Abstract
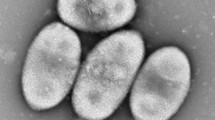
An aerobic, Gram-negative, rod-shaped, non-motile bacterium was isolated from a liquid culture of the dinoflagellate Alexandrium minutum and designated as strain LMIT003T. Analyses of 16S rRNA gene sequences revealed that this strain is affiliated with the genus Tropicibacter in the family Rhodobacteraceae of the class Alphaproteobacteria and shows high similarity (97.3%) with the type species Tropicibacter naphthalenivorans C02T. Phylogenetic analysis based on core genes showed that the isolate groups with members of the genus Tropicibacter. The G + C content of strain LMIT003T was determined to be 61.9 mol%. The major fatty acids identified included summed feature 8 (C18:1 ω7c/C18:1 ω6c), C18:0, C12:1 3-OH and C16:0. The sole respiratory lipoquinone was found to be ubiquinone-10. The major polar lipids were found to be phosphatidylcholine, phosphatidylethanolamine, phosphatidylglycerol, two unidentified aminolipids and four unidentified phospholipids. The draft genome size of strain LMIT003T was determined to be 4.8 Mbp. The average nucleotide identity values between strain LMIT003T and reference Tropicibacter species were determined to be 78.7% (T. naphthalenivorans) and 74.2% (Tropicibacter phthalicicus). Based on physiological, chemotaxonomic and phylogenetic analysis, strain LMIT003T is concluded to represent a novel species in the genus Tropicibacter, for which the name Tropicibacter alexandrii sp. nov., is proposed. The type strain is LMIT003T (= CICC 24660T = KCTC 62895T).


Similar content being viewed by others
Abbreviations
- CICC:
-
China Center for Industrial Culture Collection
- KCTC:
-
Korean Collection for Type Cultures
References
Buchan A, LeCleir GR, Gulvik CA, Gonzalez JM (2014) Master recyclers: features and functions of bacteria associated with phytoplankton blooms. Nat Rev Microbiol 12:686–698
Chun J, Oren A, Ventosa A, Christensen H, Arahal DR, Costa MS, Rooney AP, Yi H, Xu XW, Meyer SD, Trujillo ME (2018) Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol 68:461–466
Cooper MB, Smith AG (2015) Exploring mutualistic interactions between microalgae and bacteria in the omics age. Curr Opin Plant Biol 26:147–153
Croft MT, Lawrence AD, Raux-Deery E, Warren MJ, Smith AG (2005) Algae acquire vitamin B12 through a symbiotic relationship with bacteria. Nature 438:90–93
Curson AR, Todd JD, Sullivan MJ, Johnston AW (2011) Catabolism of dimethylsulphoniopropionate: microorganisms, enzymes and genes. Nat Rev Microbiol 9:849–859
Emms DM, Kelly S (2018) OrthoFinder2: fast and accurate phylogenomic orthology analysis from gene sequences. bioRxiv, https://dx.doi.org/10.1101/466201
Field CB, Behrenfeld MJ, Randerson JT, Falkowski P (1998) Primary production of the biosphere: integrating terrestrial and oceanic components. Science 281:237–240
Guillard RRL (1975) Culture of phytoplankton for feeding marine invertebrates. In: Smith WL, Chanley MH (eds) Culture of marine invertebrate animals. Plenum Press, New York, pp 29–60
Gurevich A, Saveliev V, Vyahhi N, Tesler G (2013) QUAST: quality assessment tool for genome assemblies. Bioinformatics 29:1072–1075
Harwati TU, Kasai Y, Kodama Y, Susilaningsih D, Watanabe K (2009) Tropicibacter naphthalenivorans gen. nov., sp. nov., a polycyclic aromatic hydrocarbon-degrading bacterium isolated from Semarang Port in Indonesia. Int J Syst Evol Microbiol 59:392–396
Iwaki H, Nishimura A, Hasegawa Y (2012) Tropicibacter phthalicicus sp. nov., a phthalate-degrading bacterium from seawater. Curr Microbiol 64:392–396
Katoh K, Misawa K, Kuma K, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30:3059–3066
Kim M-J, Jeong S-Y, Seong -J, Lee S-J (2008) Isolation, identification, and algicidal activity of marine bacteria against Cochlodinium polykrikoides. J Appl Phycol 20:1069–1078
Kumar S, Stecher G, Tamura K (2016) MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Mayali X, Azam F (2004) Algicidal bacteria in the sea and their impact on algal blooms. J Eukaryot Microbiol 51:139–144
Minnikin DE, O’donell AG, Goodfellow M, Alderson G, Athalye M, Schaal A, Parlett JH (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Morgan NP, Paramvir SD, Adam PA (2009) FastTree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol Biol Evol 26:1641–1650
Nedashkovskaya OI, Kim SB, Han SK, Rhee MS, Lysenko AM, Falsen E, Frolova GM, Mikhailov VV, Bae KS (2004) Ulvibacter litoralis gen. nov., sp. nov., a novel member of the family Flavobacteriaceae isolated from the green alga Ulva fenestrata. Int J Syst Evol Microbiol 54:119–123
Newton R, Griffin L, Loftis K, Meile C, Gifford S, Givens CC, Howard E, King E, Oakley C, Reisch C, Rinta-Kanto J, Sharma S, Sun S, Varaljay V, Vila-Costa M, Westrich J, Moran MA (2010) Genome characteristics of a generalist marine bacterial lineage. ISME J 4:784–798
Nurk S, Bankevich A, Antipov D, Gurevich AA (2013) Assembling genomes and mini-metagenomes from highly chimeric reads. In: Deng M, Jiang R, Sun F, Zhang X (eds) Research in computational molecular biology. Springer, Berlin, pp 158–170
Park S-K, Kim M-S, Jung M-J, Nam Y-D, Park E-J, Roh SW, Bae J-W (2011) Brachybacterium squillarum sp. nov., isolated from salt-fermented seafood. Int J Syst Evol Microbiol 61:1118–1122
Ramanan R, Kim BH, Cho DH, Oh HM, Kim HS (2016) Algae-bacteria interactions: evolution, ecology and emerging applications. Biotechnol Adv 34:14–29
Rodionov DA, Gelfand MS, Todd JD, Curson AR, Johnston AW (2006) Computational reconstruction of iron- and manganese-responsive transcriptional networks in alpha-proteobacteria. PLoS Comput Biol 2:e163
Sasser M (2001) Identification of bacteria by gas chromatography of cellular fatty acids. MIDI Technical Note 101. MIDI Inc , Newark, DE
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069
Seymour JR, Amin SA, Raina J-B, Stocker R (2017) Zooming in on the phycosphere: the ecological interface for phytoplankton-bacteria relationships. Nat Microbiol 2:17065
Shieh WY, Chen Y-W, Chaw S-M, Chiu H-H (2003) Vibrio ruber sp. nov., a red, facultatively anaerobic, marine bacterium isolated from sea water. Int J Syst Evol Microbiol 53:479–484
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Gerhardt P, et al. (eds) Methods for general and molecular bacteriology. American Society for Microbiology, Washington DC, pp 607–654
Tittsler RP, Sandholzer LA (1936) The use of semi-solid agar for the detection of bacterial motility. J Bacteriol 31:575–580
Wagner-Döbler I, Ballhausen B, Berger M, Brinkhoff T, Buchholz I, Bunk B, Cypionka H, Daniel R, Drepper T, Gerdts G, Hahnke S, Han C, Jahn D, Kalhoefer D, Kiss H, Klenk H-P, Kyrpides N, Liebl W, Liesegang H, Meincke L, Pati A, Petersen J, Piekarski T, Pommerenke C, Pradella S, Pukall R, Rabus R, Stackebrandt E, Thole S, Thompson L, Tielen P, Tomasch J, Von JM, Wanphrut N, Wichels A, Zech H, Simon M (2009) The complete genome sequence of the algal symbiont Dinoroseobacter shibae: a hitchhiker's guide to life in the sea. ISME J 4:61–77
Wang X, Li ZJ, Su JQ, Tian Y, Ning XR, Hong H, Zheng T (2010) Lysis of a red-tide causing alga, Alexandrium tamarense, caused by bacteria from its phycosphere. Biol Control 52:123–130
Wang H, Laughinghouse HD, Anderson MA, Chen F, Willliams E, Place AR, Zmora O, Zohar Y, Zheng T, Hill R (2012) Novel bacterial isolate from Permian groundwater, capable of aggregating potential biofuel-producing microalgae Nannochloropsis oceanica IMET1. Appl Environ Microb 78:1445–1453
Wang X, Yang HX, Zhang YK (2015) Luteimonas soli sp. nov., isolated from farmland soil. Int J Syst Evol Microbiol 65:4809–4815
Wayne LG (1987) International committee on systematic bacteriology. Report of the Ad Hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
Wirth JS, Whitman WB (2018) Phylogenomic analyses of a clade within the roseobacter group suggest taxonomic reassignments of species of the genera Aestuariivita, Citreicella, Loktanella, Nautella, Pelagibaca, Ruegeria, Thalassobius, Thiobacimonas and Tropicibacter, and the proposal of six novel genera. Int J Syst Evol Microbiol 68:2393–2411
Yoon S-H, Ha S-M, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 67:1613–1617
Acknowledgements
We acknowledge the Marine Culture Collection of China (MCCC) for assistance in identifying the novel strain. We also thank Dr. Mary Atieno from International Centre for Tropical Agriculture (CIAT) for English language assistance during the preparation of this manuscript.
Funding
Funding for this study was provided by National Natural Science Foundation of China (41676116), SAIL Foundation for Distinguished Scholars (Hui Wang), Pearl River Scholar for Young Investigator (Hui Wang) and Innovation Projects of Universities in Guangdong Province (2017KTSCX072).
Author information
Authors and Affiliations
Contributions
HW, ZH and XW designed the experiments. XW, JZ, JF and HW performed the experiments, collected and analysed the data. XW, AS and HW wrote the draft of the article. XW, AS, ZH and HW revised the draft of the article.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
This article does not contain any human and animals studies.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, X., Zhu, J., Feng, J. et al. Tropicibacter alexandrii sp. nov., a novel marine bacterium isolated from the phycosphere of a dinoflagellate, Alexandrium minutum. Antonie van Leeuwenhoek 113, 311–320 (2020). https://doi.org/10.1007/s10482-019-01339-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-019-01339-8