Abstract
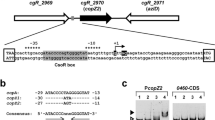
Metal responsive MerR family transcriptional regulators are widespread in bacteria and activate the transcription of genes involved in metal ion detoxification, efflux, or homeostasis, in response to the presence of cognate metal species in the cytoplasm. MerR family regulators recognize and bind to dyad symmetrical DNA sequences in specific promoters that have a spacer region between the –35 and –10 sequences which is longer than the canonical 16–18 bp spacer for other σ70–dependent promoters. In this study we report β-galactosidase assays of MerR family-regulated gene expression in the multiple metal resistant bacterium Cupriavidus metallidurans. A series of pMU2385 reporter plasmid derivatives containing cloned MerR family-activated promoters were used to determine metal ion-induced responses from different MerR family regulated promoters, as well as regulators cloned with the cognate promoter into pMU2385. Mercuric ion-responsive MerR and lead ion-responsive PbrR activity was confirmed using this assay system as well as MerR family activator activity on heterologous promoters PcopA, PcadA, and Pzcc from Escherichia coli, Pseudomonas aeruginosa and Bordetella pertussis, respectively. In C. metallidurans CH34, transcription from these promoters was activated by MerR family regulators encoded on the chromosome or megaplasmids in response to copper (PcopA), and lead (PcadA and PzccA), showing that MerR family activators in C. metallidurans can act on MerR family promoters from other organisms, which have sequence differences to the predicted C. metallidurans promoters.




Similar content being viewed by others
References
Borremans B, Hobman JL, Provoost A, Brown NL, van der Lelie D (2001) Cloning and functional analysis of the pbr lead resistance operon of Ralstonia metallidurans CH34. J Bacteriol 183:5651–5658. doi:10.1128/JB.183.19.5651-5658.2001
Brown NL, Stoyanov JV, Kidd SP, Hobman JL (2003) The MerR family of transcriptional regulators. FEMS Microbiol Rev 27:145–163. doi:10.1016/S0168-6445(03)00051-2
Brocklehurst KR, Hobman JL, Lawley B, Blank L, Marshall SJ, Brown NL, Morby AP (1999) ZntR is a Zn(II)-responsive MerR-like transcriptional regulator of zntA in Escherichia coli. Mol Microbiol 31:893–902. doi:10.1046/j.1365-2958.1999.01229.x
Brocklehurst KR, Megit SJ, Morby AP (2003) Characterisation of CadR from Pseudomonas aeruginosa: a Cd(II)-responsive MerR homologue. Biochem Biophys Res Commun 308:234–239. doi:10.1016/S0006-291X(03)01366-4
Chen P, Greenberg B, Taghavi S, Romano C, van der Lelie D, He C (2005) An exceptionally selective lead (II)-regulatory protein from Ralstonia metallidurans: Development of a fluorescent lead (II) probe. Angew Chem Int Ed 44:2715–2719. doi:10.1002/anie.200462443
Chen PR, Wasinger EC, Zhao J, van der Lelie D, Chen LX, He C (2007) Spectroscopic insights into lead (II) coordination by the selective lead(II)-binding protein PbrR691. J Am Chem Soc 129:12350–12351. doi:10.1021/ja0733890
Corbisier P, van der Lelie D, Borremans B, Provoost A, de Lorenzo V, Brown NL, Lloyd JR, Hobman JL, Csoregi E, Johansson G, Mattiasson B (1999) Whole cell and protein-based biosensors for the detection of bioavailable heavy metals in environmental samples. Anal Chim Acta 387:235–244. doi:10.1016/S0003-2670(98)00725-9
Diels L, Faelen M, Mergeay M, Nies D (1985) Mercury transposons from plasmids governing multiple resistance to heavy metals in Alcalignes eutrophus CH34. Arch Int Physiol Biochim 93:27–28. doi:10.3109/13813458509080622
Foster TJ, Ginnity F (1985) Some mercurial resistance plasmids from different incompatibility groups specify merR regulatory functions that both repress and induce the mer operon of plasmid R100. J Bacteriol 162:773–776
Groβe C, Grass G, Anton A, Franke S, Santos AN, Lawley B, Brown NL, Nies DH (1999) Transcriptional organization of the czc heavy-metal homeostasis determinant from Alcaligenes eutrophus. J Bacteriol 181:2385–2393
Hobman JL (2007) MerR family transcription activators: similar designs, different specificities. Mol Microbiol 63:1275–1278
Hobman JL, Wilkie J, Brown NL (2005) A design for life: prokaryotic metal-binding MerR family regulators. Biometals 18:429–436
Kidd SP, Brown NL (2003) ZccR- a MerR-like regulator from Bordetella pertussis, which responds to zinc, cadmium and cobalt. Biochem Biophys Res Comm 302:697–702
Mergeay M, Houba C, Gerrits J (1978) Extrachromosomal inheritance controlling resistance to cadmium, cobalt, copper and zinc ions: evidence from curing in a Pseudomonas. Arch Int Physiol Biochem 86:440–442
Mergeay M, Nies D, Schlegel HG, Gerits J, Charles P, van Gijsegem F (1985) Alcaligenes eutrophus CH34 is a facultative chemolithotroph with plasmid-bound resistance to heavy metals. J Bacteriol 162:328–334
Mergeay M, Monchy S, Vallaeys T, Auquier V, Benotmane A, Bertin P, Taghavi S, Dunn J, van der Lelie D, Wattiez R (2003) Ralstonia metallidurans, a bacterium specifically adapted to toxic metals: towards a catalogue of metal-responsive genes. FEMS Microbiol Rev 27:385–410
Miller J (1972) Experiments in molecular genetics. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Monchy S, Benotmane MA, Wattiez R, van Aelst S, Auquier V, Borremans B, Mergeay M, Taghavi S, van der Lelie D, Vallaeys T (2006a) Transcriptomic and proteomic analyses of the pMOL30-encoded copper resistance in Cupriavidus metallidurans strain CH34. Microbiology 152:1765–1776
Monchy S, Vallaeys T, Bossus A, Mergeay M (2006b) Metal transport ATPase genes from Cupriavidus metallidurans CH34: a transcriptomics approach. Intern J Environ Anal Chem 86:677–692
Monchy S, Benotmane MA, Janssen P, Vallaeys T, Taghavi S, van der Lelie D, Mergeay M (2007) Plasmids pMOL28 and pMOL30 of Cupriavidus metallidurans are specialized in the maximum viable response to heavy metals. J Bacteriol 189:7417–7425
Nies DH (1992) CzcR and CzcD, gene products affecting regulation of resistance to cobalt, zinc, and cadmium (czc system) in Alcaligenes eutrophus. J Bacteriol 174:8102–8110
Outten FW, Outten CE, Hale J, O’Halloran TV (2000) Transcriptional activation of an Escherichia coli copper efflux regulon by the chromosomal MerR homologue, CueR. J Biol Chem 275:31024–31029
Parkhill J, Lawley B, Hobman JL, Brown NL (1998) Selection and characterization of mercury-independent activation mutants of the Tn501 transcriptional regulator, MerR. Microbiology 144:2855–2864
Permina EA, Kazakov AE, Kalinina OV, Gelfand M (2006) Comparative genomics of regulation of heavy metal resistance in eubacteria. BMC Microbiol 6:49
Praszkier J, Wilson IW, Pittard AJ (1992) Mutations affecting translational coupling between the rep genes of an IncB miniplasmid. J Bacteriol 174:2376–2383
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Sarkar SK, Andoy NM, Benítez JJ, Chen PR, Kong JS, He C, Chen P (2007) Engineered Holliday junctions as single-molecule reporters for protein–DNA interactions with application to a MerR-family regulator. J Am Chem Soc 129:12461–12467
Stoyanov JV, Hobman JL, Brown NL (2001) CueR (YbbI) of Escherichia coli is a MerR family regulator controlling expression of the copper exporter CopA. Mol Microbiol 39:502–511
Stoyanov JV, Brown NL (2003) The Escherichia coli copper-responsive copA promoter is activated by gold. J Biol Chem 278:1407–1410
Taghavi S, van der Lelie D, Mergeay M (1994) Electroporation of Alcaligenes eutrophus with (mega) plasmids and genomic DNA fragments. Appl Environ Microbiol 60:3585–3591
Taghavi S, Lesaulnier C, Monchy S, Wattier R, Mergeay M, van der Lelie D (2009) Lead(II) resistance in Cupriavidus metallidurans CH34: interplay between plasmid and chromosomally located functions. Anthonie van Leeuwenhoek. doi:10.1007/s10482-008-9289-0
Vieira J, Messing J (1991) New pUC-derived cloning vectors with different selectable markers and DNA replication origins. Gene 100:189–194
Acknowledgments
This work was supported by Biotechnology and Biological Sciences Research Council grant B10333 and a BBSRC studentship to DJJ. The Birmingham Functional Genomics laboratory was supported by the Biotechnology and Biological Sciences Research Council Joint Infrastructure Fund grant JIF13209.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Julian, D.J., Kershaw, C.J., Brown, N.L. et al. Transcriptional activation of MerR family promoters in Cupriavidus metallidurans CH34. Antonie van Leeuwenhoek 96, 149–159 (2009). https://doi.org/10.1007/s10482-008-9293-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-008-9293-4