Abstract
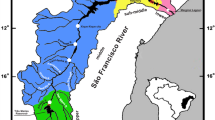
The potential for genetic contamination of stocks arising from translocation and subsequent release or escape of translocated and/or genetically mixed stocks may be a significant risk to wild populations. In this context, we undertook a population genetic survey of stocks of lumpfish (Cyclopterus lumpus) along the Norwegian coast to establish the existing genetic population structure, which will aid the development of policy of the species’ use as cleaner fish in salmonid aquaculture. This was done by using 14 microsatellite loci and 287 specimens collected at five fishing grounds, covering most of the Norwegian coastline from south to north, with additional 18 samples of first-generation reared fish from a fish farm outside Tromsø (North Norway). Overall, there was no indication of significant spatial genetic structuring or of positive correlation between geographic and genetic distance among the wild lumpfish samples. These results suggest that, should translocated individuals escape from aquaculture in Norway, this will probably have little to no impact on the genetic composition of the local fish population.




Similar content being viewed by others
Change history
26 October 2017
A correction to this article has been published.
References
Asche F, Guttormsen AG, Sebulonsen T, Sissener EH (2005) Competition between farmed and wild salmon: the Japanese salmon market. Agr Econm 33:333–340
Anon (2016a) Rognkjeks er den tredje viktigste oppdrettsarten i Norge (lumpfish is the third most important aquaculture species in Norway). Norwegian Directorate of Fisheries, Bergen, http://www.fiskeridir.no/statistikk/akvakultur/statistiskepublikasjoner/foreloepig-statistikk-for-akvakultur (in Norwegian)
Anon (2016b) Orientering om regulering av fisket etter leppefisk (Orientation on the regulation of wrasse fisheries in Norway). Norwegian Directorate of Fisheries, Bergen, Norway, nr. 19/2016 pp. 13 (in Norwegian)
Belkhir K, Borsa P, Chikhi L, Raufaste N, Bonhomme F (2004) Genetix 4.05, logiciel sous Windows TM pour la génétique des populations. Laboratoire Génome, Populations, Interactions, CNRS UMR 5000, Université de Montpellier II, Montpellier
Bekkevold D, Hansen MM, Nielsen EE (2006) Genetic impact of gadoid culture on wild fish populations: predictions, lessons from salmonids, and possibilities for minimizing adverse effects. ICES J Mar Sci 63:198–208
Benjamini Y, Yekutieli D (2001) The control of the false discovery rate in multiple testing under dependency. Ann Stat 29:1165–1188
Blonk RJW, Komen J, Kamstra A, Crooijmans RPMA, Van Arendonk JAM (2009) Levels of inbreeding in group mating captive broodstock populations of common sole, (Solea solea), inferred from parental relatedness and contribution. Aquaculture 289:26–31
Boxaspen K (2006) A review of the biology and genetics of sea lice. ICES J Sea Res 63:1304–1316
Coughlan JP, Imsland AK, Galvin PT, FitzGerald RD, Nævdal G, Cross TF (1998) Microsatellite DNA variation in wild populations and farmed strains of turbot Scophthalmus maximus from Ireland and Norway. J Fish Biol 52:916–922
Denholm I, Devine GJ, Horsberg TE, Sevatdal S, Fallang A, Nolan DV, Powell R (2002) Analysis and management of resistance to chemotherapeutants in salmon lice Lepeophtheirus salmonis (Krøyer) (Copepoda: Caligidae). Pest Man Sci 58:528–536
Earl DA, VonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Con Gen Res 4:359–361
Eriksen E, Durif CMF, Prozorkevich D (2014) Lumpfish (Cyclopterus lumpus) in the Barents Sea: development of biomass and abundance indices, and spatial distribution. ICES J Mar Sci 71:2398–2402
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and windows. Mol Ecol Res 10:564–567
Falush D, Stephens M, Pritchard J (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Falush D, Stephens M, Pritchard JK (2007) Inference of population structure using multilocus genotype data: dominant markers and null alleles. Mol Ecol Not 7:574–578
Garcia-Mayoral E, Olsen E, Hedeholm R, Post S, Nielsen EE, Bekkevold D (2016) Genetic structure of West Greenland populations of lumpfish Cyclopterus Lumpus. J Fish Biol 89:2625–2642
Gonzales EB, Knutsen H, Jorde PE (2016) Habitat discontinuities separate genetically divergent populations of a rocky shore marine fish. PLoS One 11:1–17
Hara M, Sekino M (2007) Genetic differences between hatchery stocks and natural populations in pacific abalone (Haliotis discus) estimated using microsatellite DNA markers. Mar Biotechnol 9:74–81
Hindar K, Fleming IA, McGinnity P, Diserud OH (2006) Genetic and ecological effects of salmon farming on wild salmon: modelling from experimental results. ICES J Mar Sci 63:1234–1247
Hubisz MJ, Falush D, Stephens M, Pritchard J (2009) Inferring weak population structure with the assistance of sample group information. Mol Ecol Res 9:1322–1332
Hutchings JA, Fraser DJ (2008) The nature of fisheries- and farming-induced evolution. Mol Ecol 17:294–313
Imsland AK, Reynolds P, Eliassen G, Hangstad TA, Foss A, Vikingstad E, Elvegård TA (2014a) The use of lumpfish (Cyclopterus lumpus L.) to control sea lice (Lepeophtheirus salmonis Krøyer) infestations in intensively farmed Atlantic salmon (Salmo salar L.) Aquaculture 424-425:18–23
Imsland AK, Reynolds P, Eliassen G, Hangstad TA, Nytrø AV, Foss A, Vikingstad E, Elvegård TA (2014b) Assessment of growth and sea lice infection levels in Atlantic salmon stocked in small-scale cages with lumpfish. Aquaculture 433:137–142
Imsland AK, Reynolds P, Eliassen G, Hangstad TA, Nytrø AV, Foss A, Vikingstad E, Elvegård TA (2014c) Notes on behaviour of lumpfish in sea pens with and without Atlantic salmon. J Ethol 32:117–122
Imsland AK, Reynolds P, Eliassen G, Hangstad TA, Nytrø AV, Foss A, Vikingstad E, Elvegård TA (2015a) Feeding preferences of lumpfish (Cyclopterus lumpus L.) maintained in open net-pens with Atlantic salmon (Salmo salar L.) Aquaculture 436:47–51
Imsland AK, Reynolds P, Eliassen G, Hangstad TA, Nytrø AV, Foss A, Vikingstad E, Elvegård TA (2015b) Assessment of suitable substrates for lumpfish in sea pens. Aquac Int 23:639–645
Jimenez DF, Heuch PA, Revie CW, Gettinby G (2012) Confidence in assessing the effectiveness of bath treatments for the control of sea lice on Norwegian salmon farms. Aquaculture 344:58–65
Jørgensen OA, Hvingel C, Møller PR, Treble MA (2005) Identification and mapping of bottom fish assemblages in Davis Strait and southern Baffin Bay. Can J Fish Aquat Sci 62:1833–1852
Kennedy J, Jonsson ST, Kasper JM, Olafsson HG (2015) Movements of female lumpfish (Cyclopterus lumpus) around Iceland. ICES J Mar Sci 72:880–889
Krkosek M, Lewis MA, Volpe JP, Morton A (2006) Fish farms and sea lice infestations of wild juvenile salmon in the Broughton archipelago—a rebuttal to brooks (2005). Rev Fish Sci 14:1–11
Pampoulie C, Skirnisdottir S, Olafsdottir G, Helyar SJ, Thorsteinsson V, Jónsson SÞ, Fréchet A, Durif CMF, Sherman S, Lampart-Kalużniacka M, Hedeholm R, Ólafsson H, Daníelsdóttir AK, Kasper JM (2014) Genetic structure of the lumpfish Cyclopterus lumpus across the North Atlantic. ICES J Mar Sci 71:2390–2397
Phinchongsakuldit J, Chaipakdee O, Collins JF, Jaroensutasinee M, Brookfield JFY (2013) Population genetics of cobia (Rachycentron canadum) in the Gulf of Thailand and Andaman Sea: fisheries management implications. Aquac Int 21:197–217
Peakall R, Smouse PE (2006) Genalex 6: genetic analysis in excel. Population genetic software for teaching and research. Mol Eco Not 6:288–295
Peakall R, Smouse PE (2012) GenAIEx 6.5: genetic analysis in excel. Population genetic software for teaching and research-an update. Bioinformatics 28:2537–2539
Porta J, Porta JM, Cañavate P, Martínez-Rodríguez G, Álvarez MC (2007) Substantial loss of genetic variation in a single generation of Senegalese sole (Solea senegalensis) culture: implications in the domestication process. J Fish Biol 71:223–234
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Quintela M, Danielsen EA, Lopez L, Barreiro R, Svåsand T, Knutsen H, Skiftesvik AB, Glover KA (2016) Is the ballan wrasse (Labrus bergylt) two species? Genetic analysis reveals within-species divergence associated with plain and spotted morphotype frequencies. Int Zool 11:162–172
Ribergaard MH (2014) Oceanographic investigations off West Greenland 2013. Copenhagen: Danish Meteorological Institute Centre for Ocean and Ice. Available at http://ocean.dmi.dk/staff/mhri/Docs/scr14-001.pdf/
Rousset F (2008) GENEPOPˊ007: a complete re-implementation of the GENEPOP software for windows and Linux. Mol Ecol Res 8:103–106
Sanchez P, Vinas J, Bremer JRA, Ambrosio PP, Flos R (2012) Loss of genetic variability in a hatchery strain of Senegalese sole (Solea senegalensis) revealed by sequence data of the mitochondrial DNA control region and microsatellite markers. Sco Mar 76:225–235
Sayer MDJ, Reader JP (1996) Exposure of goldsinny, rock cook and corkwing wrasse to low temperature and low salinity: survival, blood physiology and seasonal variation. J Fish Biol 49:41–63
Sekino M, Hara M, Taniguchi N (2002) Loss of microsatellite and mitochondrial DNA variation in hatchery strains of Japanese flounder Paralichthys olivaceus. Aquaculture 213:101–122
Skiftesvik AB, Blom G, Agnalt AL, Durif CMF, Browman H, Bjelland RM, Harkestad LS, Farestveit E, Paulsen OI, Fauske M, Havelin T, Johnsen K, Mortensen S (2014) Wrasse (Labridae) as cleaner fish in salmonid aquaculture—the Hardangerfjord as a case study. Mar Biol Res 10:289–300
Skirnisdottir S, Olafsdottir G, Olafsson K, Jendrossek T, Lloyd HD, Helyar S, Pampoulie C, Danielsdottir AK, Kasper JM (2013) Twenty-two novel microsatellite loci for lumpfish (Cyclopterus lumpus). Con Gen Res 5:177–179
Szpiech ZA, Jakobsson M, Rosenberg NA (2008) ADZE: a rarefaction approach for counting alleles private to combinations of populations. Bioinformatics 24:2498–2504
Treasurer JW (2002) A review of potential pathogens of sea lice and the application of cleaner fish in biological control. Pest Managem Sci 58:546–558
Tully O, Daly P, Lysaght S, Deady S, Varian SJA (1996) Use of cleaner-wrasse (Centrolubrus exoletus (L.) and Ctenolabrus rupestris (L.)) to control infestations of Caligus elongatus Nordmann on farmed Atlantic salmon. Aquaculture 142:11–24
Van Oosterhout C, Hutchinson WF, Willis DPM, Shipley P (2004) MICRO-checker: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Not 4:535–538
Acknowledgements
We are grateful to Mr. Thor Jonassen, Mr. Thor Arne Hangstad and Mrs. Mette Remen at Akvaplan-niva, Norway, and Mrs. Caroline Durif and Mr. Nola Velimir at Marine Research Institute, Bergen, Norway, for their assistance in sampling the biological samples of lumpfish. Financial support was provided by the Regional Research Fund of Northern Norway (grants 220615, POPKJEKS, and 239135, AVLUS).
Author information
Authors and Affiliations
Corresponding author
Additional information
A correction to this article is available online at https://doi.org/10.1007/s10499-017-0197-z.
Rights and permissions
About this article
Cite this article
Jónsdóttir, Ó.D.B., Schregel, J., Hagen, S.B. et al. Population genetic structure of lumpfish along the Norwegian coast: aquaculture implications. Aquacult Int 26, 49–60 (2018). https://doi.org/10.1007/s10499-017-0194-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10499-017-0194-2