Abstract
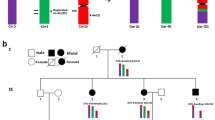
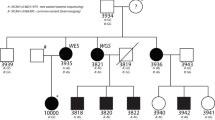
Dyslexia is one of the most common neurodevelopmental disorders where likely many genes are involved in the pathogenesis. So far six candidate dyslexia genes have been proposed, and two of these were identified by rare chromosomal translocations in affected individuals. By systematic re-examination of all translocation carriers in Denmark, we have identified 16 different translocations associated with dyslexia. In four families, where the translocation co-segregated with the phenotype, one of the breakpoints concurred (at the cytogenetic level) with either a known dyslexia linkage region—at 15q21 (DYX1), 2p13 (DYX3) and 1p36 (DYX8)—or an unpublished linkage region at 19q13. As a first exploitation of this unique cohort, we identify three novel candidate dyslexia genes, ZNF280D and TCF12 at 15q21, and PDE7B at 6q23.3, by molecular mapping of the familial translocation with the 15q21 breakpoint.





Similar content being viewed by others
References
Anthoni H, Zucchelli M, Matsson H, Muller-Myhsok B, Fransson I, Schumacher J, Massinen S, Onkamo P, Warnke A, Griesemann H, Hoffmann P, Nopola-Hemmi J, Lyytinen H, Schulte-Korne G, Kere J, Nothen MM, Peyrard-Janvid M (2007) A locus on 2p12 containing the co-regulated MRPL19 and C2ORF3 genes is associated to dyslexia. Hum Mol Genet 16(6):667–677
Bache I, Hjorth M, Bugge M, Holstebroe S, Hilden J, Schmidt L, Brondum-Nielsen K, Bruun-Petersen G, Jensen PK, Lundsteen C, Niebuhr E, Rasmussen K, Tommerup N (2006) Systematic re-examination of carriers of balanced reciprocal translocations: a strategy to search for candidate regions for common and complex diseases. Eur J Hum Genet 14(4):410–417
Bates TC, Luciano M, Castles A, Coltheart M, Wright MJ, Martin NG (2007) Replication of reported linkages for dyslexia and spelling and suggestive evidence for novel regions on chromosomes 4 and 17. Eur J Hum Genet 15(2):194–203
Bates TC, Lind PA, Luciano M, Montgomery GW, Martin NG, Wright MJ (2009) Dyslexia and DYX1C1: deficits in reading and spelling associated with a missense mutation. Mol Psychiatry Nov 10, 2009 [Epub ahead of print]
Bugge M, Bruun-Petersen G, Brondum-Nielsen K, Friedrich U, Hansen J, Jensen G, Jensen PK, Kristoffersson U, Lundsteen C, Niebuhr E, Rasmussen KR, Rasmussen K, Tommerup N (2000) Disease associated balanced chromosome rearrangements: a resource for large scale genotype-phenotype delineation in man. J Med Genet 37(11):858–865
Chapman NH, Igo RP, Thomson JB, Matsushita M, Brkanac Z, Holzman T, Berninger VW, Wijsman EM, Raskind WH (2004) Linkage analyses of four regions previously implicated in dyslexia: confirmation of a locus on chromosome 15q. Am J Med Genet B Neuropsychiatr Genet 131B(1):67–75
Cope N, Harold D, Hill G, Moskvina V, Stevenson J, Holmans P, Owen MJ, O’Donovan MC, Williams J (2005a) Strong evidence that KIAA0319 on chromosome 6p is a susceptibility gene for developmental dyslexia. Am J Hum Genet 76(4):581–591
Cope NA, Hill G, van den Bree M, Harold D, Moskvina V, Green EK, Owen MJ, Williams J, O’Donovan MC (2005b) No support for association between dyslexia susceptibility 1 candidate 1 and developmental dyslexia. Mol Psychiatry 10(3):237–238
Dahdouh F, Anthoni H, Tapia-Paez I, Peyrard-Janvid M, Schulte-Korne G, Warnke A, Remschmidt H, Ziegler A, Kere J, Muller-Myhsok B, Nothen MM, Schumacher J, Zucchelli M (2009) Further evidence for DYX1C1 as a susceptibility factor for dyslexia. Psychiatr Genet 19(2):59–63
de Kovel CG, Hol FA, Heister JG, Willemen JJ, Sandkuijl LA, Franke B, Padberg GW (2004) Genomewide scan identifies susceptibility locus for dyslexia on Xq27 in an extended Dutch family. J Med Genet 41(9):652–657
de Kovel CG, Franke B, Hol FA, Lebrec JJ, Maassen B, Brunner H, Padberg GW, Platko J, Pauls D (2008) Confirmation of dyslexia susceptibility loci on chromosomes 1p and 2p, but not 6p in a Dutch sib-pair collection. Am J Med Genet B Neuropsychiatr Genet 147(3):294–300
Deffenbacher KE, Kenyon JB, Hoover DM, Olson RK, Pennington BF, DeFries JC, Smith SD (2004) Refinement of the 6p21.3 quantitative trait locus influencing dyslexia: linkage and association analyses. Hum Genet 115(2):128–138
DeFries JC, Fulker DW, LaBuda MC (1987) Evidence for a genetic aetiology in reading disability of twins. Nature 329(6139):537–539
Fagerheim T, Raeymaekers P, Tonnessen FE, Pedersen M, Tranebjaerg L, Lubs HA (1999) A new gene (DYX3) for dyslexia is located on chromosome 2. J Med Genet 36(9):664–669
Fiegler H, Gribble SM, Burford DC, Carr P, Prigmore E, Porter KM, Clegg S, Crolla JA, Dennis NR, Jacobs P, Carter NP (2003) Array painting: a method for the rapid analysis of aberrant chromosomes using DNA microarrays. J Med Genet 40(9):664–670
Fisher SE, DeFries JC (2002) Developmental dyslexia: genetic dissection of a complex cognitive trait. Nat Rev Neurosci 3(10):767–780
Fisher SE, Francks C, Marlow AJ, MacPhie IL, Newbury DF, Cardon LR, Ishikawa-Brush Y, Richardson AJ, Talcott JB, Gayan J, Olson RK, Pennington BF, Smith SD, DeFries JC, Stein JF, Monaco AP (2002) Independent genome-wide scans identify a chromosome 18 quantitative-trait locus influencing dyslexia. Nat Genet 30(1):86–91
Francks C, Fisher SE, Olson RK, Pennington BF, Smith SD, DeFries JC, Monaco AP (2002) Fine mapping of the chromosome 2p12-16 dyslexia susceptibility locus: quantitative association analysis and positional candidate genes SEMA4F and OTX1. Psychiatr Genet 12(1):35–41
Gardner C, Robas N, Cawkill D, Fidock M (2000) Cloning and characterization of the human and mouse PDE7B, a novel cAMP-specific cyclic nucleotide phosphodiesterase. Biochem Biophys Res Commun 272(1):186–192
Gayan J, Willcutt EG, Fisher SE, Francks C, Cardon LR, Olson RK, Pennington BF, Smith SD, Monaco AP, DeFries JC (2005) Bivariate linkage scan for reading disability and attention-deficit/hyperactivity disorder localizes pleiotropic loci. J Child Psychol Psychiatry 46(10):1045–1056
Grigorenko EL, Wood FB, Meyer MS, Hart LA, Speed WC, Shuster A, Pauls DL (1997) Susceptibility loci for distinct components of developmental dyslexia on chromosomes 6 and 15. Am J Hum Genet 60(1):27–39
Grigorenko EL, Wood FB, Meyer MS, Pauls JE, Hart LA, Pauls DL (2001) Linkage studies suggest a possible locus for developmental dyslexia on chromosome 1p. Am J Med Genet 105(1):120–129
Hannula-Jouppi K, Kaminen-Ahola N, Taipale M, Eklund R, Nopola-Hemmi J, Kaariainen H, Kere J (2005) The axon guidance receptor gene ROBO1 is a candidate gene for developmental dyslexia. PLoS Genet 1(4):e50
Hsiung GY, Kaplan BJ, Petryshen TL, Lu S, Field LL (2004) A dyslexia susceptibility locus (DYX7) linked to dopamine D4 receptor (DRD4) region on chromosome 11p15.5. Am J Med Genet B Neuropsychiatr Genet 125B(1):112–119
Ingason A, Giegling I, Cichon S, Hansen T, Rasmussen HB, Nielsen J, Jurgens G, Muglia P, Hartmann AM, Strengman E, Vasilescu C, Muhleisen TW, Djurovic S, Melle I, Lerer B, Moller HJ, Francks C, Pietilainen OP, Lonnqvist J, Suvisaari J, Tuulio-Henriksson A, Walshe M, Vassos E, Di Forti M, Murray R, Bonetto C, Tosato S, Cantor RM, Rietschel M, Craddock N, Owen MJ, Peltonen L, Andreassen OA, Nothen MM, St. Clair D, Ophoff RA, Odonovan M, Collier D, Werge T, Rujescu D (2010) A large replication study and meta-analysis in European samples provides further support for association of AHI1 markers with schizophrenia. Hum Mol Genet 19(7):1379–1386
Kaplan DE, Gayan J, Ahn J, Won TW, Pauls D, Olson RK, DeFries JC, Wood F, Pennington BF, Page GP, Smith SD, Gruen JR (2002) Evidence for linkage and association with reading disability on 6p21.3-22. Am J Hum Genet 70(5):1287–1298
Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, Haussler D (2002) The human genome browser at UCSC. Genome Res 12(6):996–1006. http://genome.ucsc.edu/
Kleinjan DJ, van Heyningen V (1998) Position effect in human genetic disease. Hum Mol Genet 7(10):1611–1618
Leonard CM, Kuldau JM, Maron L, Ricciuti N, Mahoney B, Bengtson M, DeBose C (2008) Identical neural risk factors predict cognitive deficit in dyslexia and schizophrenia. Neuropsychology 22(2):147–158
Lyon GR, Shaywitz SE, Shaywitz BA (2003) A definition of dyslexia. Annals of Dyslexia 53:1–15
Marino C, Giorda R, Luisa Lorusso M, Vanzin L, Salandi N, Nobile M, Citterio A, Beri S, Crespi V, Battaglia M, Molteni M (2005) A family-based association study does not support DYX1C1 on 15q21.3 as a candidate gene in developmental dyslexia. Eur J Hum Genet 13(4):491–499
Meng H, Hager K, Held M, Page GP, Olson RK, Pennington BF, DeFries JC, Smith SD, Gruen JR (2005a) TDT-association analysis of EKN1 and dyslexia in a Colorado twin cohort. Hum Genet 118(1):87–90
Meng H, Smith SD, Hager K, Held M, Liu J, Olson RK, Pennington BF, DeFries JC, Gelernter J, O’Reilly-Pol T, Somlo S, Skudlarski P, Shaywitz SE, Shaywitz BA, Marchione K, Wang Y, Paramasivam M, LoTurco JJ, Page GP, Gruen JR (2005b) DCDC2 is associated with reading disability and modulates neuronal development in the brain. Proc Natl Acad Sci USA 102(47):17053–17058
Miscimarra L, Stein C, Millard C, Kluge A, Cartier K, Freebairn L, Hansen A, Shriberg L, Taylor HG, Lewis B, Iyengar SK (2007) Further evidence of pleiotropy influencing speech and language: analysis of the DYX8 region. Hum Hered 63(1):47–58
Nopola-Hemmi J, Taipale M, Haltia T, Lehesjoki AE, Voutilainen A, Kere J (2000) Two translocations of chromosome 15q associated with dyslexia. J Med Genet 37(10):771–775
Nopola-Hemmi J, Myllyluoma B, Haltia T, Taipale M, Ollikainen V, Ahonen T, Voutilainen A, Kere J, Widen E (2001) A dominant gene for developmental dyslexia on chromosome 3. J Med Genet 38(10):658–664
Pennington BF, Gilger JW, Pauls D, Smith SA, Smith SD, DeFries JC (1991) Evidence for major gene transmission of developmental dyslexia. JAMA 266(11):1527–1534
Petryshen TL, Kaplan BJ, Fu Liu M, de French NS, Tobias R, Hughes ML, Field LL (2001) Evidence for a susceptibility locus on chromosome 6q influencing phonological coding dyslexia. Am J Med Genet 105(6):507–517
Pfaffl MW, Tichopad A, Prgomet C, Neuvians TP (2004) Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper—excel-based tool using pair-wise correlations. Biotechnol Lett 26(6):509–515
Poelmans G, Engelen JJ, Van Lent-Albrechts J, Smeets HJ, Schoenmakers E, Franke B, Buitelaar JK, Wuisman-Frerker M, Erens W, Steyaert J, Schrander-Stumpel C (2009) Identification of novel dyslexia candidate genes through the analysis of a chromosomal deletion. Am J Med Genet B Neuropsychiatr Genet 150B(1):140–147
Raven JC (1994) Standard progressive matrices: sets A, B, C, D and E. Oxford Psychologists Press Ltd, Oxford
Raven JC, Court JH, Raven J (1988) Manual for Raven’s progressive matrices and vocabulary scales. Section 3, The standard progressive matrices. H. K. Lewis & Co. Ltd., London
Revheim N, Butler PD, Schechter I, Jalbrzikowski M, Silipo G, Javitt DC (2006) Reading impairment and visual processing deficits in schizophrenia. Schizophr Res 87(1–3):238–245
Rosen GD, Bai J, Wang Y, Fiondella CG, Threlkeld SW, LoTurco JJ, Galaburda AM (2007) Disruption of neuronal migration by RNAi of Dyx1c1 results in neocortical and hippocampal malformations. Cereb Cortex 17(11):2562–2572
Scerri TS, Fisher SE, Francks C, MacPhie IL, Paracchini S, Richardson AJ, Stein JF, Monaco AP (2004) Putative functional alleles of DYX1C1 are not associated with dyslexia susceptibility in a large sample of sibling pairs from the UK. J Med Genet 41(11):853–857
Schulte-Korne G, Grimm T, Nothen MM, Muller-Myhsok B, Cichon S, Vogt IR, Propping P, Remschmidt H (1998) Evidence for linkage of spelling disability to chromosome 15. Am J Hum Genet 63(1):279–282
Shaywitz SE, Shaywitz BA, Fletcher JM, Escobar MD (1990) Prevalence of reading disability in boys and girls. Results of the Connecticut Longitudinal Study. JAMA 264(8):998–1002
Smith SD, Kimberling WJ, Pennington BF, Lubs HA (1983) Specific reading disability: identification of an inherited form through linkage analysis. Science 219(4590):1345–1347
Taipale M, Kaminen N, Nopola-Hemmi J, Haltia T, Myllyluoma B, Lyytinen H, Muller K, Kaaranen M, Lindsberg PJ, Hannula-Jouppi K, Kere J (2003) A candidate gene for developmental dyslexia encodes a nuclear tetratricopeptide repeat domain protein dynamically regulated in brain. Proc Natl Acad Sci USA 100(20):11553–11558
Tzenova J, Kaplan BJ, Petryshen TL, Field LL (2004) Confirmation of a dyslexia susceptibility locus on chromosome 1p34-p36 in a set of 100 Canadian families. Am J Med Genet B Neuropsychiatr Genet 127B(1):117–124
Uittenbogaard M, Chiaramello A (2002) Expression of the bHLH transcription factor Tcf12 (ME1) gene is linked to the expansion of precursor cell populations during neurogenesis. Brain Res Gene Expr Patterns 1(2):115–121
Veltman IM, Veltman JA, Arkesteijn G, Janssen IM, Vissers LE, de Jong PJ, van Kessel AG, Schoenmakers EF (2003) Chromosomal breakpoint mapping by arrayCGH using flow-sorted chromosomes. Biotechniques 35(5):1066–1070
Wigg KG, Couto JM, Feng Y, Anderson B, Cate-Carter TD, Macciardi F, Tannock R, Lovett MW, Humphries TW, Barr CL (2004) Support for EKN1 as the susceptibility locus for dyslexia on 15q21. Mol Psychiatry 9(12):1111–1121
Wigg KG, Feng Y, Crosbie J, Tannock R, Kennedy JL, Ickowicz A, Malone M, Schachar R, Barr CL (2008) Association of ADHD and the Protogenin gene in the chromosome 15q21.3 reading disabilities linkage region. Genes Brain Behav 7(8):877–886
Willcutt EG, Pennington BF (2000) Comorbidity of reading disability and attention-deficit/hyperactivity disorder: differences by gender and subtype. J Learn Disabil 33(2):179–191
Williams J, O’Donovan MC (2006) The genetics of developmental dyslexia. Eur J Hum Genet 14(6):681–689
World Health Organisation (2010) International statistical classification of diseases and related health problems, 10th Revision, Version for 2007, http://apps.who.int/classifications/apps/icd/icd10online/
Zhou K, Asherson P, Sham P, Franke B, Anney RJ, Buitelaar J, Ebstein R, Gill M, Brookes K, Buschgens C, Campbell D, Chen W, Christiansen H, Fliers E, Gabriels I, Johansson L, Marco R, Mulas F, Muller U, Mulligan A, Neale BM, Rijsdijk F, Rommelse N, Uebel H, Psychogiou L, Xu X, Banaschewski T, Sonuga-Barke E, Eisenberg J, Manor I, Miranda A, Oades RD, Roeyers H, Rothenberger A, Sergeant J, Steinhausen HC, Taylor E, Thompson M, Faraone SV (2008) Linkage to chromosome 1p36 for attention-deficit/hyperactivity disorder traits in school and home settings. Biol Psychiatry 64(7):571–576
Acknowledgements
We are grateful to the patients and families for participation in this study. We thank Dr. Elena Grigorenko, Child Study Center, Department of Psychology, Yale University, New Haven, CT, for communication concerning new dyslexia linkage loci. We also thank Claus Hansen and Karen Friis Henriksen for help with gene expression analysis. Wilhelm Johannsen Centre for Functional Genome research is established by the Danish National Research Foundation who also financed this study by an International Talent Recruitment grant to RB.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by Elena Grigorenko and Brett Miller.
Rights and permissions
About this article
Cite this article
Buonincontri, R., Bache, I., Silahtaroglu, A. et al. A Cohort of Balanced Reciprocal Translocations Associated with Dyslexia: Identification of Two Putative Candidate Genes at DYX1. Behav Genet 41, 125–133 (2011). https://doi.org/10.1007/s10519-010-9389-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10519-010-9389-2