Abstract
The sudden outbreak of swine flu has increased the global demand of shikimic acid which is an industrially interesting compound, as it is used as a key starting material for the synthesis of a neuraminidase inhibitor Tamiflu®, for the treatment of antiviral infections such as swine flu. Statistical optimization and evaluation of medium components for the production of shikimic acid by Citrobacter freundii is addressed in the present investigation. Plackett–Burman design was applied for the screening of the most significant variables affecting shikimic acid production, where glucose, asparagine, KH2PO4, CaCO3 and agitation rate were the most significant factors. Response surface methodology was also employed to study the interaction among the most significant variables through which shikimic acid production increased to 12.76 g/L. Further, fed-batch studies resulted in the production of 22.32 g/L of shikimic acid. The scalability of the process was also confirmed by running 14 L bioreactor (7.5 L production medium) where 20.12 g/L of shikimic acid was produced. In addition the antibacterial activity of the shikimic acid produced was analysed against four Gram positive and four Gram negative bacteria and it was found to have a greater inhibition effect against the Gram negative bacteria.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Swine/Avian flu has emerged as a primary public health concern of the 21st century. Although, various strains of avian influenza have been recognised for decades, the lethality and mutability of H1N1 subtype of influenza virus has served as the source of human influenza pandemic—swine flu. A highly lethal but non-human-to-human transferable influenza A—H1N1, emerged and raged through South-East Asian countries, Egypt and other countries, after the preparation of the world for 2008 pandemic influenza i.e. H5N1—the bird flu (Lagace-Wiens and Rubinstein 2010).
The causative agent of swine flu is Influenza A H1N1 virus. It is a member of the Orthomyxoviridae family of viruses (Racaniello 2009). Multipartite, negative sense, single stranded RNA genome and a lipid envelope are the characteristics of influenza viruses (Beigel and Bray 2008). The symptoms of swine flu are quite similar to those of the seasonal flu (Saxena et al. 2012). Thus, it is of utmost importance to find an ultimate cure for this disease.
In this respect, researchers suggest that shikimic acid is the basic compound for the production of swine flu drug Tamiflu and is found in the seeds of an evergreen Chinese plant Star anise (Illicium vernum) (Gibson et al. 1962). Shikimic acid consists of a highly functional six membered carbocyclic ring and three asymmetric centres. Therefore, it has been proved to be an attractive enantiomerically pure building block for the synthesis of different biologically important compounds (Evans and Osman 1974). Apart from being used as a key starting material for the synthesis of the neuraminidase inhibitor (Tamiflu) (Gibson et al. 1962; Kim et al. 1997, 1998), shikimic acid also plays an important role in the biosynthesis of many other important natural products such as aromatic amino acids, alkaloids, phenolics, and phenylpropanoids (Stavric and Stolz 1976). Furthermore, shikimic acid also has several other applications in food, pharmaceutical and cosmetic industries (Bongaerts et al. 2001; Kramer et al. 2003). It is also known for its antibacterial, antifungal properties and more recently for its use in arthritis (Gibson et al. 1962; Johansson et al. 2005).
Bulk production of Tamiflu is a difficult task as it not only takes months to complete but is hazardous too. Careful handling and relatively mild reaction conditions are required for the synthesis of drug, as potentially explosive azide chemistry is involved in it. Production of large amount of drug requires enough amount of starting material (shikimic acid). The major harvesting period for star anise is from March to May which is not enough to supply large amount of shikimic acid (starting material) to meet the drug requirement worldwide (Bradley 2005). Besides star anise, Ginkgo biloba is also considered as another source of shikimic acid (Chandran et al. 2003). Since, both the raw materials are in short supplies, the pharmaceutical industry needs to find an alternative sustainable supply which is time independent and cost effective.
Interestingly, it was confirmed later that shikimic acid is also a natural intermediate in the formation of aromatic amino acids in the microorganisms. Hence, the research focus and the area of interest of shikimic acid synthesis shifted towards the microbial route. Researchers could boost the production of shikimic acid in strains of the bacterium Escherichia coli by ramping up enzyme activity and feeding the microbe a diet of carbohydrates. Recombinant and mutant strains of few bacteria such as Citrobacter freundii, Bacillus amyloliquefaciens, E. coli and Aerobacter aerogenes are also known to produce shikimic acid (Kim et al. 1997; Rohloff et al. 1998). In this respect, in order to increase the availability of shikimic acid and thus Tamiflu’s availability, fermentative production processes have been tried by different researchers (Bongaerts et al. 2001; Kramer et al. 2003; Johansson et al. 2005; Li et al. 1999; Johansson and Liden 2006; Ahn et al. 2008). The fermentative approach for the production of shikimic acid from glucose presents an excellent and even more sustainable alternative to meet the current market volume at a competitive price level.
Production of shikimic acid by fermentation has many advantages over the plant sources, owing to its simplicity and easy availability of shikimic acid at the time of need. Although, the recombinant and mutant strains of few microbes have been exploited for the production of shikimic acid by fermentation, yet an economically viable biosynthetic process for shikimic acid production from wild type bacteria is still to be developed.
Keeping this in mind, the present investigation has been carried out using a natural isolate C. freundii. Here, the significance of medium components in the production of shikimic acid from C. freundii is addressed through Plackett–Burman (PB) and Response surface methodology (RSM). To find out the optimum levels of the most significant variables as well as studying possible interactions among studied variables, Face centred central composite design (FCCCD) was employed. Production of any biomolecule basically depends on its substrate utilization. Therefore, fed-batch studies were carried out to further increase shikimic acid production. In addition to this, the scalability of the process was analysed by translating the process to 14 L bioreactor thus, providing a base for the industrial production of shikimic acid. Further, the antibacterial activity of shikimic acid produced was also analysed against four Gram positive and four Gram negative bacteria.
Materials and methods
Chemicals
Shikimic acid was purchased from Sigma (St. Louis, USA). HPLC grade water (Thomas Baker, India) was used to prepare the mobile phase for HPLC. All other chemicals used for media preparation were of analytical grade and were purchased from the local commercial sources.
Organism and growth conditions
In our previous study (Tripathi et al. 2013), C. freundii YRL11 was selected for shikimic acid production. The gene sequence of this strain was later submitted to NCBI GeneBank and was provided the Accession no. KC879274. Thus, this strain C. freundii KC879274 was taken for further investigations on shikimic acid production. The components for the shikimic acid production medium were (g/L): Glucose, 50.0; Asparagine, 45.0; CaCO3, 20.0; KH2PO4, 1.0; MgSO4, 0.4; l-Tyrosine, 0.1; l-Tryptophan, 0.1; l-Phenylalanine, 0.1; pABA, 0.001; FeSO4·7H2O, 0.0099; MnSO4·H2O, 0.0072; ZnCl2, 0.0025; CuSO4·5H2O, 0.0005 at pH 6.0 ± 0.2 (Tripathi et al. 2013). The medium was sterilised at 10 psi (at 110 °C for 20 min) followed by inoculation with 6 % (v/v) seed medium (composition (g/L): Tryptone, 10.0; Yeast Extract, 5.0; NaCl, 5.0 at pH 7.5 ± 0.2) as suggested by Shirai et al. (2001). However, the volume of the inoculum was decided on the basis of the results of the ‘one-variable-at-a-time’ approach as obtained in the previous study (Tripathi et al. 2013). Further, the inoculated medium was incubated at 30 °C and 200 rpm for 72 h. Samples (2 ml) were withdrawn at a regular interval of 12 h up to 72 h.
Four Gram positive (Bacillus cereus, Bacillus subtilis, Staphylococcus aureus & Streptococcus pyogenes) and four Gram negative bacteria (Escherichia coli, Klebsiella pneumoniae, Salmonella typhi & Pseudomonas aeruginosa) were also maintained in the nutrient agar medium with the composition (g/L): NaCl, 5.0; Peptone, 5.0; Beef extract, 1.5; Yeast extract, 1.5; Agar, 20.0 at pH 7.5 ± 0.2. However, these bacteria were grown in nutrient broth with the composition same as that of nutrient agar (without agar in it).
Preparation of sample for shikimic acid and growth estimation
Samples withdrawn after the desired incubation period were subjected to centrifugation at 1,000 rpm for 2 min in a refrigerated centrifuge (SIGMA 4K15 Germany). After centrifugation, CaCO3 present in the sample settled down and the supernatant was withdrawn with the help of micro pipette. The supernatant thus obtained was now observed for its absorbance making the appropriate dilution with the distilled water. The absorbance was read at 660 nm against control using spectrophotometer (UV–Visible 1700, Shimadzu Corp., Kyoto, Japan).
Analysis of shikimic acid
Shikimic acid produced was estimated by high performance liquid chromatography (HPLC, Shimadzu Corp., Kyoto, Japan) on Aminex HPX-87H (300 × 7.8 mm) column, refractive index detector at an oven temperature of 55 °C and 5 mM H2SO4 as the mobile phase. The supernatant obtained after centrifugation at 10,000 rpm, was filtered by 0.22 µ (mdi, USA) and 20 µl of each test sample was run on HPLC.
Experimental design
Evaluation of medium components by Plackett–Burman approach
A sequential optimization strategy based on the statistical experimental design was implemented to increase shikimic acid production. Plackett–Burman (1946) experimental design was attempted to investigate the significance of various medium components on shikimic acid production. Here, seven variables were tested in 12-set of experiments. Two levels of each variable were used: −1 for low level and +1 for high level in the Plackett–Burman design matrix (Table 1). The main effect of each variable was calculated as the difference between the average of measurements made at high level (+1) and low level (−1) of that factor.
Plackett–Burman experimental design is based on the first order model (Eq. 1):
where, Y is the predicted response (shikimic acid yield), β0 and β1 are the constant co-efficients and x1 is the coded independent variable factor.
This model is used to screen and evaluate important factors that influence the response (shikimic acid production). The regression analysis of the variables leads to the selection of factors which may have significant effect on shikimic acid production by use of RSM.
Table 2 shows the design matrix of the variables in coded units together with the experimental results of shikimic acid produced (g/L). All the experiments were performed in triplicates and the average of the observations was used as the response of the design.
Optimization of shikimic acid production using response surface methodology
Response surface methodology was adopted for improving shikimic acid production using Design-Expert Version 6.0.10, Stat-Ease Inc. Minneapolis, USA software; to find the interaction between the significant variables (obtained from Plackett–Burman experiments). The FCCCD of response surface method was used to obtain data that fills a full second order polynomial model. The coded terms and actual values are presented in Table 3. Here, the independent variables were screened in 32 combinations according to the design matrix as shown in Table 4 and regression analysis was done for the data obtained. A second order polynomial equation was used to fit the data by multiple regression procedure. The three dimensional graphical representation of model equation represents the individual and interactive effect of the test variables on the response (shikimic acid yield). For a five factor system, the model equation is:
where, Y is the predicted response, β0 model constant, A, B, C, D and E are independent variables, β1, β2, β3, β4 and β5 are linear coefficients, β12, β13, β14, β15, β23, β24, β25, β34, β35 and β45 are cross product coefficients and β11, β22, β33, β44 and β55 are the quadratic coefficients.
The proportion of variance explained by the polynomial models obtained was given by multiple co-efficient of determination, R2. The fitted polynomial equation was expressed as three dimensional response surface plots to find the concentration of each factor for maximum shikimic acid production. The relationship between the responses and the experimental levels of each factor used in the design was also analyzed. To optimize level of each factor for maximum response, numerical optimization process was employed. The combination of different optimized parameters (which gave maximum shikimic acid yield), was tested experimentally to validate the model.
Results obtained from the statistically designed experiments were subjected to standard regression analysis. Design Experiment (6.0.10) was used for data analysis, determination of coefficients, as well as polynomial model reduction. Factors having highest t value and confidence level over 95 % were considered to be highly significant on shikimic acid production.
Fed-batch of carbon and nitrogen sources for maximum shikimic acid production
Increase in the utilization of carbon and nitrogen sources during the fermentation run may result in an increased production. Thus, with the view to further enhance shikimic acid production, fed-batch studies were carried out using RSM optimized production medium.
Selection of time of fed
To begin with the fed-batch studies, it is necessary to find out the exact time of fed during the fermentation run to increase shikimic acid production yield. Thus, shikimic acid production medium containing 20 g/L of initial glucose concentration was fed with 5 g/L of glucose at different time intervals i.e. 12, 24 and 36 h of incubation, in individual sets.
Optimization of glucose (feed) concentration
The concentration of the feed may also affect the production level of any biomolecule. Therefore, it was of utmost importance to find out the optimal concentration of glucose (carbon source), which can result in an increased production of shikimic acid. Keeping this in mind, different concentrations of glucose (5, 10 and 15 g/L) were fed to the production medium at the selected fed time.
Evaluation of second fed of glucose in the production medium
On realising the fact that glucose plays a very important role in the synthesis of shikimic acid, the effect of second fed of glucose on shikimic acid production was evaluated. Wherein, the shikimic acid production medium was again fed with the selected concentration of glucose at the time when the concentration of glucose already present in the medium reaches to half of its initial concentration.
Effect of nitrogen source (asparagine) fed
Nitrogen source affects the growth of the microorganism and thus certainly affects the production of the biomolecule. Realising this, the effect of asparagine fed on shikimic acid production was analysed. The production medium was fed with 10 g/L asparagine at the selected fed time, in combination with glucose (selected concentration).
All the experiments were conducted in triplicate and results are presented along with the standard deviation.
Scale up of shikimic acid production in 14 L bioreactor
Scale up of shikimic acid production was carried out in a 14 L bioreactor (Bioferm-LS2, Scigenics, India Pvt. Ltd.) with 7.5 L of the RSM optimized production medium. The optimized medium was sterilized in situ at 110 °C for 20 min. The medium was inoculated with 10 % (v/v) of the seed inoculum (OD at 660 nm 0.6–0.8) and fermentation was carried out at 30 °C for 72 h. Foaming was controlled by adding silicon antifoam agent (0.1 ml of 10 %, v/v prepared in distilled water).The impeller speed was adjusted to 200 rpm. Samples (10 ml) were withdrawn periodically at an interval of 6 h and analyzed for shikimic acid production. The fermentation parameters, such as temperature, pH and DO were continuously monitored using microprocessor-controlled probes.
Antibacterial activity of shikimic acid
The antibacterial activity of shikimic acid was evaluated by Agar Well Diffusion Technique as described by Agarry et al. (2005). Here, melted and cooled sterile Nutrient Agar media was poured into sterilized Petri dishes and allowed to solidify. Using the sterilized cotton swabs, overnight broth culture of the each of test organisms was swabbed on the nutrient agar plates uniformly. Wells (5 mm diameter) were cut on the inoculated nutrient agar plates with the aid of a sterile stainless steel cork borer. About 0.1 ml of different concentrations (1, 3 and 5 mg/ml) of purified shikimic acid were filled into each of the wells. These plates were then incubated at 37 ± 0.2 °C. Zones of inhibition (mm) were recorded after 24 h of incubation. The presence of zone of inhibition was considered as the presence of antibacterial activity.
Results and discussion
Selection of significant factors by Plackett–Burman experimental design
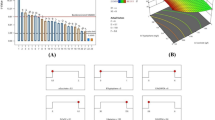
Plackett–Burman design was adopted to select the most significant factors responsible for higher shikimic acid production. Table 2 shows the design matrix along with the response of different experimental trials. However, it is clear from the regression analysis of the design matrix (Fig. 1) that among all the factor analysed, glucose, asparagine, KH2PO4 and CaCO3 showed the most significant effect(s) as compared to the effect of other three factors (MgSO4, agitation rate and inoculum level). Thus, these four factors (glucose, asparagine, KH2PO4 and CaCO3) were selected as the significant factors for shikimic acid production for further optimization by RSM. Further, on the basis of p values (<0.05), agitation rate was also selected as one of the significant factors. These results are in coherence with the statement of Sawale and Lele (2009) and Wee et al. (2009), where they stated “A large co-efficient (main effect) either positive or negative indicates that a factor has a large impact on response and a co-efficient (main effect) closer to zero means that a factor has little or no effect”.
Optimization of shikimic acid production by response surface methodology
Optimum levels of glucose, asparagine, KH2PO4, CaCO3 & agitation rate and the effect of their interactions on shikimic acid production were determined by FCCCD. Table 5 shows the details of the FCCCD design matrix employed in the RSM along with the predicted and observed responses for shikimic acid production (Y). Second order regression equation provided the levels of shikimic acid production as a function of initial values of glucose, asparagine, KH2PO4, CaCO3 and agitation rate, which can be predicted by the following Eq. (3)
where, A, B, C, D and E are glucose, KH2PO4, asparagine, CaCO3 and agitation rate, respectively.
Table 6 shows the ANOVA results for the RSM quadratic equation for response. ANOVA for shikimic acid production (Y, g/L) indicated the ‘F value’ to be 140.33, which implies that the model is significant. Model terms having values of ‘Prob > F’ less than 0.05 are considered significant, whereas those, greater than 0.10 are insignificant. The ‘Lack of Fit value’ of 2.86 implies that lack of fit is non significant relative to pure error and that the model fits. This is in agreement with the fact that, high F-value and non-significant lack of fit indicates the model to be a good fit (Isar et al. 2007). ANOVA indicated the R2 value of 0.9961 for response (shikimic acid yield). This again ensured a satisfactory adjustment of the quadratic model and could explain 99 % response variability. This is supported by the statement of Burkert et al. (2004) where, they stated that, for a good statistical model, the R2 value should be closer to one. The adequate precision was 47.799, which measures the “signal to noise” ratio. This indicates an adequate signal of accuracy as in this case a ratio of >4 is desirable (Chauhan and Gupta 2004). This model can be used to navigate the design space for the response. The ‘Pre R-Squared’ of 0.9263 is in reasonable agreement with ‘Adjusted R Squared’ of 0.9890 for the response. A good correlation between observed and predicted results reflected the accuracy and applicability of face centered central composite design for process optimization. The optimum levels of each variable for maximum shikimic acid production were determined by the three dimensional response surface plots (Fig. 2a–c), which were constructed by plotting the response (shikimic acid production) on Z-axis against any two independent variables, while maintaining the other variables at their optimum levels. Each plot represents an infinite number of combinations of two test variables with the other variables maintained at their respective central values.
Fed-batch of carbon and nitrogen sources for maximum shikimic acid production
Selection of time of fed
It was observed that on feeding the medium with glucose, in the exponential phase (12 h) a higher amount of shikimic acid (14.69 g/L) was produced. This amount of shikimic acid (14.69 g/L) is approximately 22 fold higher than the value obtained in the initial un-optimized production medium. On the other hand when glucose was fed at 6 h of incubation (early exponential phase) and 18 h of incubation (mid exponential phase) shikimic acid production declined (Fig. 3a–c). It is supported by the fact that bacteria utilise more substrate in the exponential phase. Thus, also helps in enhancing the production of the biomolecule.
Optimization of glucose (feed) concentration
Based on the glucose consumption rate data in batch process, the glucose nutrient feeding strategy was designed. Optimization of glucose concentration as a feed shows that 10 g/L is the optimal glucose concentration as it resulted in the production of 16.21 g/L of shikimic acid. However, on using 5 and 15 g/L glucose as the feed the amount of shikimic acid produced was comparatively less (Fig. 4a–c). This may be because 5 g/L glucose is lower than the concentration required for higher shikimic acid production while 15 g/L glucose is higher. Similarly, Draths et al. (1999) reported that increasing glucose availability in the medium might repress shikimic acid production.
Evaluation of second feed of glucose in the production medium
Results presented in Fig. 5 show that the second feed of glucose i.e. 10 g/L (selected during the optimization of glucose feed concentration) at 24 h resulted in the production of 18.64 g/L of shikimic acid which is approximately 1.5 fold higher than the value obtained during the single fed of glucose (10 g/L) in the production medium (16.21 g/L).
Effect of nitrogen source (asparagine) feed on shikimic acid production
It can be inferred from Fig. 6 that the addition of asparagine had a positive effect on shikimic acid production when used at a concentration of 5, 10 or 15 g/L at 24 h along with the fed of 10 g/L glucose at 12 h and 24 h. Although, the residual amount of asparagine could not be estimated. However, on the basis of shikimic acid production, it was inferred that 10 g/L is the optimal concentration of asparagines, resulting in the production of 22.32 g/L of shikimic acid in 48 h. On the other hand, only 19.87 and 20.06 g/L of shikimic acid was produced when 5 and 15 g/L of asparagine was used (respectively) in 12 h.
Increase in the amount of shikimic acid produced after the feed of glucose and asparagine is in accordance with Shukla et al. (2012) and Lin et al. (2008) where they stated that cell growth and organic acid production rates decrease with increase in carbon and nitrogen source concentrations in the fermentation medium. Thus, providing glucose in multiple steps at different incubation time of the run might have increased the amount of shikimic acid produced during the fermentation run in comparison to the amount produced while putting the total amount of glucose together initially.
Scale up of shikimic acid production in 14 L bioreactor
The optimal conditions obtained after process optimization were finally examined and verified for its scale up in a 14 L bioreactor with a working volume of 7.5 L (Fig. 7). Shikimic acid production started at 6 h, by this time, the organism enters into the exponential phase. Under these conditions, a maximum of 20.15 g/L of shikimic acid was produced with the productivity of 0.42 g/L/h, in the late exponential phase (48 h). Moreover the DO level was uncontrolled during the run but was still monitored during each sampling and it was found that at 48 h (when shikimic acid content was maximum), the DO level in the medium declined to a lower level of 30 %. Shikimic acid yield during the scale up of the process is 0.518 mol/mol, which is higher than the yield obtained by Escalante et al. (2010) i.e. 0.42 mol/mol while using the recombinant strain of E. coli. Further, the most probable reason for the successful scale up of the process is the controlled condition throughout the fermentation run. Bioreactor studies clearly show that maximum growth (OD at 660 nm 8.76) and shikimic acid production (20.12 g/L) was obtained in 48 h. This was followed by the stationary phase wherein, a steep decline in shikimic acid production was observed.
Antibacterial activity of shikimic acid
Observations presented in Table 7 clearly reveal that purified shikimic acid showed a varying degree of inhibition against the pathogenic bacteria tested at different concentrations. It is also evident that the level of inhibition increased with the increase in concentrations of shikimic acid with higher inhibition against Gram negative bacteria. Thus, it can be inferred from these results that shikimic acid may also be used as an antibiotic against different pathogenic bacteria.
Exhibition of a higher inhibition of the Gram negative bacteria as compared to the Gram positive bacteria by shikimic acid is supported by the fact that there is a thinner peptidoglycan layer in Gram negative bacteria. However, the peptidoglycan layer in Gram positive bacteria is comparatively thicker.
Conclusion
In the present study, the sequential statistical strategies, Plackett–Burman design followed by RSM were used successfully to find the optimum values of the significant factors for achieving maximum shikimic acid production. Further, fed-batch studies were also performed to increase the shikimic acid yield. These experiments resulted in the production of 22.32 g/L of shikimic acid which is approximately, 34-fold higher than the initial un-optimized medium. Moreover, the yield of shikimic acid obtained through this process in the shake flask conditions is 0.57 mol/mol. This is quite a good yield with reference to mol/mol yield as this yield is even higher than the yield obtained by Escalante et al. (Escalante et al. 2010). Subsequent scale up to a 14 L bioreactor resulted in the production of 20.15 g/L of shikimic acid. The present investigation may provide a significant tool for providing information on the use of different statistical approaches for maximum production of shikimic acid from the micro-organisms. Thus, the study is worthwhile for the statistical optimization of shikimic acid. Further, it is also showed in the results that shikimic acid may also be used as an antibiotic agent.
References
Agarry OO, Olaleye MT, Bello-Michael CO (2005) Comparative Antimicrobial Activities of Aloe vera gel and leaf. Afr J Biotechnol 4(12):1413–1414
Ahn JO, Lee HW, Saha R, Park MS, Jung JK, Lee DY (2008) Exploring the effects of carbon sources on the metabolic capacity for shikimic acid production in Escherichia coli using In silico metabolic prediction. J Microbiol Biotechnol 18(11):1773–1784
Beigel J, Bray M (2008) Current and future antiviral therapy of severe seasonal and avian influenza. Antiviral Res 78(1):91–102
Bongaerts J, Kramer M, Muller U, Raeven L, Wubbolts M (2001) Metabolic engineering for microbial production of aromatic amino acids and derived compounds. Metab Eng 3:289–300
Bradley D (2005) Star role for bacteria in controlling flu pandemic. Nat Rev Drug Discov 4(12):945–946
Burkert JF, Maureri MF, Rodrigues MI (2004) Optimization of extracellular lipase production by Geotricum sp. using factorial design. Bioresour Technol 91:77–84
Chandran SS, Yi J, Draths KM, von Daeniken R, Weber W, Frost JW (2003) Phosphoenolpyruvate availability and the biosynthesis of shikimic acid. Biotechnol Prog 19:808–814
Chauhan B, Gupta R (2004) Application of statistical experimental design for optimization of alkaline protease production from Bacillus sp. RGR-14. Process Biochem 39:2115–2122
Draths KM, Knop DR, Frost JW (1999) Shikimic acid and quinic acid: replacing isolation from plant sources with recombinant microbial biocatalysis. J Am Chem Soc 121:1603–1604
Escalante A, Calderon R, Valdiviya A, de Anda R, Hernandez G, Ramirez OT, Gosset G, Bolivar F (2010) Metabolic engineering for the production of shikimic acid in an evolved Escherichia coli strain lacking the phosphoenolpyruvate: carbohydrate phosphotransferase system. Microb Cell Factories 9:21
Evans IA, Osman MA (1974) Carcinogenicity of bracken and shikimic acid. Nature 250:348–349
Gibson MI, Gibson F, Doy CH, Morgan PN (1962) The branch point in the biosynthesis of the aromatic amino-acids. Nature 195:1173
Isar J, Agarwal L, Saran S, Kaushik R, Saxena RK (2007) A statistical approach to study the interactive effects of process parameters on succinic acid production from Bacteroides fragilis. Anaerobe 13:50–56
Johansson L, Liden G (2006) Transcriptone analysis of a shikimic acid producing strain of Escherichia coli W3110 grown under carbon- and phosphate-limited conditions. J Biotechnol 126:528–545
Johansson A, Lindskog G, Silversparre CC, Nielson KF, Liden G (2005) Shikimic acid production by a modified strain of Escherichia coli (W3110.shik1) under phosphate limited and carbon limited conditions. Biotechnol Bioeng 92:541–552
Kim CU, Lew W, Williams MA, Liu H, Zhang L, Swaminathan S, Bischhofberger N, Chen MS, Mendel DB, Tai CY, Laver WG, Stevens RC (1997) Influenza neuraminidase inhibitors possessing novel hydrophobic interaction in the enzyme active site: design, synthesis and structural analysis of carbocyclic sialic acid analogues with potent anti-influenza activity. J Am Chem Soc 119:681–690
Kim CU, Lew W, Williams MA, Wu H, Zhang L, Chen X, Escarpe PA, Mendel DB, Laver WG, Stevens RC (1998) Structure-activity relationship studies of novel carbocyclic influenza neuraminidase inhibitors. J Am Chem Soc 41:2451–2460
Kramer M, Bongaerts J, Bovenberg R, Kremer S, Muller U, Orf S, Wubbolts M, Raeven L (2003) Metabolic engineering for microbial production of shikimic acid. Metab Eng 5:277–283
Lagace-Wiens PRS, Rubinstein E (2010) Influenza epidemiology—past, present, and future. Crit Care Med 38(4):e1–e9
Li K, Mikola MR, Draths KM, Worden RM, Frost JW (1999) Fed-batch fermenter synthesis of 3-dehydroshikimic acid using recombinant Escherichia coli. Biotechnol Bioeng 64(1):61–73
Lin SKC, Du C, Koutinas A, Wang R, Webb C (2008) Substrate and product inhibition kinetics in succinic acid production by Actinobacillus succinogenes. Biochem Eng J 41:128–135
Plackett RL, Burman JP (1946) The design of optimum multifactorial experiments. Biometrica 33:305–325
Racaniello V (2009) Sturcture of influenza virus. www.virology.ws/2009/04/30/structure-of-influenza-virus/
Rohloff JC, Kent KM, Postich MJ, Becker MW, Chapman HH, Kelly DE, Lew W, Louie MS, McGee LR, Prisbe EJ, Schultze LM, Yu RH, Zhang L (1998) Practical total synthesis of the anti-influenza drug GS-4104. J Org Chem 63(13):4545–4550
Sawale SD, Lele SS (2009) Increased dextransucrase production by response surface methodology from Leuconostoc sp. Isolated from fermentated idli batter. J Biotechnol Biochem 4(2):160–167
Saxena RK, Tripathi P, Rawat G (2012) Pandemism of swine flu and its prospective drug therapy. Eur J Clin Microbiol Infect Dis 31(12):3265–3279
Shirai M, Miyata R, Sasaki S, Sakamoto K, Yahanda S, Shibayama K, Yonehara T, Ogawa K (2001) Micro-organism belonging to the genus Citrobacter and process for producing shikimic acid. European patent, EP 1 092 766 A1
Shukla R, Pandey D, Surendranathan D, Dubey AK (2012) Organic acid production by Corynebacterium vitaeruminis. Ind J Fund Appl Life Sci 2(2):36–41
Stavric B, Stolz DR (1976) Shikimic acid. Food Cosmet Toxicol 14(2):141–145
Tripathi P, Rawat G, Yadav S, Saxena RK (2013) Fermentative production of shikimic acid: a paradigm shift from plant route to microbial route. Bioprocess Biosys Eng. doi:10.1007/s00449-013-0940-4
Wee YJ, Reddy LVA, Chung KC, Ryu HW (2009) Optimization of chitosanase production from Bacillus sp. RKY3 using statistical experimental designs. J Chem Technol Biotechnol 84:1356–1363
Acknowledgments
Authors wish to acknowledge the financial support of Indian Council of Medical Research (ICMR) Project sanction to RKS. Authors also wish to acknowledge the fellowship of Jawaharlal Nehru Memorial Fund (JNMF) to carry out this work. The infrastructural support provided by Technology Based Incubator (TBI), UDSC, India, is highly acknowledged.
Conflict of interest
Authors declare that there is no conflict of interest in this publication.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tripathi, P., Rawat, G., Yadav, S. et al. Shikimic acid, a base compound for the formulation of swine/avian flu drug: statistical optimization, fed-batch and scale up studies along with its application as an antibacterial agent. Antonie van Leeuwenhoek 107, 419–431 (2015). https://doi.org/10.1007/s10482-014-0340-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-014-0340-z