Abstract
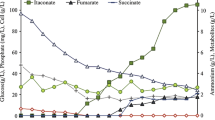
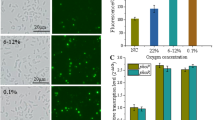
The dissolved oxygen (DO) level of a culture of Corynebacterium glutamicum (C. glutamicum) in a bioreactor has a significant impact on the cellular redox potential and the distribution of energy and metabolites. In this study, to gain a deeper understanding of the effects of DO on the metabolism of C. glutamicum, we sought to systematically explore the influence of different DO concentrations on genetic regulation and metabolism through transcriptomic analysis. The results revealed that after 20 h of fermentation, oxygen limitation enhanced the glucose metabolism, pyruvate metabolism and carbon overflow, and restricted NAD+ availability. A high oxygen supply enhanced the TCA cycle and reduced glyoxylate metabolism. Several key genes involved in response of C. glutamicum to different oxygen concentrations were examined, which provided suggestions for target site modifications in developing optimized oxygen supply strategies. These data provided new insights into the relationship between oxygen supply and metabolism of C. glutamicum.






Similar content being viewed by others
Change history
08 February 2020
The authors have retracted this article.
08 February 2020
The authors have retracted this article.
References
Audic S, Claverie JM (1997) The significance of digital gene expression profiles. Genome Res 7:986–995
Bai Y, Zhou PP, Fan P, Zhu YM, Tong Y, Wang HB, Yu LJ (2015) Four-stage dissolved oxygen strategy based on multi-scale analysis for improving spinosad yield by Saccharopolyspora spinosa ATCC49460. Microb Biotechnol 8:561–568
Benjamini Y, Yekutieli D (2001) The control of the false discovery rate in multiple testing under dependency. Ann Stat 29:1165–1188
Berríos-Rivera Bennett GN, San KY (2002) Metabolic engineering of Escherichia coli: increase of NADH availability by overexpressing an NAD(+)-dependent formate dehydrogenase. Metab Eng 4:217–229
Bott M, Niebisch A (2003) The respiratory chain of Corynebacterium glutamicum. J Biotechnol 104:129–153
Brinkrolf K, Brune I, Tauch A (2007) The transcriptional regulatory network of the amino acid producer Corynebacterium glutamicum. J Biotechnol 129:191–211
Buchholz J, Graf M, Freund A, Busche T, Kalinowski J, Blombach B, Takors R (2014) CO2/HCO3 − perturbations of simulated large scale gradients in a scale-down device cause fast transcriptional responses in Corynebacterium glutamicum. Appl Microbiol Biotechnol 98:8563–8572
Conesa A, Gotz S, Garcia-Gomez JM, Terol J, Talon M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21:3674–3676
Eikmanns BJ, Rittmann D, Sahm H (1995) Cloning, sequence analysis, expression, and inactivation of the Corynebacterium glutamicum icd gene encoding isocitrate dehydrogenase and biochemical characterization of the enzyme. J Bacteriol 177:774–782
Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 95:14863–14868
Garcia-Ochoa F, Gomez E (2009) Bioreactor scale-up and oxygen transfer rate in microbial processes: an overview. Biotechnol Adv 27:153–176
Garcia-Ochoa F, Gomez E, Alcon A, Santos VE (2013) The effect of hydrodynamic stress on the growth of Xanthomonas campestris cultures in a stirred and sparged tank bioreactor. Bioprocess Biosyst Eng 36:911–925
Garcia-Ochoa F, Gomez E, Santos VE, Merchuk JC (2010) Oxygen uptake rate in microbial processes: an overview. Biochem Eng J 49:289–307
Ge XY, Xu Y, Chen X, Zhang LY (2015) Regulation of metabolic flux in Lactobacillus casei for lactic acid production by overexpressed ldhL gene with two-stage oxygen supply strategy. J Microbiol Biotechnol 25:81–88
Gopinath V, Murali A, Dhar KS, Nampoothiri KM (2012) Corynebacterium glutamicum as a potent biocatalyst for the bioconversion of pentose sugars to value-added products. Appl Microbiol Biotechnol 93:95–106
Hara KY, Kondo A (2015) ATP regulation in bioproduction. Microb Cell Fact 14:198
Hermann T (2003) Industrial production of amino acids by coryneform bacteria. J Biotechnol 104:155–172
Hong EJ, Kim Kim ES, Kim Y, Lee HS (2016) Involvement of the osrR gene in the hydrogen peroxide-mediated stress response of Corynebacterium glutamicum. Res Microbiol 167:20–28
II’chenko AP, Shishkanova NV, Chernyavskaya OG, Finogenova TV (1998) Oxygen concentration as a factor controlling central metabolism and citric acid biosynthesis in the yeast Yarrowia lipolytica grown on ethanol. Microbiology 67:241–244
Joshi J, Elias C, Patole M (1996) Role of hydrodynamic shear in the cultivation of animal, plant and microbial cells. Biochem Eng J 62:121–141
Kalinowski J, Bathe B, Bartels D, Bischoff N, Bott M, Burkovski A, Dusch N, Eggeling L, Eikmanns BJ, Gaigalat L, Goesmann A, Hartmann M, Huthmacher K, Krämer R, Linke B, McHardy AC, Meyer F, Möckel B, Pfefferle W, Pühler A, Rey DA, Rückert C, Rupp O, Sahm H, Wendisch VF, Wiegräbe I, Tauch A (2003) The complete Corynebacterium glutamicum ATCC 13032 genome sequence and its impact on the production of l-aspartate-derived amino acids and vitamins. J Biotechnol 104:5–25
Kim HI, Nam JY, Cho JY, Lee CS, Park YJ (2013) Next-generation sequencing-based transcriptome analysis of l-lysine-producing Corynebacterium glutamicum ATCC 21300 strain. J Microbiol 51:877–880
Kim SY, Kim JH, Oh DK (1997) Improvement of xylitol production by controlling oxygen supply in Candida parapsilosis. J Ferment Bioeng 83:267–270
Koch-Koerfges A, Pfelzer N, Platzen L, Oldiges M, Bott M (2013) Conversion of Corynebacterium glutamicum from an aerobic respiring to an aerobic fermenting bacterium by inactivation of the respiratory chain. Biochim Biophys Acta 1827:699–708
Li R, Yu C, Li Y, Lam TW, Yiu SM, Kristiansen K, Wang J (2009) SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25:1966–1967
Lietzan AD, Lin Y, St Maurice M (2014) The role of biotin and oxamate in the carboxyltransferase reaction of pyruvate carboxylase. Arch Biochem Biophys 562:70–79
Lindner SN, Knebel S, Pallerla SR, Schoberth SM, Wendisch VF (2010) Cg2091 encodes a polyphosphate/ATP-dependent glucokinase of Corynebacterium glutamicum. Appl Microbiol Biotechnol 87:703–713
Liu X, Yang Y, Zhang W, Sun Y, Peng F, Jeffrey L, Harvey L, McNeil B, Bai Z (2015) Expression of recombinant protein using Corynebacterium Glutamicum: progress, challenges and applications. Crit Rev Biotechnol 25:1–13
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−∆∆CT method. Methods 25:402–408
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31:426–428
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5:621–628
Park SY, Kim HK, Yoo SK, Oh TK, Lee JK (2000) Characterization of glk, a gene coding for glucose kinase of Corynebacterium glutamicum. FEMS Microbiol Lett 188:209–215
Pauling J, Röttger R, Tauch A, Azevedo V, Baumbach J (2012) CoryneRegNet 6.0—updated database content, new analysis methods and novel features focusing on community demands. Nucleic Acids Res 40:D610–D614
Pfeifer-Sancar K, Mentz A, Rückert C, Kalinowski J (2013) Comprehensive analysis of the Corynebacterium glutamicum transcriptome using an improved RNAseq technique. BMC Genom 14:888
Pinto AC, Melo-Barbosa HP, Miyoshi A, Silva A, Azevedo V (2011) Application of RNA-seq to reveal the transcript profile in bacteria. Genet Mol Res 10:1707–1718
Saier MH Jr, Reizer J (1994) The bacterial phosphotransferase system: new frontiers 30 years later. Mol Microbiol 13:755–764
Saldanha AJ (2004) Java Treeview—extensible visualization of microarray data. Bioinformatics 20:3246–3248
Shimizu H, Tanaka H, Nakato A, Nagahisa K, Kimura E, Shioya S (2003) Effects of the changes in enzyme activities on metabolic flux redistribution around the 2-oxoglutarate branch in glutamate production by Corynebacterium glutamicum. Bioprocess Biosyst Eng 25:291–298
Siezen RJ, Wilson G, Todt T (2010) Prokaryotic whole-transcriptome analysis: deep sequencing and tiling arrays. J Microbial Biotechnol 3:125–130
Tang Y, Zhong J (2003) Role of oxygen supply in submerged fermentation of Ganoderma lucidum for production of Ganoderma polysaccharide and ganoderic acid. Enzyme Microb Tech 32:478–484
Tsai PS, Hatzimanikatis V, Bailey JE (1996) Effect of Vitreoscilla hemoglobin dosage on microaerobic Escherichia coli carbon and energy metabolism. Biotechnol Bioeng 49:139–150
Wang JY (2002) Biochemistry. Higher Education Press, Beijing (in Chinese)
Wang Z, Gerstein M, Snyder M (2009) RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet 10:57–63
Wendisch VF, Bott M, Kalinowski J, Oldiges M, Wiechert W (2006) Emerging Corynebacterium glutamicum systems biology. J Biotechnol 124:74–92
Wimpenny JWT, Firth A (1972) Levels of nicotinamide adenine dinucleotide and reduced nicotinamide adenine dinucleotide in facultative bacteria and the effect of oxygen. J Bacteriol 111:24–32
Xu H, Dou W, Xu H, Zhang X, Rao Z, Shi Z (2009) A two-stage oxygen supply strategy for enhanced l-arginine production by Corynebacterium crenatum based on metabolic fluxes analysis. J Biochem Eng J 43:41–51
Xu Y, Zhong JJ (2011) Significance of oxygen supply in production of a novel antibiotic by Pseudomonas sp. Bioresour Technol 102:9167–9174
Yamamoto S, Sakai M, Inui M, Yukawa H (2011) Diversity of metabolic shift in response to oxygen deprivation in Corynebacterium glutamicum and its close relatives. Appl Microbiol Biotechnol 90:1051–1061
Yang JK, Xiong W, Xu L, Li J, Zhao XJ (2015) Constitutive expression of Campylobacter jejuni truncated hemoglobin CtrHb improves the growth of Escherichia coli cell under aerobic and anaerobic conditions. Enzyme Microb Technol 75–76:64–70
Ye J, Fang L, Zheng H, Zhang Y, Chen J, Zhang Z, Wang J, Li S, Li R, Bolund L, Wang J (2006) WEGO: a web tool for plotting GO annotations. Nucleic Acids Res (Web Server issue) 34:W293–297
Yegneswaran PK, Gray MR, Thompson BG (1991) Effect of dissolved oxygen control on growth and antibiotic production in Streptomyces clavuligerus fermentations. Biotechnol Progr 7:246–250
Yu WB, Gao SH, Yin CY, Zhou Y, Ye BC (2011) Comparative transcriptome analysis of Bacillus subtilis responding to dissolved oxygen in adenosine fermentation. PLoS One 6:e20092
Zhang Y, French SL, Beyer AL, Schneider DA (2016) The transcription factor THO promotes transcription initiation and elongation by RNA polymerase I. J Biol Chem 291:3010–3018
Acknowledgements
This study was funded by the National Basic Research Program of China (973 Program) (Grant Number 2013CB733602), the Fundamental Research Funds for the Central Universities (Grant Number JUSRP51401A), the National Natural Science Foundation of China (Grant Number 31570034), and the Natural Science Foundation of Jiangsu Province (Grant Number BK20150148).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
X. Liu and S. Yang are contributed equally to this work.
The authors have retracted this article [1] because a significant portion of the study has been previously published in Chinese [2]. This article is therefore redundant. All authors agree to this retraction.
[1] Liu, X., Yang, S., Wang, F. et al. J Ind Microbiol Biotechnol (2017) 44: 181. https://doi.org/10.1007/s10295-016-1854-3
[2] Yang YK, Wang F, Sun Y. Effect of different dissolved oxygen concentrations on metabolism in Corynebacterium glutamicum. Microbiology China. (2016) ;43:11. 10.13344/j.microbiol.china.150975.
Electronic supplementary material
Below is the link to the electronic supplementary material.
About this article
Cite this article
Liu, X., Yang, S., Wang, F. et al. RETRACTED ARTICLE: Comparative analysis of the Corynebacterium glutamicum transcriptome in response to changes in dissolved oxygen levels. J Ind Microbiol Biotechnol 44, 181–195 (2017). https://doi.org/10.1007/s10295-016-1854-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-016-1854-3