Abstract
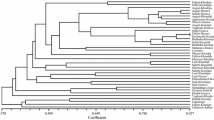
Northeastern Turkey is recognized as one of the most important germplasm centers for the grape in the world. In the present study, simple sequence repeat markers were used to investigate the genetic diversity between four Vitis vinifera cv. Kabarcik populations sampled from the Coruh Valley in Turkey, at altitudes of 800–1,150 m. The mean observed number of alleles per locus varied from 2 (loci VVMD7 and VVMD24) to 6 (VVS2) among populations. The population from the highest altitude showed the greatest average number of alleles, 4.5. With regard to the six loci examined in all populations, the mean observed heterozygosity was higher than the expected heterozygosity. Among the loci, VVS2 (probability of identity = 0.137) was found to be the most informative among populations. Genetic distances between populations ranged from 0.072 to 0.216. Genetic differentiation among populations was strongly related to geographic distances in all populations.

Similar content being viewed by others
References
Agar G, Halasz J, Ercisli S (2011) Genetic relationships among wild and cultivated blackberries (Rubus caucasicus L.) based on amplified fragment length polymorphism markers. Plant Biosyst 145(2):347–352
Arnold C, Rossetto M, McNally J, Henry RJ (2002) The application of SSRs characterized for grape (Vitis vinifera) to conservation studies in Vitaceae. Am J Bot 89:22–28
Barreneche T, Bodenes C, Lexer C, Trontin JF, Fluch S, Streiff R, Plomion C, Roussel G, Steinkellner H, Burg K, Favre JM, Glössl J, Kremer A (1998) A genetic linkage map of Quercus robur L. (pedunculate oak) based on RAPD, SCAR, microsatellite, minisatellite, isozyme and 5S rDNA markers. Theor Appl Genet 97:1090–1103
Benjak A, Ercisli S, Vokurka A, Maletic E, Pejic I (2005) Genetic relationships among grapevine cultivars native to Croatia, Greece and Turkey. Vitis 44(2):73–77
Borrego J, Rodriguez I, Andres MT, Martin J, Chavez J, Cabello F, Ibanez J (2001) Characterization of the most important Spanish grape varieties through isoenzyme and microsatellite analysis. Proc Int Symp Mol Markers, Acta Hortic 546:371–375
Bowcock AM, Ruiz-Linares A, Tomfohrde J, Minch E, Kidd JR, Cavalli-Sforza LL (1994) High resolution of human evolutionary trees with polymorphic microsatellites. Nature 368:455–457
Bowers JE, Bandman EB, Meredith CP (1993) DNA fingerprint characterization of some wine grape cultivars. Am J Enol Viticult 44(3):166–274
Bowers JE, Dangl GS, Vignani R, Meredith CP (1996) Isolation and characterization of new polymorphic simple sequence repeat loci in grape (Vitis vinifera L.). Genome 39:628–633
Buckler ES, Thornsberry JM, Kresovich S (2001) Molecular diversity, structure and domestication of grasses. Genet Res 77:213–218
Chen F, Wang A, Chen K, Wan D, Liu J (2009) Genetic diversity and population structure of the endangered and medically important Rheum tanguticum (Polygonaceae) revealed by SSR markers. Biochem Syst Ecol 37(5):613–621
Crespan M, Milani N (2001) The muscats: a molecular analysis of synonyms, homonyms and genetic relationships within a large family of grapevine cultivars. Vitis 40:23–30
Doebley J (1989) Isozymic evidence and evolution of crop plants. In: Soltis ED, Soltis PM (eds) Isozymes in plant biology. Dioscorides, Portland, pp 165–191
Fatahi R, Ebadi A, Basil N, Mehlenbacher SA, Zamani Z (2003) Characterization of Iranian grapevine cultivars using microsatellite markers. Vitis 42(4):185–192
Hernandez-Verdugo S, Luna-Reyes R, Oyama K (2001) Genetic structure and differentiation of wild and domesticated populations of Capsicum annuum from Mexico. Plant Syst Evol 226:129–142
Ibanez J, Andres MT, Molino A, Borrego J (2003) Genetic study of key Spanish grapevine varieties using microsatellite analysis. Am J Enol Viticult 54(1):22–30
Kafkas S, Ozgen M, Dogan Y, Ozcan B, Ercisli S, Serce S (2008) Molecular characterization of mulberry accessions in Turkey by AFLP markers. J Am Soc Hortic Sci 133(4):593–597
Lefort F, Roubelakis-Angelakis KA (2001) Genetic comparison of Greek cultivars of Vitis vinifera L. by nuclear microsatellite profiling. Am J Ecol Vitic 52(2):101–108
Lefort F, Lally M, Thompson D, Douglas GC (1998) Morphological traits, microsatellite fingerprinting and genetic relatedness of a stand of elite oaks (Quercus robur L.) at Tullynally, Ireland. Silvae Genet 47(56):257–262
Lopes MS, Sefc KM, Dias EE, Steinkellner H, Machado MLD, Machado AD (1999) The use of microsatellites for germplasm management in a Portuguese grapevine collection. Theor Appl Genet 99(3–4):733–739
Merdinoglu D, Butterlin G, Bauer C, Balthazard DJ (2000) Comparison of RAPD, AFLP and SSR (microsatellite) markers for genetic diversity analysis in Vitis vinifera L. Proc 7th Int Symp Grapevine Genet Breed, Acta Hortic 528:193–197
Minch E, Ruiz-Linares A, Goldstein DB, Feldman M, Cavalli-Sforza LL (1995) Microsat (version 1.4d): a computer program for calculating various statistics on microsatellite allele data. Stanford University, Stanford
Morgante M, Olivieri AM (1993) PCR amplification of microsatellite markers in plant genetics. Plant J 3:175–182
Mullins MG, Bouquet A, Williams LE (1992) Biology of grapevine. Cambridge University Press, Cambridge
Nunez Y, Fresno J, Torres V, Ponz F, Gallego FJ (2004) Practical use of microsatellite markers to manage Vitis vinifera germplasm: molecular identification of grapevine samples collected blindly in D.O. “El Bierzo” (Spain). J Hortic Sci Biotech 79(3):437–440
Ohsawa T, Ide Y (2008) Global patterns of genetic variation in plant species along vertical and horizontal gradients on mountains. Global Ecol Biogeogr 17:152–162
Otero-Arnaiz A, Casas A, Hamrick JL, Cruse-Sanders J (2005) Genetic variation and evolution of Polaskia chichipe under domestication in Tehuacan Valley, central Mexico. Mol Ecol 14:1603–1611
Paetkau D, Calvert W, Stirling I, Strobeck C (1995) Microsatellite analysis of population structure in Canadian polar bears. Mol Ecol 4:347–354
Pedryc A, Ruthner S, Herman R, Krska B, Hegedus A, Halasz J (2009) Genetic diversity of apricot revealed by a set of SSR markers from linkage group G1. Sci Hortic 121:19–26
Pickersgill B (1969) The domestication of chili peppers. In: Ucko PJ, Dimbley GW (eds) The domestication and exploitation of plants and animals. Duckworth, London, pp 443–450
Powell W, Machray GC, Provan J (1996a) Polymorphism revealed by simple sequence repeats. Trends in Plant Sci 1:215–222
Powell W, Morgante M, Andre C, Hanafey M, Vogel J, Tingey S, Rafalski A (1996b) The comparison of RFLP, RAPD, AFLP and SSR (microsatellite) markers for germplasm analysis. Mol Breed 2:225–238
Rafalski JA, Vogel JM, Morgante M, Powell W, Andre C, Tingey SV (1996) Generating and using DNA markers in plants. In: Birren B, Lai E (eds) Nonmammalian genomic analysis: a practical guide. Academic Press, London, pp 75–134
Rohlf FJ (1988) NTSYSpc: Numerical taxonomy and multiware analysis system version 2.0. Applied Biostatistics, Setauket
Rudmann-Maurer K, Weyand A, Fischer M, Stocklin J (2007) Microsatellite diversity of the agriculturally important alpine grass Poa alpina in relation to land use and natural environment. Ann Bot 100:1249–1258
Russell JR, Fuller JD, Macaulay M, Hatz BG, Jahoor A, Powell W, Waugh R (1997) Direct comparison of levels of genetic variation among barley accessions detected by RFLPs, AFLPs, SSRs and RAPDs. Theor Appl Genet 95:714–722
Scott KD, Eggler P, Seaton G, Rossetto M, Ablett EM, Lee LS, Henry RJ (2000) Analysis of SSRs derived from grape ESTs. Theor Appl Genet 100:723–726
Sefc KM, Lefort F, Grando MS, Scott KD, Steinkellner H, Thomas MR (2001) Microsatellite markers for grapevine: a state of the art. In: Roubelakis-Angelakis KA (ed) Molecular biology and biotechnology of the grapevine. Kluwer Academic, The Netherlands, pp 1–29
Selli F, Bakir M, Inan G, Aygün H, Boz Y, Yasasin AS, Ozer C, Akman B, Söylemezoglu G, Kazan K, Ergül A (2007) Simple sequence repeat-based assessment of genetic diversity in ‘Dimrit’ and ‘Germe’ grapevine accessions from Turkey. Vitis 46:182–187
Sneath PHA, Sokal RR (1973) Numerical taxonomy: the principles and practice of numerical classification. Freeman, San Francisco, p 573
Szczys P, Hughes CR, Kessel RV (2005) Novel microsatellite markers used to determine the population genetic structure of the endangered roseate tern, Sterna dougallii, in Northwest Atlantic and Western Australia. Conserv Genet 6:461–466
Tangolar SG, Soydam S, Bakir M, Karaagac E, Tangolar S, Ergul A (2009) Genetic analysis of grapevine cultivars from the eastern Mediterranean region of Turkey, based on SSR markers. Tarim Bilim Derg 15(1):1–8
Thomas MR, Matsumoto S, Cain P, Scott NS (1993) Repetitive DNA of grapevine: classes present and sequences suitable for cultivar identification. Theor Appl Genet 86:173–180
Wagner HW, Sefc KM (1999) Identity 1.0. Centre for Applied Genetics. University of Agricultural Science, Vienna
Wang L, Yang J, Guo J, Zhao G (2007) Genetic structure and differentiation of Psathyrostachys huashanica populations detected with RAPD markers. Frontier Biol China 2(1):39–45
Wang YH, Korpelainen H, Li CY (2006) Microsatellite polymorphism in the edaphic spruce, Picea asperata, originating from the mountains of China. Silva Fenn 40(4):561–575
Zeder MA, Emshwiller E, Smith BD, Bradley DG (2006) Documenting domestication: the intersection of genetics and archaeology. Trends Genet 22:139–155
Zheng W, Wang LY, Meng LH, Liu JQ (2008) Genetic variation in the endangered Anisodus tanguticus (Solanaceae), an alpine perennial endemic to the Qinghai-Tibetan Plateau. Genetica 132:123–129
Acknowledgments
This study was supported by grants from the Research Funds appropriated to Atatürk University (Project no. 2008/78).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Agar, G., Yildirim, N., Ercisli, S. et al. Determination of Genetic Diversity of Vitis vinifera cv. Kabarcik Populations from the Coruh Valley Using SSR Markers. Biochem Genet 50, 476–483 (2012). https://doi.org/10.1007/s10528-011-9492-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-011-9492-y