 337:37–43
Archelas A, Furstoss R (2001) Synthetic applications of epoxide hydrolases. Curr Opin Chem Biol 5:112–119
Barth S, Fischer M, Schmid RD, Pleiss J (2004) The database of epoxide hydrolase and haloalkane dehalogenases: one structure, many functions. Bioinformatics 20:2845–2847
Cao L, Lee J, Chen JW, Wood TK (2006) Enantioconvergent production of (R)-1-phenyl-1,2-ethanediol from styrene oxide by combining the Solanum tuberosum and an evolved Agrobacterium radiobacter AD1 epoxide hydrolases. Biotechnol Bioeng 94:522–529
Cedrone F, Niel S, Ait-abdelkader N, Torre C, Krumm H, Roca S, Bhatnagar T, Maichele A, Reetz MT, Baratti JC (2003) Directed evolution of the epoxide hydrolase from Aspergillus niger. Biocatal Biotransformation 21:357–364
Hwang S, Choi CY, Lee EY (2008a) One-pot biotransformation of racemic styrene oxide into (R)-1,2-phenylethandiol by two recombinant microbial epoxide hydrolases. Biotechnol Bioprocess Eng 13:274–278
Hwang Y-O, Kang SG, Woo J-H, Kwon KK, Sato T, Lee EY, Han MS, Kim S-J (2008b) Screening enantioselective epoxide hydrolase activities from marine microorganisms: detection of activities in Erythrobacter spp. Mar Biotechnol 10:366–373
Karboune S, Archelas A, Baratti J (2006) Properties of epoxide hydrolase from Aspergillus niger for the hydrolytic kinetic resolution of epoxides in pure organic media. Enzyme Microb Technol 39:318–324
Kim HS, Lee SJ, Lee EY (2006) Development and characterization of recombinant whole-cell biocatalysts expressing epoxide hydrolase from Rhodotorula glutinis for enantioselective resolution of racemic epoxides. J Mol Catal B 43:2–8
Kim HS, Lee OK, Hwang S, Kim BJ, Lee EY (2008) Biosynthesis of (R)-phenyl-1,2-ethanediol from racemic styrene oxide by using bacterial and marine fish epoxide hydrolases. Biotechnol Lett 30:127–133
Kronenburg NAE, de Bont JAM (2001) Effects of detergents on specific activity and enantioselectivity of the epoxide hydrolase from Rhodotorula glutinis. Enzyme Microb Technol 28:210–217
Lee EY (2007) Enantioselective hydrolysis of epichlorohydrin in organic solvents using recombinant epoxide hydrolase. J Ind Eng Chem 13:159–162
Lee EY (2008) Epoxide hydrolase-mediated enantioconvergent bioconversions to prepare chiral epoxides and alcohols. Biotechnol Lett 30:1509–1514
Lee EY, Shuler ML (2007) Molecular engineering of epoxide hydrolase and its application to asymmetric and enantioconvergent hydrolysis. Biotechnol Bioeng 98:318–327
Lee SJ, Kim HS, Kim SJ, Park S, Kim BJ, Shuler ML, Lee EY (2007) Cloning, expression and enantioselective hydrolytic catalysis of a microsomal epoxide hydrolase from a marine fish, Mugil cephalus. Biotechnol Lett 29:237–246
Nardini M, Ridder IS, Rozeboom HJ, Kalk KH, Rink R, Janssen DB, Dijkstra BW (1999) The X-ray structure of epoxide hydrolase from Agrobacterium radiobacter AD1. J Biol Chem 274:14579–14596
Reetz MT (2006) Directed evolution of enantioselective enzymes as catalysts for organic synthesis. Adv Catal 49:1–69
Reetz MT, Torre C, Eipper A, Lohmer R, Hermes M, Brunner B, Maichele A, Bocola M, Arand M, Cronin A, Genzel Y, Archelas A, Furstoss R (2004) Enhancing the enantioselectivity of an epoxide hydrolase by directed evolution. Org Lett 6:177–180
Rink R, Spelberg JHL, Pieters RJ, Kingma J, Nardini M, Kellogg RM, Dijkstra BW, Janssen DB (1999) Mutation of tyrosine residues involved in the alkylation half reaction of epoxide hydrolase from Agrobacterium radiobacter AD1 results in improved enantioselectivity. J Am Chem Soc 121:7417–7418
Rui L, Cao L, Chen W, Reardon KF, Wood TK (2004) Active site engineering of the epoxide hydrolase from Agrobacterium radiobacter AD1 to enhance aerobic mineralization of cis-1,2-dichloroethylene in cells expressing an evolved toluene ortho-monooxygenase. J Biol Chem 279:46810–46817
Rui L, Cao L, Chen W, Reardon KF, Wood TK (2005) Protein engineering of epoxide hydrolase from Agrobacterium radiobacter AD1 for enhanced activity and enantioselective production of (R)-1-phenylethane-1,2-diol. Appl Environ Microbiol 71:3995–4003
Schrader W, Eipper A, Pugh Dj, Reetz MT (2002) Second-generation MS-based high-throughput screening system for enantioselective catalysis and biocatalysis. Can J Chem 80:626–632
Strauss UT, Felfer U, Faber K (1999) Biocatalytic transformation of racemates into chiral building blocks in 100% chemical yield and 100% enantiomeric excess. Tetrahedron 10:107–117
van Loo B, Spelberg JHL, Kingma J, Sonke T, Wubbolts MG, Janssen DB (2004) Directed evolution of epoxide hydrolase from A. radiobacter toward higher enantioselectivity by error-prone PCR and DNA shuffling. Chem Biol 11:981–990
Zou J, Hallberg BM, Bergfors T, Oesch F, Arand M, Mowbray SL, Jones TA (2000) Structure of Aspergillus niger epoxide hydrolase at 1.8 Å resolution: implication for the structure and function of the mammalian microsomal class of epoxide hydrolases. Structure 8:111–122
Acknowledgments
This work was supported by the Marine and Extreme Genome Research Center Program, Ministry of Land, Transportation and Maritime Affairs, Republic of Korea. The stipend for S. H. Choi was partially supported by the Ministry of Knowledge Economy (MKE) and Korea Industrial Technology Foundation (KOTEF) through the Human Resource Training Project for Strategic Technology.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Choi, S.H., Kim, H.S. & Lee, E.Y. Comparative homology modeling-inspired protein engineering for improvement of catalytic activity of Mugil cephalus epoxide hydrolase. Biotechnol Lett 31, 1617–1624 (2009). https://doi.org/10.1007/s10529-009-0055-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-009-0055-9