Abstract
A plethora of studies have offered crucial insights in the phylogeographic status of Western Palearctic bird species. However, an overview integrating all this information and analyzing the combined results is still missing. In this study, we compiled all published peer-reviewed and grey literature available on the phylogeography of Western Palearctic bird species. Our literature review indicates a total number of 198 studies, with the overwhelming majority published as journal articles (n = 186). In total, these literature items offer information on 145 bird species. 85 of these species are characterized by low genetic differentiation, 46 species indicate genetic variation but no geographic structuring i.e. panmixia, while 14 species show geographically distinct lineages and haplotypes. Majority of bird species inhabiting the Western Palearctic display genetic admixture. The glaciation cycles in the past few million years were pivotal factors in shaping this situation: during warm periods many species expanded their distribution range to the north over wide areas of Eurasia; whereas, during ice ages most areas were no longer suitable and species retreated to refugia, where lineages mixed.
Zusammenfassung
Gemeinsame Muster in der molekularen Phylogeographie der westpaläarktischen Vögel: Ein umfassender Überblick
Eine Fülle von genetischen Untersuchungen konnte entscheidende Einblicke in den phylogeographischen Status der westpaläarktischen Vogelarten geliefert. Es fehlt jedoch immer noch eine Übersicht, die all diese Informationen integriert und die Ergebnisse gemeinsam analysiert. In dieser Studie haben wir die gesamte veröffentlichte, begutachtete und graue Literatur zur Phylogeographie westpaläarktischer Vogelarten zusammengestellt. Unsere Literaturübersicht kommt auf eine Gesamtzahl von 198 Untersuchungen, wobei die überwiegende Mehrheit als Zeitschriftenartikel veröffentlicht wurde (n = 186). Insgesamt bieten diese Publikationen Informationen zu 145 Vogelarten. 85 dieser Arten zeichnen sich durch eine geringe genetische Differenzierung aus, 46 Arten zeigen genetische Variation, aber keine geografische Strukturierung, d. h. Panmixie, während nur14 Arten geografisch unterschiedliche Linien und Haplotypen aufweisen. Die Mehrheit der Vogelarten, die die Westpaläarktis besiedeln, zeigen eine genetische Vermischung. Die Vergletscherungszyklen der letzten paar Millionen Jahre waren ausschlaggebend für diese Situation: Während Warmzeiten dehnten viele Arten ihr Verbreitungsgebiet über weite Teile Eurasiens nach Norden aus; während der Eiszeiten waren die meisten Gebiete zur Besiedlung nicht mehr geeignet und die Arten zogen sich in Refugien zurück. Diese klimatischen Zyklen und periodischen Verbreitungsänderungen innerhalb der letzten zwei Millionen Jahre führten zur Durchmischung der genetischen Entwicklungslinien, wie wir sie heute feststellen können.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Past climatic events are embedded in the DNA of organisms (Hewitt 2000). Across the paleogeographic areas of the world, local fauna shares molecular evidence indicating how they responded to the major successions of cold and warm periods (Webb and Bartlein 1992). In the past three decades, the emergence of fine-tuned molecular techniques triggered a renaissance in zoology. Based on DNA sequencing, scientists were finally able to decipher the chronological and spatial evolution of species and subspecies (Hewitt 1999). With the possibility of assessing the progress of a species in both time and space, the field of phylogeography emerged (Avise et al. 1987). This served as an unprecedented opportunity and soon after the first studies began to appear (Cwynar and MacDonald 1987; Martin and Simon 1990; Reeb and Avise 1990; Nevo and Beiles 1992; Prinsloo and Robinson 1992; Quinn 1992; Melnick et al. 1993). Initial studies had a limited focus, dealing with small sample sizes and generally employed a single molecular marker, most often a mitochondrial DNA (hereafter mtDNA) gene. Moreover, in the first few years, both software and statistical techniques were a crude limitation to data analysis (Edwards and Bensch 2009).
As advanced DNA sequencing techniques became widely available, phylogeography became more popular, and the field entered a period of formidable growth (Hickerson et al. 2010), with thousands of studies currently being available. One would expect that a considerable amount of work would also have been done in bringing together and interpreting this colossal volume of data. However, such reviews are very scarce, and they generally have a wide focus, from assessing the hypotheses behind the genetic lineages, to number of papers and species. To date, review articles have included the status of phylogeographic research for oceanic habitats at global level (Bowen et al. 2016), or for archipelagos (Shaw and Gillespie 2016). Reviews are also available for research on continental areas (Riddle 2016), or studies dealing exclusively with the Southern hemisphere (Beheregaray 2008). One review examined exclusively the terrestrial taxa in the Aegean archipelago and surrounding regions (Poulakakis et al. 2015). Among all the vertebrates which became a research focus in phylogeography, birds proved to be the most iconic (Weiss and Ferrand 2007).
Numerous avian species had their phylogeography revealed. In Eurasia, the bird families which received most attention are the raptors, especially the Accipitridae, the crows (Corvidae), flycatchers and wheatears (both in Muscicapidae), as well as species from Phasianidae and the waders (Scolopacidae). To date, surprisingly few review and comparative studies have focused on birds. The first, authored by Robert M. Zink (Zink 1996), analyzed mtDNA geographic patterns of five North American bird species, to determine that their absence of genetic structure is linked to recent population expansion. The second, a book section in 1997, written by the same author (Zink 1997), offers an improved version of the first review, with more species included, which overall indicates the same population genetic structure. The third, in 1998, written by John C. Avise and DeEtte Walker (Avise and Walker 1998), compares 63 species from a phylogeographic and speciation perspective, confirming that the Pleistocene had a decisive effect on avian speciation. The fourth, as a chapter in the PhD thesis of Alexandra Pavlova (Pavlova 2004), has compared phylogeographic information of 28 Eurasian bird species, concluding that the South contains regions with higher genetic diversity and phylogeographic endemism, with the overall genetic structure being shaped by the post-Pleistocene recolonization of Eurasia. In 2009, a review focused on the phylogeography of birds from the Australo-Papuan region, indicated extensive paraphyly among birds in Australia (Joseph and Omland 2009). Following the chronological order, the next review brings into attention the horizontal and elevational phylogeographical patterns of Himalayan and Southeast Asian birds (Päckert et al. 2012). The seventh study collected phylogeographic data for 210 bird species in the New World, to reveal that species from lower latitudes have higher genetic diversity (Smith et al. 2017). The most recent study, authored by Alexey P. Kryukov (Kryukov 2019) has an exclusive focus on Palearctic corvid species and reviews published data on the natural hybrid zones between crow species.
To the best of our knowledge, no study aimed specifically at deciphering the avian phylogeography of the Western Palearctic has been published. This region has received extensive attention, triggered both by the geographic composition e.g. various islands promoting endemism, as well as the density of research groups and availability of funding. In this study, we compiled a comprehensive body of published scientific literature with a clear focus on phylogeography of bird species inhabiting the Western Palearctic.
Survey methodology
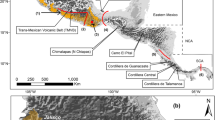
Between November 2018 and June 2020, we conducted an extensive search for relevant literature on: (1) ISI Web of knowledge, (2) Google Scholar, (3) Research Gate and (4) Google. We used the following keywords: “phylogeography” AND (“bird” OR “avian”) AND (“Western Palearctic” OR “Europe”). Our initial searches were made using English, French, Spanish, German and Russian, which are recognized as some of the most used languages for scientific publishing in our target region (Ammon 2001; Ammon and McConnell 2002). However, as we only found proper results in English, we excluded the other languages from further searches. Moreover, we considered both peer-reviewed and grey literature. We extracted additional papers from the references of the articles revealed by our systematic review, when the title or citation context indicated a bird phylogeography investigation. This study is limited to the area of the Western Palearctic, as presented in Shirihai & Svensson (Shirihai and Svensson 2018), consisting of Europe, North Africa, the Middle East and Asia Minor, the Cape Verde, Azores and Canary Islands, Madeira, Jan Mayen, Svalbard and Iceland. As the Eastern limit, we expanded until the Caspian Sea and the Ural Mountains. For taxonomy, we followed the IOC World Bird List (Gill et al. 2020).
The papers revealed by our on-line searches were further filtered according to the following criteria: (i) to focus on naturally occurring species in the Western Palearctic and (ii) to have samples originating from at least three geographically distinct populations. These facts were obtained upon reading the abstract plus materials and methods section. We later extracted information on the phylogeographic status of the studied species and assigned it one of the three categories: (i) panmixia, when the haplotypes are randomly distributed across the sampled area and no structure can be observed; (ii) low differentiation, if a certain degree of geographic delimitation of the haplotypes occur yet the lineage sorting is incomplete (e.g. Western vs. Eastern European haploclades, continental Europe vs. UK lineages) or (iii) geographically distinct lineages, for cases where certain haplotypes can be safely attributed to a geographic area and monophyletic groups are present (e.g. haplotypes found exclusively in one mountain range).
In addition, for each bird species, we noted the type of molecular technique (markers) used for obtaining the data and the migratory status of the respective species. The later information was retrieved from the IUCN Red List (IUCN 2019). We used the program R (R Core Team 2019) for data visualization.
To offer a better visualization of the phylogeographic differences among the three defined categories, we selected one species for each category, constructed its haplotype network and positioned all three networks side by side. We chose the European Turtle Dove (Streptopelia turtur) to illustrate panmixia, the European Green Woodpecker (Picus viridis) for low geographic differentiation and the African Blue Tit (Cyanistes teneriffae) as model for the geographically distinct lineages category. Complete details on the GenBank sequences used for the visualization are found in Table S4. Mainly, the sequences are derived from the following studies: (Calderón et al. 2016) for the dove, (Perktas et al. 2011) for the woodpecker and (Dietzen et al. 2008) for the tit. After downloading the sequences from GenBank, we grouped all files belonging to one species into a fasta file, using MEGA X (Kumar et al. 2018). We further assessed the number of haplotypes in DNA SP (Rozas et al. 2017) and finally employed the PopArt software (Leigh and Bryant 2015) to create the networks.
Results
In total, 145 bird species from the Western Palearctic have been the target of phylogeographic studies (Tables 1, 2, 3, Table S1). A number of 198 literature items (Table S2), including 186 peer-reviewed articles, one preprint, four PhD theses and one Master thesis, four articles in conference proceedings, and two book chapters matched our literature selection criteria. The year of publication ranges from 1993 until 2020. The Western Capercaillie (Tetrao urogallus) has been the focus of ten publications, which makes it the most investigated species in our dataset.
In terms of molecular markers, for the majority of bird species (132 out of 145) the choice has been mtDNA (Fig. 1), which is also one of the three markers used since the first studies in 1993. Secondly, microsatellites have been used for 36 species and nuclear DNA sequences (hereafter nuDNA) for 36 as well. Seven studies have employed sex chromosomes and a further seven studies used next-generation DNA sequencing (hereafter NGS). In the NGS category we included research with whole-genome sequencing, ddRAD sequencing and SNPs. One study (Resano-Mayor et al. 2017) has used hydrogen isotopes in combination with mtDNA. Furthermore, one study (Lagerholm et al. 2017) employed ancient DNA extracted from fossil bones. For each marker, the above-mentioned values consist of both the occasions where the respective marker has been applied alone or in combination with another marker.
Across the Western Palearctic, 85 avian species show signs of low genetic differentiation (Fig. 2), while 46 are genetically diverse but do not show a geographic structuring—indicating panmixia, and the remaining 14 species display geographically distinct lineages. Regarding the migratory behavior of the species comprising each category, we observed that the majority of the birds with low genetic differentiation and panmixia are migratory, while the species showing geographically distinct lineages are mainly resident and/or inhabitants of Oceanic islands (Fig. 2).
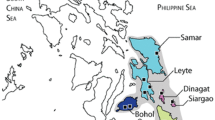
Our haplotype network comparison revealed substantial differences among the three selected species. To visualize the situation for birds with panmixia, we chose the European Turtle Dove (Streptopelia turtur). For species with low geographic differentiation, we selected the European Green Woodpecker (Picus viridis). We chose the African Blue Tit (Cyanistes teneriffae), as a model for the geographically distinct lineages category. Figure 3 (and in higher quality as figure S3) contains our visual comparison, which points to the differences in the distribution of haplotypes, among the three species. For the European Turtle Dove, all seven countries share haplotypes, regardless of the distance in between e.g. UK and Bulgaria. The network for the European Green Woodpecker indicates that several haplotypes are shared among the various populations, but some countries have specific haplotypes e.g. Italy, Turkey, Greece. It should be mentioned, that the Iberian population of the Green Woodpecker was found to be genetically different and has been consequently split as a new species Picus sharpei (Perktas et al. 2011; Pons et al. 2011). For the African Blue Tit, which is found in the Canary Islands, plus in Morocco, Algeria, Tunisia and Libya, the haplotype distribution is very clear. Majority of the islands in the Canary archipelago have distinct haplotypes, which are not found on the other islands. On the African continent, Libya has its own haplotypes, while Moroccan birds appear to be sharing some genetic background with birds from Fuerteventura. Complete details on the GenBank sequences used for the visualization are found in Table S4.
Comparative haplotype networks for the three species of birds, representative for each phylogeographic category: a European Turtle Dove (Streptopelia turtur) indicating panmixia, b European Green Woodpecker (Picus viridis) showing low geographic differentiation and c African Blue Tit (Cyanistes teneriffae) indicating geographically distinct lineages. The sequences are derived from the following studies: Calderón et al. 2016 (Calderón et al. 2016) for the dove (Cytb sequences), Perktaș et al. 2011 (Perktas et al. 2011) for the woodpecker (ND2 sequences) and Dietzen et al. 2008 (Dietzen et al. 2008) for the tit (Cytb sequences). This figure is available in higher quality as Figure S3
Discussion
Phylogeographical situation of Western Palearctic birds
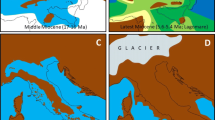
Substantial research has been carried out to understand the phylogeographic history of Western Palearctic avifauna, resulting in 145 bird species being studied. This represents roughly 20% of the 720 bird species found in the region (Shirihai and Svensson 2018). Our review indicates that majority of the species are characterized by similar patterns of genetic variation and admixture. This situation is inextricably connected to the climatic past of the Western Palearctic (Lisiecki and Raymo 2007), with the evidence being encrypted in the DNA of the species inhabiting this region (Taberlet et al. 1998; Weiss and Ferrand 2007). The periods mainly responsible for shaping the genetic background for the current avifauna are the Pleistocene and Pliocene (Rand 1948; Avise and Walker 1998; Zink et al. 2004). During these two eras, the climate oscillated between glaciations and warm cycles, accompanied by shifts in the composition of the vegetation (Frenzel et al. 1992). In turn, these oscillations led to massive bird population crashes at the arrival of each glaciation, or great population expansions, when the ice sheet retreated (Hewitt 1999). Subsequently, these demographic processes triggered population admixture both in the refugia (see Fig. 4) and at the contact zones (during population expansion times). As a result, the overwhelming majority of bird species in our dataset (i.e. 131 out of 145, or 90.3%) are characterized by high levels of genetic admixture, with either complete panmixia or low differentiation among various breeding populations.
Furthermore, the majority (i.e. 97 out of 131, or 74%) of species with genetic admixture are migrants or partial migrants (see Fig. 2). This suggests that the legacy of migration, which is strongly connected to the population expansion history in the Pleistocene and Pliocene (Bell 2000), had a central contribution to the species’ current genetic structure.
To better underline this situation, we also indicate that in our dataset, 12 out of the 14 species with geographically distinct genetic lineages are resident, which brings further evidence that movement patterns plays an important role in shaping genetic diversity. Our results, in conjunction with previous research, indicate that most of the migrants from warm areas are genetically mixed, while species better adapted to cold climates show less admixture. This implies that species like the Hazel Grouse, Western Capercaillie, and ptarmigans could have spent some of the ice ages in small mountain refugia, inside the ice sheet (Lagerholm et al. 2017) (see Fig. 4). From a different perspective, some steppe species (Garcia et al. 2011) had their maximum distribution during the glaciations, when much of today’s central and Southern Europe had scarce vegetation and resembled a steppe formation (Frenzel et al. 1992). Similarly, the genus Prunella has been shown to have colonized new areas mainly during the glacial periods (Liu et al. 2017). Regarding the migratory movements of birds in the past 50,000 years, a recent study proposed that Old World species had relatively short movements (Somveille et al. 2020). In comparison, the same simulation-based study shows that birds in the New World were already doing transcontinental migrations. Overall, this brings further evidence that, in the past thousands of years, migratory behavior had an important role in shaping today’s genetic background of bird species. Additional factors like the time of glacial isolation, habitat selection, variation of the ecological niche through time (Eyres et al. 2021), geographical barriers and hybridization have also been crucial in species’ phylogeographical structure (Avise 2000).
In the dataset we compiled, several bird species show two main mtDNA haploclades. The Eurasian Collared Dove (Bagi et al. 2018), Common Redstart (Hogner et al. 2012) and Red-backed Shrike (Pârâu et al. 2019) are characterized by two main haplotype clades, with no geographic structure. However, the Little Owl (Pellegrino et al. 2014) and Great Reed Warbler (Hansson et al. 2008), also with two main mtDNA groups, display Western and Eastern European specific haploclades. In addition, species like the Skylark (Zink et al. 2008) and Great Grey Shrike (Olsson et al. 2010) which are spread across the entire Palearctic also show two divergent clades: Western Palearctic versus East Asian individuals. Previous research on the North American continent has revealed a similar situation for the Snow Goose (Quinn 1992) and Common Raven (Webb et al. 2011), which show two haploclades, with a relatively clear geographic structure. The most plausible explanation for the occurrence of two or more haploclades are the glacial refugia (Weiss and Ferrand 2007). The thousand years spent in these Southern refugia e.g. Iberian Peninsula or the Balkans in Europe, have acted as a selective force on genetic lineages. During the cyclical back-and-forth population expansion processes associated with ice ages, only certain populations survived, became in contact, and interbred. Although in the past 12 thousand years there was no ice age in the Western Palearctic and birds from different refugia freely mixed, the genetic legacy of the cold ages is still deeply rooted in the DNA. Furthermore, several species still have refugia-specific haplotypes, like in the case of the Rook (Corvus frugilegus) (Salinas et al. 2021).
However, in terms of species with several haploclades or sister species sharing haplotypes, hybridization and introgression of gene flow also represent a valuable explanation. Avian hybridization has long been a point of interest for ornithologists and geneticists, as evidence for active speciation, or in simple terms—seeing in real-time how a species is borne (McCarthy 2006; Ottenburghs et al. 2015). Hybridization is known to occur in 9% of the bird species (Grant and Grant 1992). In these cases, genetic material from one species is incorporated into another, potentially enforcing speciation (Rheindt and Edwards 2011; Ottenburghs et al. 2017). In Europe, the classic example is the Italian Sparrow (Passer italiae), which still shares mtDNA haplotypes with its parent species, the Spanish Sparrow (Passer hispaniolensis) and the House Sparrow (Passer domesticus), representing an admixture of both genetic and phenotypic factors (Hermansen et al. 2011; Trier et al. 2014; Sætre et al. 2017). The dawn of the genomic era already provides strong molecular evidence into the complexity of genomic regions directly responsible for speciation, and promising a much better understanding in the years to come (Joseph 2018).
The Western Palearctic also includes a series of oceanic islands situated at low latitudes i.e. Macaronesia, which experienced different climate conditions during the Pliocene and Pleistocene (Webb and Bartlein 1992). These oceanic islands, which were not connected by land bridges with the continent were not affected by glaciations, which represents the triggering factor for genetic admixture in the avifauna of continental Western Palearctic (Wink 2018a). As a result, bird populations developed independently on these islands for several million years and acquired distinct genetic lineages (Avise and Walker 1998). Species like the Common Chaffinch (Rodrigues et al. 2014b), Common Blackbird (Rodrigues et al. 2016), European Robin (Dietzen et al. 2003, 2015; Rodrigues et al. 2013) and the Goldcrest (Rodrigues et al. 2014a) show island or archipelago unique haplotypes across Macaronesia, which are not shared with the continental conspecific population. Furthermore, due to recent molecular studies, island taxa which were previously considered subspecies have been elevated to species level: the Cape Verde Shearwater (Gómez-Díaz et al. 2006) and both the Tenerife and Gran Canaria Blue Chaffinches (Lifjeld et al. 2016; Sangster et al. 2016). Taking into consideration that the current taxonomic trend is to split species which have geographically distinct lineages, we expect to reach an increased number of approximately 18 thousand bird species (Barrowclough et al. 2016), from the 11 thousand currently recognized.
This circumstance of continental bird species being characterized by genetic admixture does not represent a special feature of the Western Palearctic avifauna, but rather the general situation for birds in the Northern temperate areas. Research on the North American continent has revealed similar patterns in several bird species (Zink 1996; Avise and Walker 1998; Dohms 2016), with most populations sharing haplotypes and only a handful of examples for geographically distinct genetic lineages.
For avian species in areas not affected by glaciations in the past few million years e.g. tropical areas and oceanic islands (see above), genetic structure and differentiation is, in many species, distinct. For example, the gnatcatchers and gnatwrens (Polioptilidae), pectoral sparrows (Arremon taciturnus), tyrant-manakins (Pipridae) and the Straight-billed Hermit (Phaethornis bourcieri) from South America show very distinct genetic groups, with almost no gene flow (Araújo-Silva et al. 2017; Capurucho et al. 2018; de Melo et al. 2018,2020; Smith et al. 2018). A similar situation has been described for the Wedge-billed Woodcreeper (Glyphorynchus spirurus) (Fernandes et al. 2013), the Southern Chestnut-tailed Antbird (Sciaphylax hemimelaena) (Fernandes et al. 2012), both Spotted and Spot-backed Antbirds (Hylophylax naevioides/ naevius) (Fernandes et al. 2014) and lowland antpittas (Grallariidae) (Carneiro et al. 2018). The consensus of the above-mentioned research is that genetic diversification in South American bird species was mainly triggered by the consolidation of Amazonian rivers and drainage system, which acted as dispersal barriers (Haffer 1969; Silva et al. 2019). These geological events took place during the Miocene and Pliocene (Rull 2011), which gave several million years of additional speciation for neotropical birds, in comparison to the birds from the Western Palearctic. Furthermore, the South American continent had a less fluctuating climate during these eras, a crucial factor contributing to species delimitation. A recent study indicated similar effects of the major geological events in the past million years on local birds phylogeography in Australia (Dolman and Joseph 2015). For the African continent, both river barriers and large vegetation shifts promoted bird speciation (Voelker et al. 2010, 2013).
In regard to other fauna and flora taxa inhabiting the Western Palearctic, certain degrees of genetic variation can be observed and very often, individuals can be linked to certain populations or geographic areas. This is illustrated by recent studies on Brown Hares (Lepus europaeus) (Minoudi et al. 2018), Stone Martins (Martes foina) (Tsoupas et al. 2019), plus Balkan Mole (Talpa stankovici) and European Mole (Talpa europaea) (Tryfonopoulos et al. 2010), which revealed that populations from the Balkans have region specific haplotypes. However, the Wild Cat (Felis silvestris) only shows five main geographic groups across the whole of Europe, with some populations hybridizing with domestic cats (Mattucci et al. 2016). Similarly, the European Roe Deers (Capreolus capreolus) and European Wild Boars (Sus scrofa) are characterized by a three clade pattern (Scandura et al. 2008; Sommer et al. 2009). Overall, the carnivores tend to exhibit region specific lineages, as has been shown in the Golden Jackals (Canis aureus) (Rutkowski et al. 2015), the Brown Bears (Ursus arctos) (Swenson et al. 2011) and the Grey Wolves (Canis lupus) (Pilot et al. 2010). Smaller mammals, such as the Field Vole (Microtus agrestis) and the Wood Mouse (Apodemus sylvaticus) equally show geographically specific clades across Europe (Jaarola and Searle 2002; Michaux et al. 2003). Reptiles and amphibians, which show very limited mobility, are defined by even more distinct genetic lineages (Joger et al. 2010), as indicated by the Tree Frogs (Hyla arborea) (Dufresnes et al. 2019), the Blotched Snakes (Elaphe sauromates) (Jablonski et al. 2019), the Grass Snakes (Natrix natrix) (Kindler et al. 2017) and the Ocellated Skinks (Chalcides ocellatus) (Kornilios et al. 2010), to name just a few. Several studies in plants have also revealed comparable genetic differentiation (see ivy Hedera sp. (Grivet and Petit 2002) and numerous tree species (Petit et al. 2005)). One study based on mtDNA indicates that the European Stag Beetle (Lucanus cervus) has two main lineages, one restricted to the Balkan peninsula, while the second one is widely distributed in Europe (Cox et al. 2019). For the European Stone Crayfish (Austropotamobius torrentium), several region specific lineages were also uncovered by sequencing the mtDNA (Pârvulescu et al. 2019).
The above examples offer evidence that a species’ phylogeographic status is shaped by a combination of factors, including its locomotive capacities, fidelity to both breeding and migratory areas, as well as the age of the respective species. Furthermore, the current genetic population structure of a species is just a temporary step in its evolution (Avise 2000). To illustrate, species that are characterized by panmixia in present might be undergoing an active process of speciation and lineage sorting, such as the Great Tit (Parus major), which has been shown to have differentiating genomic elements in peripheral populations (Spurgin et al. 2019).
To conclude, genetic admixture in Western Palearctic birds (except the birds from the Macaronesia islands) represents a result of past climatic events, which occurred during the Pleistocene and Pliocene, as well as the high vagility of birds (unparalleled by other taxa), which helped to achieve such high gene flow.
The choice of molecular markers
To date, the majority (i.e. 132 out of 198) of avian phylogeographic studies have employed nucleotide sequences of mtDNA as a molecular marker. In avian and other taxa population history, mtDNA has been a pivotal method which helped the field of phylogeography flourish (Avise 2004; Beheregaray 2008). It has been primarily used in initial surveys of population demography and biography, due to its low cost and efficiency (Mindell 1997). However, mtDNA does have limitations, as it is non-recombinant and maternally inherited (Krebs et al. 2018). These shortcomings have ignited a number of debates (Hebert et al. 2003; Ballard and Whitlock 2004; Hurst and Jiggins 2005; Edwards and Bensch 2009), but we consider that the advantages easily overcome the drawbacks (Rubinoff and Holland 2005; Sequeira et al. 2008; Zink and Barrowclough 2008). To bring further support for mtDNA, we argue that in our dataset, two different studies on the Saker Falcon (Falco cherrug), one employing mtDNA (Nittinger et al. 2007) and the more recent one, using SNPs (Zhan et al. 2015), produced the same phylogeography for the species. Similarly, two studies on the Eurasian Curlew (Numenius arquata), one using nuclear and mitochondrial DNA (Rodrigues et al. 2019), and one with NGS (Tan et al. 2019), yielded comparable results.
After mtDNA, microsatellites and nuDNA are the most commonly used markers. With the accompanying benefits of these two last markers (Avise 2004), the main restrain is that the focus is on a small strain of DNA.
Concerning the low scale usage of NGS (seven out of 198) across the studies included in our review, this is a relatively novel technology (see Fig. 1) and we argue that the costs still represent an impediment for many research groups. Although the sequencing costs are dropping fast, harnessing and affording high-quality computational analyses represents the major drawback. Unfortunately, bioinformatic expert support is still a luxury for many bird research labs.
As a final regard, we envision mtDNA will continue to provide robust first phylogeographic assessments for many years to come and we expect an increase in studies based on NGS data.
What the future holds?
Until present, the bulk of animal phylogeography research has been dominated by mtDNA (Emerson and Hewitt 2005; Avise et al. 2016). With the advent of sequencing techniques, coupled with a decrease in running costs, we expect that the era of big “omics” data will revolutionize the phylogeographic research. However, regarding the later costs, the expenses for computational analyses, computer clusters and human bioinformatic resources are increasing (Muir et al. 2016). Whole-genome sequencing will challenge the present image offered by mtDNA (Kraus and Wink 2015; Ottenburghs et al. 2019) and we anticipate that a number of species currently characterized by panmixia will reveal a certain degree of differentiation. Furthermore, the maturation of phylogeography will benefit not only from developments in DNA sequencing techniques, but also from the advancement of theory and statistical analyses in this field (Stiller and Zhang 2019).
Finally, bird populations are very dynamic and their distribution in space and time are affected by multiple factors such as climate, availability of habitat and food but also, in the past hundred years, anthropogenic threats. This complex network of factors has a fundamental influence on shaping their future phylogeography (Kumar and Kumar 2018). With climate change, unprecedented human-driven alteration of the environment and overall decrease of food stocks (e.g. insects), genetic consequences on bird populations might not be that far away.
Conclusions
After 30 years of studies in phylogeography, we have a good understanding of avian population history in the Western Palearctic, based on the 145 species whose genetic background has been elucidated. The majority of them show high levels of genetic admixture, whereas the species inhabiting the oceanic islands (i.e. Macaronesia) are resident and developed distinct genetic lineages. The panmixia is the legacy of the Pleistocene and Pliocene climatic fluctuations, which forced the birds to cyclically retreat in refugia only to subsequently expand and recolonize higher latitudes, after the ice sheet retreat. These events caused population admixture, both in the refugia and at the contact zones, during population expansion. The bulk of the avian phylogeographic information comes from nucleotide sequences of mtDNA, which, with few limitations, has proven to be a robust and trustworthy molecular marker. With the current dawn of big genomic data in bird research, which offers a much higher resolution than previous studies, we envision a steep increase of NGS-driven phylogeography studies. These new studies have both the power to offer initial population structure surveys and, most important, to challenge previous views based on other markers. Nonetheless, the utility of mtDNA when used with due understanding and in conjunction with NGS is clearly very high. We, therefore, urge our peers not to forget mtDNA completely, which has been a great companion in the past 30 years.
References
Abdilzadeh R, Aliabadian M, Olsson U (2020) Molecular assessment of the distribution and taxonomy of the Lesser Whitethroat Sylvia curruca complex in Iran, with particular emphasis on the identity of the contentious taxon, zagrossiensis Sarudny, 1911. J Ornithol 161:665–676. https://doi.org/10.1007/s10336-020-01772-x
Alaei Kakhki N, Aliabadian M, Förschler MI et al (2018) Phylogeography of the Oenanthe hispanica-pleschanka-cypriaca complex (Aves, Muscicapidae: Saxicolinae): diversification history of open-habitat specialists based on climate niche models, genetic data, and morphometric data. J Zool Syst Evol Res 56:408–427. https://doi.org/10.1111/jzs.12206
Albayrak T, Gonzalez J, Drovetski SV, Wink M (2012) Phylogeography and population structure of Krüper’s Nuthatch Sitta krueperi from Turkey based on microsatellites and mitochondrial DNA. J Ornithol 153:405–411. https://doi.org/10.1007/s10336-011-0756-4
Albrecht F, Hering J, Fuchs E, et al (2018) Phylogeography of the Eurasian Wren Troglodytes troglodytes—complex diversification patterns of Ibero-Maghrebian populations. In: Proceedings of the 151th meeting of the German Ornithologists‘ Society. In Vogelwarte, 56. p 338
Albrecht F, Hering J, Fuchs E et al (2020) Phylogeny of the Eurasian Wren Nannus troglodytes (Aves: Passeriformes: Troglodytidae) reveals deep and complex diversification patterns of Ibero-Maghrebian and Cyrenaican populations. PLoS ONE 15:e0230151
Ammon U (2001) The dominance of english as a language of science. De Gruyer Mouton, Berlin
Ammon U, McConnell G (2002) English as an academic language in Europe. Peter Lang, Bern
Araújo-Silva LE, Miranda LS, Carneiro L, Aleixo A (2017) Phylogeography and diversification of an Amazonian understory hummingbird: paraphyly and evidence for widespread cryptic speciation in the Plio-Pleistocene. Ibis 38:42–49. https://doi.org/10.1111/ibi.12500
Arbabi T, Gonzalez J, Witt HH et al (2014b) Mitochondrial phylogeography of the Eurasian Reed Warbler Acrocephalus scirpaceus and the first genetic record of A. s. fuscus in Central Europe. Ibis 156:799–811. https://doi.org/10.1111/ibi.12174
Arbabi T, Gonzalez J, Wink M (2014a) Mitochondrial evidence for genetic diversity and low phylogeographic differentiation in the Marsh Warbler Acrocephalus palustris (Aves: Acrocephalidae). Org Divers Evol 14:409–417. https://doi.org/10.1007/s13127-014-0177-3
Arshad M, Pedall I, Gonzalez J et al (2009) Genetic variation of four Gyps species (Gyps bengalensis, G. africanus, G. indicus and G. fulvus) based on microsatellite analysis. J Raptor Res 43:227–236
Avise JC (2000) Phylogeography: the history and formation of species. Harvard University Press, Cambridge
Avise JC (2004) Molecular markers, natural history, and evolution, 2nd edn. Sinauer Associates Inc, Sunderland
Avise JC, Walker D (1998) Pleistocene phylogeographic effects on avian populations and the speciation process. Proc R Soc B Biol Sci 265:457–463. https://doi.org/10.1098/rspb.1998.0317
Avise JC, Arnold J, Ball RM et al (1987) Intraspecific Phylogeography: the mitochondrial DNA bridge between population genetics and systematics. Annu Rev Ecol Syst 18:489–522. https://doi.org/10.1146/annurev.es.18.110187.002421
Avise JC, Bowen BW, Ayala FJ (2016) In the light of evolution X: Comparative phylogeography. Proc Natl Acad Sci 113:7957–7961. https://doi.org/10.1073/pnas.1604338113
Backström N, Sætre G-P, Ellegren H (2013) Inferring the demographic history of European Ficedula flycatcher populations. BMC Evol Biol 13:2. https://doi.org/10.1186/1471-2148-13-2
Bagi Z, Dimopoulos EA, Loukovitis D et al (2018) MtDNA genetic diversity and structure of Eurasian Collared Dove (Streptopelia decaocto). PLoS ONE. https://doi.org/10.1371/journal.pone.0193935
Bajc M, Čas M, Ballian D et al (2011) Genetic differentiation of the Western Capercaillie highlights the importance of South-eastern Europe for understanding the species phylogeography. PLoS ONE 6:e23602. https://doi.org/10.1371/journal.pone.0023602
Baker AJ, Piersma T, Rosenmeier L (1994) Unraveling the intraspecific phylogeography of Knots Calidris canutus: a progress report on the search for genetic markers. J Für Ornithol 135:599–608. https://doi.org/10.1007/BF01649854
Ballard JWO, Whitlock MC (2004) The incomplete natural history of mitochondria. Mol Ecol 13:729–744. https://doi.org/10.1046/j.1365-294X.2003.02063.x
Barbanera F, Forcina G, Guerrini M, Dini F (2011) Molecular phylogeny and diversity of the Corsican red-legged partridge: hybridization and management issues. J Zool 285:56–65. https://doi.org/10.1111/j.1469-7998.2011.00813.x
Barrowclough GF, Cracraft J, Klicka J, Zink RM (2016) How many kinds of birds are there and why does it matter? PLoS ONE 11:1–15. https://doi.org/10.1371/journal.pone.0166307
Beheregaray LB (2008) Twenty years of phylogeography: The state of the field and the challenges for the Southern Hemisphere. Mol Ecol 17:3754–3774. https://doi.org/10.1111/j.1365-294X.2008.03857.x
Bell CP (2000) Process in the evolution of bird migration and pattern in avian ecogeography. J Avian Biol 31:258–265. https://doi.org/10.1034/j.1600-048X.2000.310218.x
Bensch S, Hasselquist D (1999) Phylogeographic population structure of Great reed warblers: an analysis of mtDNA control region sequences. Biol J Linn Soc 66:171–185. https://doi.org/10.1111/j.1095-8312.1999.tb01882.x
Bensch S, Andersson T, Åkesson S (1999) Morphological and molecular variation across a migratory divide in willow warblers, Phylloscopus trochilus. Evolution (n Y) 53:1925–1935. https://doi.org/10.1111/j.1558-5646.1999.tb04573.x
Bowen BW, Gaither MR, DiBattista JD et al (2016) Comparative phylogeography of the ocean planet. Proc Natl Acad Sci 113:7962–7969. https://doi.org/10.1073/pnas.1602404113
Brambilla M, Vitulano S, Spina F et al (2008) A molecular phylogeny of the Sylvia cantillans complex: cryptic species within the Mediterranean basin. Mol Phylogenet Evol 48:461–472. https://doi.org/10.1016/j.ympev.2008.05.013
Brito PH (2005) The influence of Pleistocene glacial refugia on Tawny owl genetic diversity and phylogeography in western Europe. Mol Ecol 14:3077–3094. https://doi.org/10.1111/j.1365-294X.2005.02663.x
Cadahía L, Negro JJ, Urios V (2007) Low mitochondrial DNA diversity in the endangered Bonelli’s Eagle (Hieraaetus fasciatus) from SW Europe (Iberia) and NW Africa. J Ornithol 148:99–104. https://doi.org/10.1007/s10336-006-0107-z
Cagnon C, Lauga B, Hemery G, Mouches C (2004) Phylogeographic differentiation of storm petrels (Hydrobates pelagicus) based on cytochrome b mitochondrial DNA variation. Mar Biol 145:1257–1264. https://doi.org/10.1007/s00227-004-1407-6
Çakmak E, Pekçen ÇA, Kirazlı C et al (2019) Extremely Low Mitochondrial DNA Diversity in a near Threatened Species, the Cinereous Vulture Aegypius monachus in Turkey. Ardea 107:85–92
Calderón L, Campagna L, Wilke T et al (2016) Genomic evidence of demographic fluctuations and lack of genetic structure across flyways in a long distance migrant, the European turtle dove. BMC Evol Biol 16:237. https://doi.org/10.1186/s12862-016-0817-7
Capurucho JMG, Ashley MV, Ribas CC, Bates JM (2018) Connecting Amazonian, Cerrado, and Atlantic forest histories: Paraphyly, old divergences, and modern population dynamics in tyrant-manakins (Neopelma/Tyranneutes, Aves: Pipridae). Mol Phylogenet Evol 127:696–705. https://doi.org/10.1016/j.ympev.2018.06.015
Cardia P, Fráguas B, Pais M et al (2000) Preliminary genetic analysis of some western Palaeartic populations of Bonelli’s Eagle, Hieraetus fasciatus. In: Chancellor RD, Meyburg BU (eds) Raptors at risk. Hancock, WWGBP/Hancock House, pp 845–851
Carneiro L, Bravo GA, Aristizábal N et al (2018) Molecular systematics and biogeography of lowland antpittas (Aves, Grallariidae): The role of vicariance and dispersal in the diversification of a widespread Neotropical lineage. Mol Phylogenet Evol 120:375–389. https://doi.org/10.1016/j.ympev.2017.11.019
Corrales C, Pavlovska M, Höglund J (2014) Phylogeography and subspecies status of Black Grouse. J Ornithol 155:13–25. https://doi.org/10.1007/s10336-013-0982-z
Cox K, McKeown N, Antonini G et al (2019) Phylogeographic structure and ecological niche modelling reveal signals of isolation and postglacial colonisation in the European stag beetle. PLoS ONE 14:e0215860. https://doi.org/10.1371/journal.pone.0215860
Cwynar LC, MacDonald GM (1987) Geographical variation of lodgepole pine in relation to population history. Am Nat 129:463–469. https://doi.org/10.1086/284651
de Melo C, Moura C, de Araujo HF, Aleixo A et al (2018) The role of landscape change and paleoclimatic events in shaping the evolutionary history of the Polioptila gnatcatchers (Passeriformes, Polioptilidae) with emphasis on species associated with open habitats. J Avian Biol 49:1–12. https://doi.org/10.1111/jav.01692
de Melo C, Moura C, Bastian H-V, Bastian A et al (2019) Pliocene origin, ice ages and postglacial population expansion have influenced a panmictic phylogeography of the European Bee-Eater Merops apiaster. Diversity 11:12. https://doi.org/10.3390/d11010012
De Melo C, Moura C, Fernandes AM, Aleixo A et al (2020) Evolutionary history of the Pectoral Sparrow Arremon taciturnus : Evidence for diversification during the Late Pleistocene. Ibis. https://doi.org/10.1111/ibi.12813
Dietzen C, Witt H-H, Wink M (2003) The phylogeographic differentiation of the European robin Erithacus rubecula on the Canary Islands revealed by mitochondrial DNA sequence data and morphometrics: evidence for a new robin taxon on Gran Canaria? Avian Sci 3:115–131
Dietzen C, Voigt C, Wink M et al (2006) Phylogeography of island canary (Serinus canaria) populations. J Ornithol 147:485–494. https://doi.org/10.1007/s10336-005-0044-2
Dietzen C, Garcia-del-Rey E, Castro GD, Wink M (2008) Phylogeography of the blue tit (Parus teneriffae-group) on the Canary Islands based on mitochondrial DNA sequence data and morphometrics. J Ornithol 149:1–12. https://doi.org/10.1007/s10336-007-0192-7
Dietzen C, Michels JP, Wink M (2015) Formal description of a new subspecies of the European Robin from Gran Canaria Island, Spain ( Aves : Muscicapidae : Erithacus rubecula marionae subsp. nov.). J Open Ornithol 8:39–42
Dohms KM (2016) Phylogeography of three high latitude resident corvids : Clark’s nutcracker (Nucifraga columbiana), Eurasian nutcracker (Nucifraga caryocatactes), and Gray jay (Perisoreus canadensis). University of Lethbridge, Lethbridge
Dohms KM, Burg TM (2014) Limited geographic genetic structure detected in a widespread Palearctic corvid Nucifraga Caryocatactes. PeerJ 2:e371. https://doi.org/10.7717/peerj.371
Dolman G, Joseph L (2015) Evolutionary history of birds across southern Australia: structure, history and taxonomic implications of mitochondrial DNA diversity in an ecologically diverse suite of species. Emu - Austral Ornithol 115:35–48. https://doi.org/10.1071/MU14047
Drovetski SV, Zink MR, Sievert R et al (2004) Complex biogeographic history of a Holarctic passerine. Proc R Soc London Ser B Biol Sci 271:545–551. https://doi.org/10.1098/rspb.2003.2638
Drovetski SV, Raković M, Semenov G et al (2014) Limited phylogeographic signal in sex-linked and autosomal loci despite geographically, ecologically, and phenotypically concordant structure of mtDNA variation in the Holarctic Avian genus Eremophila. PLoS ONE 9:1–10. https://doi.org/10.1371/journal.pone.0087570
Drovetski SV, Fadeev IV, Raković M et al (2018) A test of the European Pleistocene refugial paradigm, using a Western Palaearctic endemic bird species. Proc R Soc B Biol Sci. https://doi.org/10.1098/rspb.2018.1606
Dufresnes C, Beddek M, Skorinov V, et al (2019) Diversification and speciation in tree frogs from the Maghreb (Hyla meridionalis sensu lato), with description of a new African endemic. Mol Phylogenet Evol 134:291–299. https://doi.org/10.1016/j.ympev.2019.02.009
Duriez O, Sachet J-M, Ménoni E et al (2007) Phylogeography of the Capercaillie in Eurasia: what is the conservation status in the Pyrenees and Cantabrian Mounts? Conserv Genet 8:513–526. https://doi.org/10.1007/s10592-006-9165-2
Edwards S, Bensch S (2009) Looking forwards or looking backwards in avian phylogeography? A comment on Zink and Barrowclough 2008. Mol Ecol 18:2930–2933
Elverici C (2018) Phylogeography of Rock Nuthatches: an integrated approach. Hacettepe University, Hacettepe
Emerson BC, Hewitt GM (2005) Phylogeography. Curr Biol 15:R367–R371
Eyres A, Eronen JT, Hagen O et al (2021) Climatic effects on niche evolution in a passerine bird clade depend on paleo-climate reconstruction method. Evolution (n Y). https://doi.org/10.1111/evo.14209
Fernandes AM, Wink M, Aleixo A (2012) Phylogeography of the Chestnut-tailed Antbird (Myrmeciza hemimelaena) clarifies the role of rivers in Amazonian biogeography. J Biogeogr 39:1524–1535. https://doi.org/10.1111/j.1365-2699.2012.02712.x
Fernandes AM, Gonzalez J, Wink M, Aleixo A (2013) Multilocus phylogeography of the Wedge-billed Woodcreeper Glyphorynchus spirurus (Aves, Furnariidae) in lowland Amazonia: widespread cryptic diversity and paraphyly reveal a complex diversification pattern. Mol Phylogenet Evol 66:270–282. https://doi.org/10.1016/j.ympev.2012.09.033
Fernandes AM, Wink M, Sardelli CH, Aleixo A (2014) Multiple speciation across the Andes and throughout Amazonia: the case of the Spot-backed Antbird species complex (Hylophylax naevius/Hylophylax naevioides). J Biogeogr 41:1094–1104. https://doi.org/10.1111/jbi.12277
Frenzel B, Pécsi M, Velichko AA (1992) Atlas of paleoclimates and paleoenvironments of the Northern hemisphere. Late Pleistocene– Holocene, Geographic. Geographical Research Institute; Hungarian Academy of Sciences. Gustav Fischer Verlag, Budapest, Stuttgart
Garcia JT, Mañosa S, Morales MB et al (2011) Genetic consequences of interglacial isolation in a steppe bird. Mol Phylogenet Evol 61:671–676. https://doi.org/10.1016/j.ympev.2011.07.017
Garcia-del-Rey E, Delgado G, Gonzalez J, Wink M (2007) Canary Island Freat spotted woodpecker (Dendrocopos major) has distinct mtDNA. J Ornithol 148:531–536. https://doi.org/10.1007/s10336-007-0142-4
Garcia-del-Rey E, Marthinsen G, Calabuig P et al (2013) Reduced genetic diversity and sperm motility in the endangered Gran Canaria Blue Chaffinch Fringilla teydea polatzeki. J Ornithol 154:761–768. https://doi.org/10.1007/s10336-013-0940-9
Gay L, Defos Du Rau P, Mondain-Monval J-Y, Crochet P-A (2004) Phylogeography of a game species: the Red-crested pochard (Netta rufina) and consequences for its management. Mol Ecol 13:1035–1045. https://doi.org/10.1111/j.1365-294X.2004.02117.x
Ghorbani F, Aliabadian M, Olsson U et al (2020a) Mitochondrial phylogeography of the genus Eremophila confirms underestimated species diversity in the Palearctic. J Ornithol 161:297–312. https://doi.org/10.1007/s10336-019-01714-2
Ghorbani F, Aliabadian M, Zhang R et al (2020b) Densely sampled phylogenetic analyses of the Lesser Short-toed Lark (Alaudala rufescens)—Sand Lark (A. raytal) species complex (Aves, Passeriformes) reveal cryptic diversity. Zool Scr 49:427–439. https://doi.org/10.1111/zsc.12422
Gill F, Donsker D, Rasmussen P (2020) IOC World Bird List v10.1. https://www.worldbirdnames.org/. Accessed 31 Jan 2020
Godoy JA, Negro JJ, Hiraldo F, Donázar JA (2004) Phylogeography, genetic structure and diversity in the endangered bearded vulture (Gypaetus barbatus, L.) as revealed by mitochondrial DNA. Mol Ecol 13:371–390. https://doi.org/10.1046/j.1365-294X.2003.02075.x
Gómez-Díaz E, González-Solís J, Peinado MA, Page RDM (2006) Phylogeography of the Calonectris shearwaters using molecular and morphometric data. Mol Phylogenet Evol 41:322–332. https://doi.org/10.1016/j.ympev.2006.05.006
Gonzalez J, Wink M, Garcia-del-Rey E, Delgado Castro G (2008) Evidence from DNA nucleotide sequences and ISSR profiles indicates paraphyly in subspecies of the Southern Grey Shrike (Lanius meridionalis). J Ornithol 149:495–506. https://doi.org/10.1007/s10336-008-0293-y
Grant PR, Grant BR (1992) Hybridization of bird species. Science. https://doi.org/10.1126/science.256.5054.193
Grivet D, Petit RJ (2002) Phylogeography of the common ivy (Hedera sp.) in Europe: genetic differentiation through space and time. Mol Ecol 11:1351–1362. https://doi.org/10.1046/j.1365-294X.2002.01522.x
Grosso AR, Bastos-Silveira C, Coelho MM, Dias D (2006) Columba palumbus Cyt b-like Numt sequence: comparison with functional homologue and the use of universal primers. Folia Zool 55:131–144
Guillaumet A, Pons J-M, Godelle B, Crochet P-A (2006) History of the Crested Lark in the Mediterranean region as revealed by mtDNA sequences and morphology. Mol Phylogenet Evol 39:645–656. https://doi.org/10.1016/j.ympev.2006.01.002
Haase M, Höltje H, Blahy B et al (2019) Shallow genetic population structure in an expanding migratory bird with high breeding site fidelity, the Western Eurasian Crane Grus grus grus. J Ornithol 160:965–972. https://doi.org/10.1007/s10336-019-01688-1
Haffer J (1969) Speciation in amazonian forest birds. Science 165:131–137. https://doi.org/10.1126/science.165.3889.131
Hansson B, Hasselquist D, Tarka M et al (2008) Postglacial colonisation patterns and the role of isolation and expansion in driving diversification in a passerine bird. PLoS ONE 3:1–8. https://doi.org/10.1371/journal.pone.0002794
Haring E, Gamauf A, Kryukov A (2007) Phylogeographic patterns in widespread corvid birds. Mol Phylogenet Evol 45:840–862. https://doi.org/10.1016/j.ympev.2007.06.016
Harris RB, Alström P, Ödeen A, Leaché AD (2018) Discordance between genomic divergence and phenotypic variation in a rapidly evolving avian genus (Motacilla). Mol Phylogenet Evol 120:183–195. https://doi.org/10.1016/j.ympev.2017.11.020
Hausknecht R, Jacobs S, Müller J et al (2014) Phylogeographic analysis and genetic cluster recognition for the conservation of Ural Owls (Strix uralensis) in Europe. J Ornithol 155:121–134. https://doi.org/10.1007/s10336-013-0994-8
Hebert PDN, Cywinska A, Ball SL, DeWaard JR (2003) Biological identifications through DNA barcodes. Proc R Soc London Ser B Biol Sci 270:313–321. https://doi.org/10.1098/rspb.2002.2218
Hermansen JOS, Sæther SA, Elgvin TO et al (2011) Hybrid speciation in sparrows I: phenotypic intermediacy, genetic admixture and barriers to gene flow. Mol Ecol 20:3812–3822. https://doi.org/10.1111/j.1365-294X.2011.05183.x
Hernández MÁ, Campos F, Santamaría T et al (2012) Genetic differences among Iberian White-Throated Dipper Cinclus cinclus populations based on the cytochrome b sequence. Ardeola 59:111–122
Hernández MA, Campos F, Santamaría T et al (2016) Is isolation by distance the cause of the genetic structure of the Iberian white-throated dipper populations? J Zool 299:27–36. https://doi.org/10.1111/jzo.12315
Hewitt GM (1999) Post-glacial re-colonization of European biota. Biol J Linn Soc 68:87–112. https://doi.org/10.1111/j.1095-8312.1999.tb01160.x
Hewitt G (2000) The genetic legacy of the Quaternary ice ages. Nature 405:907–913. https://doi.org/10.1038/35016000
Hickerson MJ, Carstens BC, Cavender-Bares J et al (2010) Phylogeography’s past, present, and future: 10 years after Avise, 2000. Mol Phylogenet Evol 54:291–301. https://doi.org/10.1016/j.ympev.2009.09.016
Höglund J, Larsson JK, Jansman HAH, Segelbacher G (2007) Genetic variability in European black grouse (Tetrao tetrix). Conserv Genet 8:239–243. https://doi.org/10.1007/s10592-006-9158-1
Höglund J, Johansson T, Beintema A, Schekkerman H (2009) Phylogeography of the Black-tailed Godwit Limosa limosa: substructuring revealed by mtDNA control region sequences. J Ornithol 150:45–53. https://doi.org/10.1007/s10336-008-0316-8
Hogner S, Laskemoen T, Lifjeld JT et al (2012) Deep sympatric mitochondrial divergence without reproductive isolation in the Common redstart Phoenicurus phoenicurus. Ecol Evol 2:2974–2988. https://doi.org/10.1002/ece3.398
Honnen A-C, Hailer F, Kenntner N et al (2010) Mitochondrial DNA and nuclear microsatellites reveal high diversity and genetic structure in an avian top predator, the White-tailed sea eagle, in central Europe. Biol J Linn Soc 99:727–737. https://doi.org/10.1111/j.1095-8312.2010.01373.x
Hou Z-C, Yang F-X, Qu L-J et al (2011) Genetic structure of Eurasian and North American mallard ducks based on mtDNA data. Anim Genet 43:352–355. https://doi.org/10.1111/j.1365-2052.2011.02248.x
Hourlay F, Libois R, D’Amico F et al (2008) Evidence of a highly complex phylogeographic structure on a specialist river bird species, the Dipper (Cinclus cinclus). Mol Phylogenet Evol 49:435–444. https://doi.org/10.1016/j.ympev.2008.07.025
Hung CM, Drovetski SV, Zink RM (2012b) Recent allopatric divergence and niche evolution in a widespread Palearctic bird, the Common rosefinch (Carpodacus erythrinus). Mol Phylogenet Evol 66:103–111. https://doi.org/10.1016/j.ympev.2012.09.012
Hung C-M, Drovetski SV, Zink RM (2012a) Multilocus coalescence analyses support a mtDNA-based phylogeographic history for a widespread Palearctic passerine bird, Sitta europaea. Evolution (n Y) 66:2850–2864. https://doi.org/10.1111/j.1558-5646.2012.01657.x
Hurst GDD, Jiggins FM (2005) Problems with mitochondrial DNA as a marker in population, phylogeographic and phylogenetic studies: The effects of inherited symbionts. Proc R Soc B Biol Sci 272:1525–1534. https://doi.org/10.1098/rspb.2005.3056
Illera JC, Koivula K, Broggi J et al (2011) A multi-gene approach reveals a complex evolutionary history in the Cyanistes species group. Mol Ecol 20:4123–4139. https://doi.org/10.1111/j.1365-294X.2011.05259.x
Illera JC, Palmero AM, Laiolo P et al (2014) Genetic, morphological, and acoustic evidence reveals lack of diversification in the colonization process in an island bird. Evolution (n Y). https://doi.org/10.1111/evo.12429
Illera JC, Rando JC, Rodriguez-Exposito E et al (2018) Acoustic, genetic, and morphological analyses of the Canarian common chaffinch complex Fringilla coelebs ssp. reveals cryptic diversification. J Avian Biol 49:1–12. https://doi.org/10.1111/jav.01885
Illera JC, Ramírez Á, Rodríguez L et al (2020) Maternal genetic structure reveals an incipient differentiation in the Canary Islands Chiffchaff Phylloscopus canariensis. Ardeola 67:401–414. https://doi.org/10.13157/arla.67.2.2020.ra11
IUCN (2019) The IUCN Red List of Threatened Species. Version 2019–2. In: IUCN Red List Threat. Species. Version 2019–2. http://www.iucnredlist.org. Accessed 10 Jul 2019
Jaarola M, Searle JB (2002) Phylogeography of Field voles (Microtus agrestis) in Eurasia inferred from mitochondrial DNA sequences. Mol Ecol 11:2613–2621. https://doi.org/10.1046/j.1365-294X.2002.01639.x
Jablonski D, Kukushkin OV, Avcı A et al (2019) The biogeography of Elaphe sauromates (Pallas, 1814), with a description of a new Rat snake species. PeerJ 7:e6944. https://doi.org/10.7717/peerj.6944
Joger U, Fritz U, Guicking D et al (2010) Relict populations and endemic clades in Palaearctic reptiles: evolutionary history and implications for conservation. In: Habel JC, Assman T (eds) Relict species-phylogeography and conservation. Springer, Berlin , pp 119–144
Joseph L (2018) Phylogeography and the role of hybridization in speciation. In: Tietze DT (ed) Bird species: how they arise, modify and vanish. Springer International Publishing, Cham, pp 165–194
Joseph L, Omland KE (2009) Phylogeography: its development and impact in Australo-Papuan ornithology with special reference to paraphyly in Australian birds. Emu 109:1–23
Jowers MJ, Sánchez-Ramírez S, Lopes S et al (2019) Unravelling population processes over the Late Pleistocene driving contemporary genetic divergence in Palearctic Buzzards. Mol Phylogenet Evol 134:269–281. https://doi.org/10.1016/j.ympev.2019.02.004
Kamp L, Pasinelli G, Milanesi P et al (2018) Significant Asia-Europe divergence in the Middle spotted woodpecker (Aves, Picidae). Zool Scr. https://doi.org/10.1111/zsc.12320
Kindler C, Chèvre M, Ursenbacher S et al (2017) Hybridization patterns in two contact zones of grass snakes reveal a new Central European snake species. Sci Rep 7:7378. https://doi.org/10.1038/s41598-017-07847-9
Klinga P, Mikoláš M, Delegan IV et al (2020) Temporal landscape genetic data indicate an ongoing disruption of gene flow in a relict bird species. Conserv Genet 21:329–340. https://doi.org/10.1007/s10592-020-01253-x
Koopman EM, McDonald BD, Hayward DG et al (2005) Genetic similarity among Eurasian subspecies of Boreal owls Aegolius funereus. J Avian Biol 36:179–183. https://doi.org/10.1111/j.0908-8857.2005.03509.x
Korepov MV, Stryukov SA, Korepova DA et al (2017) Genetic differentiation and polymorphism of the Volga population of the Imperial Eagle (Aquila heliaca, Falconiformes, Accipitridae) according to a mitochondrial DNA analysis. Univ Proceedings Volga Reg Nat Sci. https://doi.org/10.21685/2307-9150-2017-3-1
Kornilios P, Kyriazi P, Poulakakis N et al (2010) Phylogeography of the ocellated skink Chalcides ocellatus (Squamata, Scincidae), with the use of mtDNA sequences: a hitch-hiker’s guide to the Mediterranean. Mol Phylogenet Evol 54:445–456. https://doi.org/10.1016/j.ympev.2009.09.015
Kraus RHS, Wink M (2015) Avian genomics: fledging into the wild! J Ornithol 156:851–865. https://doi.org/10.1007/s10336-015-1253-y
Krebs JE, Goldstein ES, Kilpatrick ST (2018) Lewin’s genes XII. Jones & Bartlett Learning, Burlington
Kryukov AP (2019) Phylogeography and hybridization of corvid birds in the Palearctic Region. Vavilov J Genet Breed 23:232–238
Kryukov AP, Spiridonova LN, Mori S et al (2017) Deep phylogeographic breaks in Magpie Pica pica across the Holarctic: concordance with bioacoustics and phenotypes. Zoolog Sci 34:185–200
Kulikova IV, Poysa H, Zhuravlev YN (2012) Phylogeography of the mallard Anas platyrhynchos from Eurasia inferred from sequencing of the mtDNA control region. Russ J Genet 48:705–712. https://doi.org/10.1134/S1022795412070095
Kulikova IV, Zhuravlev YN, Korobitsyn IG et al (2019) Holarctic phylogeographic structure of Eurasian wigeon (Mareca penelope). Vavilov J Genet Breed 23:362–369
Kumar R, Kumar V (2018) A review of phylogeography: biotic and abiotic factors. Geol Ecol Landscapes 2:268–274. https://doi.org/10.1080/24749508.2018.1452486
Kumar S, Stecher G, Li M et al (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Kunz F, Gamauf A, Zachos FE, Haring E (2019) Mitochondrial phylogenetics of the goshawk Accipiter [gentilis] superspecies. J Zool Syst Evol Res 57:942–958. https://doi.org/10.1111/jzs.12285
Kvist L (2000) Phylogeny and phylogeography of European Parids. University of Oulu, Finland
Kvist L, Ruokonen M, Lumme J, Orell M (1999) The colonization history and present-day population structure of the European great tit (Parus major major). Heredity (edinb) 82:495–502. https://doi.org/10.1038/sj.hdy.6885130
Kvist L, Martens J, Ahola A, Orell M (2001) Phylogeography of a Palaearctic sedentary passerine, the willow tit (Parus montanus). J Evol Biol 14:930–941. https://doi.org/10.1046/j.1420-9101.2001.00354.x
Kvist L, Viiri K, Dias PC et al (2004) Glacial history and colonization of Europe by the Blue tit Parus caeruleus. J Avian Biol 35:352–359. https://doi.org/10.1111/j.0908-8857.2004.03297.x
Kvist L, Broggi J, Illera JC, Koivula K (2005) Colonisation and diversification of the blue tits (Parus caeruleus teneriffae-group) in the Canary Islands. Mol Phylogenet Evol 34:501–511. https://doi.org/10.1016/j.ympev.2004.11.017
Kvist L, Giralt D, Valera F et al (2011) Population decline is accompanied by loss of genetic diversity in the Lesser Grey Shrike Lanius minor. Ibis 153:98–109. https://doi.org/10.1111/j.1474-919X.2010.01091.x
Lagerholm VK, Sandoval-Castellanos E, Vaniscotte A et al (2017) Range shifts or extinction? Ancient DNA and distribution modelling reveal past and future responses to climate warming in cold-adapted birds. Glob Chang Biol 23:1425–1435. https://doi.org/10.1111/gcb.13522
Langguth T, Honnen A-C, Hailer F et al (2013) Genetic structure and phylogeography of a European flagship species, the White-tailed sea eagle Haliaeetus albicilla. J Avian Biol 44:263–271. https://doi.org/10.1111/j.1600-048X.2012.00075.x
Lauga B, Cagnon C, D’Amico F et al (2005) Phylogeography of the White-throated dipper Cinclus cinclus in Europe. J Ornithol 146:257–262. https://doi.org/10.1007/s10336-005-0087-4
LeBlanc NM, Stewart DT, Pálsson S et al (2017) Population structure of Purple Sandpipers (Calidris maritima) as revealed by mitochondrial DNA and microsatellites. Ecol Evol 7:3225–3242. https://doi.org/10.1002/ece3.2927
Lehtonen PK, Laaksonen T, Artemyev AV et al (2009) Geographic patterns of genetic differentiation and plumage colour variation are different in the Pied flycatcher (Ficedula hypoleuca). Mol Ecol 18:4463–4476. https://doi.org/10.1111/j.1365-294X.2009.04364.x
Leigh JW, Bryant D (2015) POPART: Full-feature software for haplotype network construction. Methods Ecol Evol 6:1110–1116. https://doi.org/10.1111/2041-210X.12410
Li X, Dong F, Lei F et al (2016) Shaped by uneven Pleistocene climate: mitochondrial phylogeographic pattern and population history of White wagtail Motacilla alba (Aves: Passeriformes). J Avian Biol 47:263–274. https://doi.org/10.1111/jav.00826
Liebers D, Helbig AJ (2002) Phylogeography and colonization history of Lesser Black-backed Gulls (Larus fuscus) as revealed by mtDNA sequences. J Evol Biol 15:1021–1033. https://doi.org/10.1046/j.1420-9101.2002.00454.x
Liebers D, Helbig AJ, De Knijff P (2001) Genetic differentiation and phylogeography of gulls in the Larus cachinnans–fuscus group (Aves: Charadriiformes). Mol Ecol 10:2447–2462. https://doi.org/10.1046/j.0962-1083.2001.01370.x
Lifjeld JT, Anmarkrud JA, Calabuig P et al (2016) Species-level divergences in multiple functional traits between the two endemic subspecies of Blue Chaffinches Fringilla teydea in Canary Islands. BMC Zool 1:4. https://doi.org/10.1186/s40850-016-0008-4
Lisiecki LE, Raymo ME (2007) Plio-Pleistocene climate evolution: trends and transitions in glacial cycle dynamics. Quat Sci Rev 26:56–69. https://doi.org/10.1016/j.quascirev.2006.09.005
Liu B, Alström P, Olsson U et al (2017) Explosive radiation and spatial expansion across the cold environments of the Old World in an avian family. Ecol Evol 7:6346–6357. https://doi.org/10.1002/ece3.3136
Liukkonen-Anttila T, Uimaniemi L, Orell M, Lumme J (2002) Mitochondrial DNA variation and the phylogeography of the Grey partridge (Perdix perdix) in Europe: from Pleistocene history to present day populations. J Evol Biol 15:971–982. https://doi.org/10.1046/j.1420-9101.2002.00460.x
Liukkonen-Anttila T, Rätti O, Kvist L et al (2004) Lack of genetic structuring and subspecies differentiation in the Capercaillie (Tetrao urogallus) in Finland. Ann Zool Fennici 41:619–633
Lopes RJ, Hortas F, Wennerberg L (2008) Geographical segregation in Dunlin Calidris alpina populations wintering along the East Atlantic migratory flyway—evidence from mitochondrial DNA analysis. Divers Distrib 14:732–741. https://doi.org/10.1111/j.1472-4642.2008.00480.x
Lucchini V, Randi E (1998) Mitochondrial DNA sequence variation and phylogeographical structure of rock partridge (Alectoris graeca) populations. Heredity (edinb) 81:528–536. https://doi.org/10.1038/sj.hdy.6884130
Marion L, Le Gentil J (2006) Ecological segregation and population structuring of the Cormorant Phalacrocorax carbo in Europe, in relation to the recent introgression of continental and marine subspecies. Evol Ecol 20:193–216. https://doi.org/10.1007/s10682-005-5828-6
Marthinsen G, Wennerberg L, Lifjeld JT (2007) Phylogeography and subspecies taxonomy of dunlins (Calidris alpina) in western Palearctic analysed by DNA microsatellites and amplified fragment length polymorphism markers. Biol J Linn Soc 92:713–726. https://doi.org/10.1111/j.1095-8312.2007.00931.x
Marthinsen G, Wennerberg L, Solheim R, Lifjeld JT (2009) No phylogeographic structure in the circumpolar Snowy owl (Bubo scandiacus). Conserv Genet 10:923–933. https://doi.org/10.1007/s10592-008-9581-6
Martin A, Simon C (1990) Differing levels of among-population divergence in the mitochondrial DNA of periodical cicadas related to historical biogeography. Evolution (n Y) 44:1066–1080. https://doi.org/10.2307/2409567
Martinez-Cruz B, Godoy JA, Negro JJ (2004) Population genetics after fragmentation: the case of the endangered Spanish imperial eagle (Aquila adalberti). Mol Ecol 13:2243–2255. https://doi.org/10.1111/j.1365-294X.2004.02220.x
Mattucci F, Oliveira R, Lyons LA et al (2016) European wildcat populations are subdivided into five main biogeographic groups: consequences of Pleistocene climate changes or recent anthropogenic fragmentation? Ecol Evol 6:3–22. https://doi.org/10.1002/ece3.1815
McCarthy EM (2006) Handbook of avian hybrids of the world. Oxford University Press, Oxford
Melnick DJ, Hoelzer GA, Absher R, Ashley MV (1993) mtDNA diversity in rhesus monkeys reveals overestimates of divergence time and paraphyly with neighboring species. Mol Biol Evol 10:282–295. https://doi.org/10.1093/oxfordjournals.molbev.a040004
Michaux JR, Magnanou E, Paradis E et al (2003) Mitochondrial phylogeography of the woodmouse (Apodemus sylvaticus) in the Western Palearctic region. Mol Ecol 12:685–697. https://doi.org/10.1046/j.1365-294X.2003.01752.x
Mindell DP (ed) (1997) Avian molecular evolution and systematics. Academic Press, London
Minoudi S, Papapetridis I, Karaiskou N et al (2018) Genetic analyses of brown hare (Lepus europaeus) support limited migration and translocation of Greek populations. PLoS ONE 13:e0206327. https://doi.org/10.1371/journal.pone.0206327
Monti F, Duriez O, Arnal V et al (2015) Being cosmopolitan: evolutionary history and phylogeography of a specialized raptor, the Osprey Pandion haliaetus. BMC Evol Biol 15:255. https://doi.org/10.1186/s12862-015-0535-6
Mori A, Baldaccini NE, Baratti M et al (2014) A first assessment of genetic variability in the Eurasian Stone-curlew Burhinus oedicnemus. Ibis 156:687–692. https://doi.org/10.1111/ibi.12164
Mori A, Giunchi D, Rodríguez-Godoy F et al (2017) Multilocus approach reveals an incipient differentiation process in the Stone-curlew, Burhinus oedicnemus around the Mediterranean basin. Conserv Genet 18:197–209. https://doi.org/10.1007/s10592-016-0894-6
Moussy C, Arlettaz R, Copete JL et al (2018) The genetic structure of the European breeding populations of a declining farmland bird, the Ortolan bunting (Emberiza hortulana), reveals conservation priorities. Conserv Genet 19:909–922. https://doi.org/10.1007/s10592-018-1064-9
Muir P, Li S, Lou S et al (2016) The real cost of sequencing: scaling computation to keep pace with data generation. Genome Biol 17:53. https://doi.org/10.1186/s13059-016-0917-0
Nebel C, Gamauf A, Haring E et al (2015) Mitochondrial DNA analysis reveals Holarctic homogeneity and a distinct Mediterranean lineage in the Golden eagle (Aquila chrysaetos). Biol J Linn Soc 116:328–340. https://doi.org/10.1111/bij.12583
Nebel C, Gamauf A, Haring E et al (2019) New insights into population structure of the European golden eagle (Aquila chrysaetos) revealed by microsatellite analysis. Biol J Linn Soc. https://doi.org/10.1093/biolinnean/blz130
Nemesházi E, Kövér S, Zachos FE et al (2016) Natural and anthropogenic influences on the population structure of White-tailed eagles in the Carpathian Basin and central Europe. J Avian Biol 47:795–805. https://doi.org/10.1111/jav.00938
Neto JM, Arroyo JL, Bargain B et al (2012) Phylogeography of a habitat specialist with high dispersal capability: the Savi’s Warbler Locustella luscinioides. PLoS ONE 7:e38497. https://doi.org/10.1371/journal.pone.0038497
Nevo E, Beiles A (1992) MtDNA polymorphisms: evolutionary significance in adaptation and speciation of subterranean mole rats. Biol J Linn Soc 47:385–405. https://doi.org/10.1111/j.1095-8312.1992.tb00676.x
Nittinger F, Gamauf A, Pinsker W et al (2007) Phylogeography and population structure of the Saker falcon (Falco cherrug) and the influence of hybridization: mitochondrial and microsatellite data. Mol Ecol 16:1497–1517. https://doi.org/10.1111/j.1365-294X.2007.03245.x
Olsson U, Alström P, Svensson L et al (2010) The Lanius excubitor (Aves, Passeriformes) conundrum-Taxonomic dilemma when molecular and non-molecular data tell different stories. Mol Phylogenet Evol 55:347–357. https://doi.org/10.1016/j.ympev.2009.11.010
Olsson U, Leader PJ, Carey GJ et al (2013) New insights into the intricate taxonomy and phylogeny of the Sylvia curruca complex. Mol Phylogenet Evol 67:72–85. https://doi.org/10.1016/j.ympev.2012.12.023
Olsson U, Rguibi-Idrissi H, Copete JL et al (2016) Mitochondrial phylogeny of the Eurasian/African reed warbler complex (Acrocephalus, Aves). Disagreement between morphological and molecular evidence and cryptic divergence: A case for resurrecting Calamoherpe ambigua Brehm 1857. Mol Phylogenet Evol 102:30–44. https://doi.org/10.1016/j.ympev.2016.05.026
Ottenburghs J, Ydenberg RC, Van Hooft P et al (2015) The Avian Hybrids Project: gathering the scientific literature on avian hybridization. Ibis 157:892–894. https://doi.org/10.1111/ibi.12285
Ottenburghs J, Kraus RHS, van Hooft P et al (2017) Avian introgression in the genomic era. Avian Res 8:30. https://doi.org/10.1186/s40657-017-0088-z
Ottenburghs J, Lavretsky P, Peters JL et al (2019) Population genomics and phylogeography. In: Kraus RHS (ed) Avian genomics in ecology and evolution. Springer International Publishing, Cham, pp 237–265
Ottvall R, Höglund J, Bensch S, Larsson K (2005) Population differentiation in the redshank (Tringa totanus) as revealed by mitochondrial DNA and amplified fragment length polymorphism markers. Conserv Genet 6:321–331. https://doi.org/10.1007/s10592-005-4973-3
Päckert M, Martens J, Sun YH et al (2012) Horizontal and elevational phylogeographic patterns of Himalayan and Southeast Asian forest passerines (Aves: Passeriformes). J Biogeogr 39:556–573. https://doi.org/10.1111/j.1365-2699.2011.02606.x
Päckert M, Bader-Blukott M, Künzelmann B et al (2020) A revised phylogeny of nuthatches (Aves, Passeriformes, Sitta) reveals insight in intra- and interspecific diversification patterns in the Palearctic. Vertebr Zool 70:241–262. https://doi.org/10.26049/VZ70-2-2020-10
Padilla DP, Spurgin LG, Fairfield EA et al (2015) Population history, gene flow, and bottlenecks in island populations of a secondary seed disperser, the Southern grey shrike (Lanius meridionalis koenigi). Ecol Evol 5:36–45. https://doi.org/10.1002/ece3.1334
Pârâu LG, Frias-Soler RC, Wink M (2019) High Genetic Diversity among Breeding Red-Backed Shrikes Lanius collurio in the Western Palearctic. Diversity 11:31. https://doi.org/10.3390/d11030031
Pârvulescu L, Pérez-Moreno JL, Panaiotu C et al (2019) A journey on plate tectonics sheds light on European crayfish phylogeography. Ecol Evol 9:1957–1971. https://doi.org/10.1002/ece3.4888
Pasquet E, Thibault J-C (1997) Genetic differences among mainland and insular forms of the Citril Finch Serinus citrinella. Ibis 139:679–684. https://doi.org/10.1111/j.1474-919X.1997.tb04691.x
Pavlova A (2004) Comparative phylogeography of Eurasian birds. University of Minnesota, Minnesota
Pavlova A, Zink RM, Drovetski SV et al (2003) Phylogeographic patterns in Motacilla flava and Motacilla citreola: species limits and population history. Auk 120:744–758
Pavlova A, Zink RM, Rohwer S (2005a) Evolutionary history, population genetics, and gene flow in the Common rosefinch (Carpodacus erythrinus). Mol Phylogenet Evol 36:669–681. https://doi.org/10.1016/j.ympev.2005.02.010
Pavlova A, Zink RM, Rohwer S et al (2005b) Mitochondrial DNA and plumage evolution in the White wagtail Motacilla alba. J Avian Biol 36:322–336. https://doi.org/10.1111/j.0908-8857.2005.03373.x
Pavlova A, Rohwer S, Drovetski SV, Zink RM (2006) Different post-pleistocene histories of eurasian parids. J Hered 97:389–402. https://doi.org/10.1093/jhered/esl011
Pavlova A, Zink RM, Drovetski SV, Rohwer S (2008) Pleistocene evolution of closely related Sand martins Riparia riparia and R. diluta. Mol Phylogenet Evol 48:61–73. https://doi.org/10.1016/j.ympev.2008.03.030
Pellegrino I, Negri A, Cucco M et al (2014) Phylogeography and Pleistocene refugia of the Little Owl Athene noctua inferred from mtDNA sequence data. Ibis 156:639–657. https://doi.org/10.1111/ibi.12162
Pellegrino I, Negri A, Boano G et al (2015) Evidence for strong genetic structure in European populations of the Little owl Athene noctua. J Avian Biol 46:462–475. https://doi.org/10.1111/jav.00679
Pentzold S, Tritsch C, Martens J et al (2013) Where is the line? Phylogeography and secondary contact of western Palearctic coal tits (Periparus ater: Aves, Passeriformes, Paridae). Zool Anz 252:367–382. https://doi.org/10.1016/j.jcz.2012.10.003
Perez-Tris J, Bensch S, Carbonell R et al (2004) Historical diversification of migration patterns in a passerine bird. Evolution (n Y) 58:1819–1832
Perktas U, Quintero E (2013) A wide geographical survey of mitochondrial DNA variation in the Great spotted woodpecker complex, Dendrocopos major ( Aves : Picidae ). Biol J Linn Soc 108:173–188
Perktas U, Barrowclough GF, Groth JG (2011) Phylogeography and species limits in the Green woodpecker complex (Aves: Picidae): multiple Pleistocene refugia and range expansion across Europe and the Near East. Biol J Linn Soc 104:710–723. https://doi.org/10.1111/j.1095-8312.2011.01750.x
Pestano J, Brown RP, Rodriguez F, Moreno A (2000) Mitochondrial DNA control region diversity in the endangered blue chaffinch, Fringilla teydea. Mol Ecol 9:1421–1425. https://doi.org/10.1046/j.1365-294x.2000.00990.x
Petit RJ, Hampe A, Cheddadi R (2005) Climate changes and tree phylogeography in the Mediterranean. Taxon 54:877–885. https://doi.org/10.2307/25065474
Pilot M, Branicki W, Jędrzejewski W et al (2010) Phylogeographic history of Grey wolves in Europe. BMC Evol Biol 10:104. https://doi.org/10.1186/1471-2148-10-104
Pitra C, Lieckfeldt D, Alonso JC (2000) Population subdivision in Europe’s great bustard inferred from mitochondrial and nuclear DNA sequence variation. Mol Ecol 9:1165–1170. https://doi.org/10.1046/j.1365-294x.2000.00983.x
Pons J-M, Olioso G, Cruaud C, Fuchs J (2011) Phylogeography of the Eurasian green woodpecker (Picus viridis). J Biogeogr 38:311–325. https://doi.org/10.1111/j.1365-2699.2010.02401.x
Pons JM, Thibault JC, Fournier J et al (2015) Genetic variation among Corsican and continental populations of the Eurasian treecreeper (Aves: Certhia familiaris) reveals the existence of a palaeoendemic mitochondrial lineage. Biol J Linn Soc 115:134–153. https://doi.org/10.1111/bij.12485
Pons J-M, Thibault J-C, Aymí R et al (2016) The role of western Mediterranean islands in the evolutionary diversification of the Spotted flycatcher Muscicapa striata, a long-distance migratory passerine species. J Avian Biol 47:386–398. https://doi.org/10.1111/jav.00859
Pons JM, Masson C, Olioso G, Fuchs J (2019b) Gene flow and genetic admixture across a secondary contact zone between two divergent lineages of the Eurasian Green Woodpecker Picus viridis. J Ornithol 160:935–945. https://doi.org/10.1007/s10336-019-01675-6
Pons J-M, Cibois A, Fournier J et al (2019a) Gene flow and genetic divergence among mainland and insular populations across the south-western range of the Eurasian treecreeper (Certhia familiaris, Aves). Biol J Linn Soc 126:447–461. https://doi.org/10.1093/biolinnean/bly200
Poulakakis N, Antoniou A, Mantziou G et al (2008) Population structure, diversity, and phylogeography in the near-threatened Eurasian Black Vultures Aegypius monachus (Falconiformes; Accipitridae) in Europe: insights from microsatellite and mitochondrial DNA variation. Biol J Linn Soc 95:859–872. https://doi.org/10.1111/j.1095-8312.2008.01099.x
Poulakakis N, Kapli P, Lymberakis P et al (2015) A review of phylogeographic analyses of animal taxa from the Aegean and surrounding regions. J Zool Syst Evol Res 53:18–32. https://doi.org/10.1111/jzs.12071
Prinsloo P, Robinson TJ (1992) Geographic mitochondrial DNA variation in the Rock hyrax, Procavia capensis. Mol Biol Evol 9:447–456. https://doi.org/10.1093/oxfordjournals.molbev.a040728
Questiau S, Gielly L, Clouet M, Taberlet P (1999) Phylogeographical evidence of gene flow among Common Crossbill (Loxia curvirostra, Aves, Fringillidae) populations at the continental level. Heredity (edinb) 83:196–205. https://doi.org/10.1046/j.1365-2540.1999.00551.x
Quinn TW (1992) The genetic legacy of Mother Goose—phylogeographic patterns of lesser snow goose Chen caerulescens caerulescens maternal lineages. Mol Ecol 1:105–117
R Development Core Team (2019) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Raković M, Neto JM, Lopes RJ et al (2019) Geographic patterns of mtDNA and Z-linked sequence variation in the Common Chiffchaff and the ‘chiffchaff complex.’ PLoS ONE 14:1–20. https://doi.org/10.1371/journal.pone.0210268
Rand AL (1948) Glaciation, an isolating factor in speciation. Evolution (n Y) 2:314–321. https://doi.org/10.2307/2405522
Randi E, Tabarroni C, Rimondi S et al (2003) Phylogeography of the Rock partridge (Alectoris graeca). Mol Ecol 12:2201–2214. https://doi.org/10.1046/j.1365-294X.2003.01899.x
Randler C, Förschler MI, Gonzalez J et al (2012) Phylogeography, pre-zygotic isolation and taxonomic status in the endemic Cyprus Wheatear Oenanthe cypriaca. J Ornithol 153:303–312. https://doi.org/10.1007/s10336-011-0744-8
Reeb CA, Avise JC (1990) A genetic discontinuity in a continuously distributed species: mitochondrial DNA in the American oyster, Crassostrea virginica. Genetics 124:397–406
Resano-Mayor J, Fernández-Martín Á, Hernández-Gómez S et al (2017) Integrating genetic and stable isotope analyses to infer the population structure of the White-winged Snowfinch Montifringilla nivalis in Western Europe. J Ornithol 158:395–405. https://doi.org/10.1007/s10336-016-1413-8
Rheindt FE, Edwards SV (2011) Genetic introgression: an integral but neglected component of speciation in birds. Auk 128:620–632. https://doi.org/10.1525/auk.2011.128.4.620
Riddle BR (2016) Comparative phylogeography clarifies the complexity and problems of continental distribution that drove A. R. Wallace to favor islands. Proc Natl Acad Sci 113:7970–7977. https://doi.org/10.1073/pnas.1601072113
Rodrigues P, Lopes RJ, Drovetski SV et al (2013) Phylogeography and genetic diversity of the Robin (Erithacus rubecula) in the Azores Islands: Evidence of a recent colonisation. J Ornithol 154:889–900. https://doi.org/10.1007/s10336-013-0953-4
Rodrigues P, Lopes RJ, Reis S et al (2014b) Genetic diversity and morphological variation of the Common chaffinch Fringilla coelebs in the Azores. J Avian Biol 45:167–178. https://doi.org/10.1111/j.1600-048X.2013.00229.x
Rodrigues P, Lopes RJ, Micael J et al (2014a) Genetic and morphometric diversity of the goldcrest (Regulus regulus) populations in the Azores. Zoology 117:383–391. https://doi.org/10.1016/j.zool.2014.07.001
Rodrigues P, Lopes RJ, Resendes R et al (2016) Genetic diversity of the Azores Blackbirds Turdus merula reveals multiple founder events. Acta Ornithol 51:221–234
Rodrigues TM, Andrade P, Vidal M et al (2019) No genetic differentiation, but less diversity, in the Iberian breeding population of the Eurasian Curlew (Numenius arquata). J Ornithol 160:17–25. https://doi.org/10.1007/s10336-018-1598-0
Rodríguez-Muñoz R, Mirol PM, Segelbacher G et al (2007) Genetic differentiation of an endangered Capercaillie (Tetrao urogallus) population at the Southern edge of the species range. Conserv Genet 8:659–670. https://doi.org/10.1007/s10592-006-9212-z
Rönkä N, Pakanen V-M, Blomqvist D et al (2019) Near panmixia at the distribution-wide scale but evidence of genetic differentiation in a geographically isolated population of the Terek Sandpiper Xenus cinereus. Ibis 161:632–647. https://doi.org/10.1111/ibi.12651
Roques S, Negro JJ (2005) MtDNA genetic diversity and population history of a dwindling raptorial bird, the Red kite (Milvus milvus). Biol Conserv 126:41–50. https://doi.org/10.1016/j.biocon.2005.04.020
Rösner S, Cimiotti DV, Brandl R (2014) Two sympatric lineages of the Raven Corvus corax jordansi coexist on the Eastern Canary Islands. J Ornithol 155:243–251. https://doi.org/10.1007/s10336-013-1006-8
Rozas J, Ferrer-Mata A, Sánchez-DelBarrio JC et al (2017) DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol Biol Evol 34:3299–3302. https://doi.org/10.1093/molbev/msx248
Rubinoff D, Holland BS (2005) Between two extremes: mitochondrial DNA is neither the panacea nor the nemesis of phylogenetic and taxonomic inference. Syst Biol 54:952–961. https://doi.org/10.1080/10635150500234674
Rull V (2011) Neotropical biodiversity: Timing and potential drivers. Trends Ecol Evol 26:508–513. https://doi.org/10.1016/j.tree.2011.05.011
Ruokonen M, Aarvak T, Madsen J (2005) Colonization history of the high-arctic Pink-footed goose Anser brachyrhynchus. Mol Ecol 14:171–178. https://doi.org/10.1111/j.1365-294X.2004.02380.x
Rutkowski R, Niewęgłowski H, Dziedzic R et al (2005) Genetic variability of polish population of the Capercaillie Tetrao urogallus. Acta Ornithol 40:27–34
Rutkowski R, Keller M, Jagołkowska P (2012) Population genetics of the Hazel Hen Bonasa bonasia in Poland assessed with non-invasive samples. Cent Eur J Biol 7:759–775. https://doi.org/10.2478/s11535-012-0051-2
Rutkowski R, Krofel M, Giannatos G et al (2015) A European concern? genetic structure and expansion of Golden jackals (Canis aureus) in Europe and the Caucasus. PLoS ONE 10:1–22. https://doi.org/10.1371/journal.pone.0141236
Rutkowski R, Jagołkowska P, Zawadzka D, Bogdanowicz W (2016) Impacts of forest fragmentation and post-glacial colonization on the distribution of genetic diversity in the Polish population of the Hazel grouse Tetrastes bonasia. Eur J Wildl Res 62:293–306. https://doi.org/10.1007/s10344-016-1002-4
Rutkowski R, Zawadzka D, Suchecka E, Merta D (2017) Conservation genetics of the Capercaillie in Poland—delineation of conservation units. PLoS ONE 12:1–23. https://doi.org/10.1371/journal.pone.0174901
Rutkowski R, Pałucki A, Dulisz B et al (2019) Conservation genetics of the Black Grouse Tetrao tetrix in Poland — distribution of genetic diversity among the last populations. Acta Ornithol 53:181–204
Sætre G-P, Cuevas A, Hermansen JS et al (2017) Rapid polygenic response to secondary contact in a hybrid species. Proc R Soc B Biol Sci 284:20170365. https://doi.org/10.1098/rspb.2017.0365
Sahlsten J, Thörngren H, Höglund J (2008) Inference of hazel grouse population structure using multilocus data: a landscape genetic approach. Heredity (edinb) 101:475
Salinas P, Morinha F, Literak I et al (2021) Genetic diversity, differentiation and historical origin of the isolated population of rooks Corvus frugilegus in Iberia. J Avian Biol. https://doi.org/10.1111/jav.02689
Salzburger W, Martens J, Nazarenko AA et al (2002) Phylogeography of the Eurasian Willow Tit (Parus montanus) based on DNA sequences of the mitochondrial cytochrome b gene. Mol Phylogenet Evol 24:26–34. https://doi.org/10.1016/S1055-7903(02)00266-X
Sangster G, Rodríguez-Godoy F, Roselaar CS et al (2016) Integrative taxonomy reveals Europe’s rarest songbird species, the Gran Canaria blue chaffinch Fringilla polatzeki. J Avian Biol 47:159–166. https://doi.org/10.1111/jav.00825
Scandura M, Iacolina L, Crestanello B et al (2008) Ancient vs. recent processes as factors shaping the genetic variation of the European wild boar: are the effects of the last glaciation still detectable? Mol Ecol 17:1745–1762. https://doi.org/10.1111/j.1365-294X.2008.03703.x
Schweizer M, Shirihai H, Schmaljohann H, Kirwan GM (2018) Phylogeography of the House Bunting complex: discordance between species limits and genetic markers. J Ornithol 159:47–61. https://doi.org/10.1007/s10336-017-1501-4
Segelbacher G, Piertney S (2007) Phylogeography of the European capercaillie (Tetrao urogallus) and its implications for conservation. J Ornithol 148:269–274. https://doi.org/10.1007/s10336-007-0153-1
Segelbacher G, Storch I (2002) Capercaillie in the Alps: genetic evidence of metapopulation structure and population decline. Mol Ecol 11:1669–1677. https://doi.org/10.1046/j.1365-294X.2002.01565.x
Segelbacher G, Höglund J, Storch I (2003) From connectivity to isolation: genetic consequences of population fragmentation in capercaillie across Europe. Mol Ecol 12:1773–1780. https://doi.org/10.1046/j.1365-294X.2003.01873.x
Senar JC, Borras A, Cabrera J et al (2006) Local differentiation in the presence of gene flow in the citril finch Serinus citrinella. Biol Lett 2:85–87. https://doi.org/10.1098/rsbl.2005.0412
Sequeira F, Alexandrino J, Weiss S, Ferrand N (2008) Documenting the advantages and limitations of different classes of molecular markers in a well-established phylogeographic context: Lessons from the Iberian endemic Golden-striped salamander, Chioglossa lusitanica (Caudata: Salamandridae). Biol J Linn Soc 95:371–387. https://doi.org/10.1111/j.1095-8312.2008.01060.x
Shannon TJ, McGowan RY, Zonfrillo B et al (2014) A genetic screen of the island races of Wren Troglodytes troglodytes in the North-east Atlantic. Bird Study 61:135–142. https://doi.org/10.1080/00063657.2014.894493
Shaw KL, Gillespie RG (2016) Comparative phylogeography of oceanic archipelagos: hotspots for inferences of evolutionary process. Proc Natl Acad Sci 113:7986–7993. https://doi.org/10.1073/pnas.1601078113
Shirihai H, Svensson L (2018) Handbook of western palearctic birds. Bloomsbury Publishing , London
Silva SM, Peterson AT, Carneiro L et al (2019) A dynamic continental moisture gradient drove Amazonian bird diversification. Sci Adv. https://doi.org/10.1126/sciadv.aat5752
Sittenthaler M, Kunz F, Szymusik A et al (2018) Fine-scale genetic structure in an eastern Alpine black grouse Tetrao tetrix metapopulation. J Avian Biol 49:1–14. https://doi.org/10.1111/jav.01681
Smith BT, Seeholzer GF, Harvey MG et al (2017) A latitudinal phylogeographic diversity gradient in birds. PLoS Biol 15:e2001073–e2001073. https://doi.org/10.1371/journal.pbio.2001073
Smith BT, Bryson RW, Mauck WM et al (2018) Species delimitation and biogeography of the gnatcatchers and gnatwrens (Aves: Polioptilidae). Mol Phylogenet Evol 126:45–57. https://doi.org/10.1016/j.ympev.2018.03.012
Sommer RS, Fahlke JM, Schmölcke U et al (2009) Quaternary history of the European roe deer Capreolus capreolus. Mamm Rev 39:1–16. https://doi.org/10.1111/j.1365-2907.2008.00137.x
Somveille M, Wikelski M, Beyer RM et al (2020) Simulation-based reconstruction of global bird migration over the past 50,000 years. Nat Commun 11:801. https://doi.org/10.1038/s41467-020-14589-2
Song G, Zhang R, Dubay SG et al (2016) East Asian allopatry and north Eurasian sympatry in Long-tailed Tit lineages despite similar population dynamics during the late Pleistocene. Zool Scr 45:115–126. https://doi.org/10.1111/zsc.12148
Song G, Zhang R, Alström P et al (2018) Complete taxon sampling of the avian genus Pica (magpies) reveals ancient relictual populations and synchronous Late-Pleistocene demographic expansion across the Northern Hemisphere. J Avian Biol. https://doi.org/10.1111/jav.01612
Song G, Zhang R, Machado-Stredel F et al (2020) Great journey of Great Tits (Parus major group): Origin, diversification and historical demographics of a broadly distributed bird lineage. J Biogeogr 47:1585–1598. https://doi.org/10.1111/jbi.13863
Sonsthagen SA, Chesser RT, Bell DA, Dove CJ (2012) Hybridization among Arctic white-headed gulls (Larus spp.) obscures the genetic legacy of the Pleistocene. Ecol Evol 2:1278–1295. https://doi.org/10.1002/ece3.240
Spurgin LG, Bosse M, Adriaensen F et al (2019) The Great tit HapMap project: a continental-scale analysis of genomic variation in a songbird. bioRxiv. https://doi.org/10.1101/561399
Stervander M, Illera JC, Kvist L et al (2015) Disentangling the complex evolutionary history of the Western Palearctic blue tits (Cyanistes spp. )—phylogenomic analyses suggest radiation by multiple colonization events and subsequent isolation. Mol Ecol 24:2477–2494. https://doi.org/10.1111/mec.13145
Stiller J, Zhang G (2019) Comparative phylogenomics, a stepping stone for bird biodiversity studies. Diversity 11:115. https://doi.org/10.3390/d11070115
Suárez NM, Betancor E, Klassert TE et al (2009) Phylogeography and genetic structure of the Canarian common chaffinch (Fringilla coelebs) inferred with mtDNA and microsatellite loci. Mol Phylogenet Evol 53:556–564. https://doi.org/10.1016/j.ympev.2009.07.018
Swenson JE, Taberlet P, Bellemain E (2011) Genetics and conservation of European brown bears Ursus arctos. Mamm Rev 41:87–98. https://doi.org/10.1111/j.1365-2907.2010.00179.x
Taberlet P, Fumagalli L, Wust-Saucy AG, Cosson JF (1998) Comparative phylogeography and postglacial colonization routes in Europe. Mol Ecol 7:453–464. https://doi.org/10.1046/j.1365-294x.1998.00289.x
Tan HZ, Ng EYX, Tang Q et al (2019) Population genomics of two congeneric Palaearctic shorebirds reveals differential impacts of Quaternary climate oscillations across habitats types. Sci Rep 9:18172. https://doi.org/10.1038/s41598-019-54715-9
Thibault J-C, Cibois A, Prodon R, Pasquet E (2016) Quaternary history of an endemic passerine bird on Corsica Island: glacial refugium and impact of recent forest regression. Quat Res 85:271–278. https://doi.org/10.1016/j.yqres.2016.01.002
Thies L, Tomkovich P, dos Remedios N et al (2018) Population and subspecies differentiation in a high latitude breeding wader, the Common Ringed Plover Charadrius hiaticula. Ardea 106:163–176
Tiedemann R, Paulus KB, Scheer M et al (2004) Mitochondrial DNA and microsatellite variation in the Eider duck (Somateria mollissima) indicate stepwise postglacial colonization of Europe and limited current long-distance dispersal. Mol Ecol 13:1481–1494. https://doi.org/10.1111/j.1365-294X.2004.02168.x
Trier CN, Hermansen JS, Sætre G-P, Bailey RI (2014) Evidence for mito-nuclear and sex-linked reproductive barriers between the hybrid italian sparrow and its parent species. PLOS Genet 10:e1004075
Tritsch C, Stuckas H, Martens J et al (2018) Gene flow in the European coal tit, Periparus ater (Aves: Passeriformes): low among Mediterranean populations but high in a continental contact zone. Biol J Linn Soc 124:319–338. https://doi.org/10.1093/biolinnean/bly043
Tryfonopoulos GA, Thanou EG, Fraguedakis-Tsolis SE, Chondropoulos BP (2010) New data on the distribution and genetic structure of Greek moles of the genus Talpa (Mammalia, Talpidae). J Zool Syst Evol Res 48:188–193. https://doi.org/10.1111/j.1439-0469.2009.00533.x
Tsoupas A, Andreadou M, Papakosta MA et al (2019) Phylogeography of Martes foina in Greece. Mamm Biol 95:59–68. https://doi.org/10.1016/j.mambio.2019.02.004
Väli Ü (2004) The Greater Spotted Eagle Aquila clanga and the lesser spotted eagle A. pomarina: taxonomy, phylogeography and ecology. Tartu University, Tartu
Väli Ü, Treinys R, Poirazidis K (2004) Genetic structure of Greater Aquila clanga and Lesser Spotted Eagle A. pomarina populations: implications for phylogeography and conservation. In: Chancellor R, Meyburg B-U (eds) Raptors worldwide: proceedings of the VI World Conference on Birds of Prey and Owls, Budapest, Hungary, 18–23 May 2003. Raptors Worldwide WWGBP/MME, pp 473–482
Verkuil YI, Piersma T, Jukema J et al (2012) The interplay between habitat availability and population differentiation: a case study on genetic and morphological structure in an inland wader (Charadriiformes). Biol J Linn Soc 106:641–656. https://doi.org/10.1111/j.1095-8312.2012.01878.x
Vili N, Chavko J, Szabó K et al (2009) Genetic structure of the Imperial Eagle (Aquila heliaca) population in Slovakia. Slovak Raptor J 3:21–28
Voelker G, Outlaw RK, Bowie RCK (2010) Pliocene forest dynamics as a primary driver of African bird speciation. Glob Ecol Biogeogr 19:111–121. https://doi.org/10.1111/j.1466-8238.2009.00500.x
Voelker G, Marks BD, Kahindo C et al (2013) River barriers and cryptic biodiversity in an evolutionary museum. Ecol Evol 3:536–545. https://doi.org/10.1002/ece3.482
von Rönn JAC, Shafer ABA, Wolf JBW (2016) Disruptive selection without genome-wide evolution across a migratory divide. Mol Ecol 25:2529–2541. https://doi.org/10.1111/mec.13521
Wang E, van Wijk RE, Braun MS, Wink M (2017) Gene flow and genetic drift contribute to high genetic diversity with low phylogeographical structure in European Hoopoes (Upupa epops). Mol Phylogenet Evol 113:113–125. https://doi.org/10.1016/j.ympev.2017.05.018
Wang E, Bairlein F, Wink M (2018) Phylogeography of the Northern Wheatear Oenanthe oenanthe inferred from genome sequencing data. Vogelwarte 56:387–388
Wang E, Zhang D, Braun MS et al (2020) Can Mitogenomes of the Northern Wheatear (Oenanthe oenanthe) reconstruct its phylogeography and reveal the origin of migrant birds? Sci Rep 10:9290. https://doi.org/10.1038/s41598-020-66287-0
Webb T, Bartlein PJ (1992) Global changes during the last 3 million years: climatic controls and biotic responses. Annu Rev Ecol Syst 23:141–173. https://doi.org/10.1146/annurev.es.23.110192.001041
Webb WC, Marzluff JM, Omland KE (2011) Random interbreeding between cryptic lineages of the Common Raven: evidence for speciation in reverse. Mol Ecol 20:2390–2402. https://doi.org/10.1111/j.1365-294X.2011.05095.x
Weiss S, Ferrand N (2007) Phylogeography of Southern European Refugia. Springer, Dordrecht
Wenink PW, Baker AJ, Tilanus MG (1993) Hypervariable-control-region sequences reveal global population structuring in a long-distance migrant shorebird, the Dunlin (Calidris alpina). Proc Natl Acad Sci 90:94–98. https://doi.org/10.1073/pnas.90.1.94
Wenink PW, Baker AJ, Tilanus MG (1994) Mitochondrial control-region sequences in two shorebird species, the Turnstone and the Dunlin, and their utility in population genetic studies. Mol Biol Evol 11:22–31. https://doi.org/10.1093/oxfordjournals.molbev.a040089
Wenink PW, Baker AJ, Rösner H-U, Tilanus MGJ (1996) Global mitochondrial DNA phylogeography of Holarctic breeding Dunlins (Calidris alpina). Evolution (n Y) 50:318–330. https://doi.org/10.1111/j.1558-5646.1996.tb04495.x
Wennerberg L, Marthinsen G, Lifjeld JT (2008) Conservation genetics and phylogeography of Southern dunlins Calidris alpina schinzii. J Avian Biol 39:423–437. https://doi.org/10.1111/j.2008.0908-8857.04351.x
Wink M (2008) Taxonomy and genetics: Phylogenetic and phylogeographic relationships. In: Van Nieuwenhuyse D, Genot JC, Johnson DH (eds) The Little Owl-Conservation, Ecology and Behaviour of Athene noctua. Cambridge University Press, Cambridge, pp 38–44
Wink M (2018b) Phylogeny of Falconidae and phylogeography of Peregrine Falcons. Ornis Hungarica 26:27–37. https://doi.org/10.1515/orhu-2018-013
Wink M (2018a) Biodiversity on oceanic islands—evolutionary records of past migration events. In: Funke J, Wink M (eds) Heidelberger Jahrbücher online: Perspektiven der Mobilität. Gesellschaft der Freunde Universität Heidelberg, Heidelberger, pp 119–155
Wink M, Sauer-Gürth H, Pepler D (2004) Phylogeographic relationships of the Lesser Kestrel Falco naumanni in breeding and wintering quarters, inferred from nucleotide sequences of mitochondrial cytochrome b gene. In: Chancellor R, Meyburg B-U (eds) Raptors worldwide: proceedings of the VI World Conference on Birds of Prey and Owls, Budapest, Hungary, 18–23 May 2003. Raptors Worldwide WWGBP/MME, pp 505–510
Winney BJ, Litton CD, Parkin DT, Feare CJ (2001) The subspecific origin of the inland breeding colonies of the cormorant Phalacrocorax carbo in Britain. Heredity (edinb) 86:45–53. https://doi.org/10.1046/j.1365-2540.2001.00807.x
Zehtindjiev P, Ilieva M, Hansson B et al (2011) Population genetic structure in the paddyfield warbler (Acrocephalus agricola Jerd.). Curr Zool 57:63–71. https://doi.org/10.1093/czoolo/57.1.63
Zhan X, Dixon A, Batbayar N et al (2015) Exonic versus intronic SNPs: contrasting roles in revealing the population genetic differentiation of a widespread bird species. Heredity (edinb) 114:1–9
Zink RM (1996) Comparative phylogeography in North American birds. Evolution (n Y) 50:308–317. https://doi.org/10.2307/2410802
Zink RM (1997) Phylogeographic studies of North American birds. In: Mindell DP (ed) Avian molecular evolution and systematics. Academic Press, London, pp 301–324
Zink RM, Barrowclough GF (2008) Mitochondrial DNA under siege in avian phylogeography. Mol Ecol 17:2107–2121. https://doi.org/10.1111/j.1365-294X.2008.03737.x
Zink RM, Drovetski SV, Rohwer S (2002a) Phylogeographic patterns in the Great spotted woodpecker Dendrocopos major across Eurasia. J Avian Biol 33:175–178. https://doi.org/10.1034/j.1600-048X.2002.330208.x
Zink RM, Rohwer S, Drovetski S et al (2002b) Holarctic phylogeography and species limits of three-toed woodpeckers. Condor 104:167–170. https://doi.org/10.1650/0010-5422(2002)104[0167:hpaslo]2.0.co;2
Zink RM, Drovetski SV, Questiau S et al (2003) Recent evolutionary history of the Bluethroat (Luscinia svecica) across Eurasia. Mol Ecol 12:3069–3075. https://doi.org/10.1046/j.1365-294X.2003.01981.x
Zink RM, Klicka J, Barber BR (2004) The tempo of avian diversification during the Quaternary. Philos Trans R Soc London Ser B Biol Sci 359:215–220. https://doi.org/10.1098/rstb.2003.1392
Zink RM, Drovetski SV, Rohwer S (2006) Selective neutrality of mitochondrial ND2 sequences, phylogeography and species limits in Sitta europaea. Mol Phylogenet Evol 40:679–686. https://doi.org/10.1016/j.ympev.2005.11.002
Zink RM, Pavlova A, Drovetski S, Rohwer S (2008) Mitochondrial phylogeographies of five widespread Eurasian bird species. J Ornithol 149:399–413. https://doi.org/10.1007/s10336-008-0276-z
Zuccon D, Pons J-M, Boano G et al (2020) Type specimens matter: new insights on the systematics, taxonomy and nomenclature of the subalpine warbler (Sylvia cantillans) complex. Zool J Linn Soc 190:314–341. https://doi.org/10.1093/zoolinnean/zlz169
Acknowledgements
We would like to express our sincere gratitude to Carina Carneiro de Melo Moura, who offered great support and valuable ideas at an incipient stage of the study. We further thank Erjia Wang, Roberto Carlos Frias-Soler, Hedwig Sauer-Gürth, Beate Waibel and Ivan Starikov for burgeoning discussions on the topic of phylogeography and for useful ideas to improve this review. We also acknowledge Petra Fellhauer for technical support. Alexandru Tomazatos provided constructive comments on an earlier draft of the manuscript. Kylynn Clare kindly helped with proofreading and English grammar corrections. We acknowledge the help of Franz Bairlein, Jan T. Lifjeld, Per Alström and one anonymous reviewer for their contribution to the peer-review of this work. Mare Haider, Pelin and Irmgard Yildiz, Philipp Scheffzek, Dietlinde Schiebel and Markus Schirmer provided logistical support at the initial stage of the study. LP would like to dedicate this article to his esteemed ornithology mentor, Prof. Eugen Petrescu from Danube Delta—Romania, who triggered his passion and curiosity on bird science, providing constant guidance in the past 19 years.
Funding
Open Access funding enabled and organized by Projekt DEAL. LP received financial support from the Deutsche Ornithologen-Gessellschaft (DO-G)—Abs-Halbstipendium, the Erwin-Stresemann Förderung, the Irmgard und Michael Abs Stiftung, Graduate Academy Heidelberg, DAAD & STIBET PhD Completion Grants. MW received additional funding from the German Science Foundation (DFG), German Academic Exchange Service (DAAD), COST, Chinese Scholarship Council (CSC), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), National Council of Science and Technology (CONACYT), Science and Technology Development Fund (STDF) and German Ornithologist Society (DO-G).
Author information
Authors and Affiliations
Contributions
LP and MW conceived the study. LP collected and analyzed the data, and wrote the manuscript. MW provided regular intellectual input, reviewed drafts of the paper and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interests
The authors declare no conflict of interests.
Additional information
Communicated by J. T. Lifjeld.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pârâu, L.G., Wink, M. Common patterns in the molecular phylogeography of western palearctic birds: a comprehensive review. J Ornithol 162, 937–959 (2021). https://doi.org/10.1007/s10336-021-01893-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10336-021-01893-x