Abstract
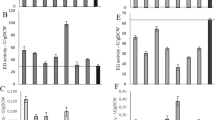
Enzyme engineering was performed to link the β-glucosidase enzyme (BGL1) from Saccharomycopsis fibuligera to the cellulose-binding domain (CBD2) of Trichoderma reesei cellobiohydrolase (CBHII) to investigate the effect of a fungal CBD on the enzymatic characteristics of this non-cellulolytic yeast enzyme. Recombinant enzymes were constructed with single and double copies of CBD2 fused at the N-terminus of BGL1 to mimic the two-domain organization displayed by cellulolytic enzymes in nature. The engineered S. fibuligera β-glucosidases were expressed in Saccharomyces cerevisiae under the control of phosphoglycerate-kinase-1 promoter (PGK1 P ) and terminator (PGK1 T ) and yeast mating pheromone α-factor secretion signal (MFα1 S ). The secreted enzymes were purified and characterized using a range of cellulosic and non-cellulosic substrates to illustrate the effect of the CBD on their enzymatic activity. The results indicated that the recombinant enzymes of BGL1 displayed a 2–4-fold increase in their hydrolytic activity toward cellulosic substrates like avicel, amorphous cellulose, bacterial microcrystalline cellulose, and carboxy methyl cellulose in comparison with the native enzyme. The organization of the CBD in these recombinant enzymes also resulted in enhanced substrate affinity, molecular flexibility and synergistic activity, thereby improving the ability of the enzymes to act on and hydrolyze cellulosic substrates, as characterized by adsorption, kinetics, thermal stability, and scanning electron microscopic analyses.



Similar content being viewed by others
References
Arai T, Araki R, Tanaka A, Karita S, Kimura T, Sakka K, Ohmiya K (2003) Characterization of a cellulase containing a family 30 carbohydrate-binding module (CBM) derived from Clostridium thermocellum CelJ: importance of the CBM to cellulose hydrolysis. J Bacteriol 185:504–512
Bellstedt DU, Human PA, Rowland GF, van der Merwe KJ (1987) Acid-treated, naked bacteria as immune carriers for protein antigens. J Immunol Methods 98:249–255
Boisset C, Fraschini C, Schulein M, Henrissat B, Chanzy H (2000) Imaging the enzymatic digestion of bacterial cellulose ribbons reveals the endo character of the cellobiohydrolase Cel6A from Humicola insolens and its mode of synergy with cellobiohydrolase Cel7A. Appl Environ Microbiol 66:1444–1452
Gal L, Pages S, Gaudin C, Belaich A, Reverbel-Leroy C, Tardif C, Belaich JP (1997) Characterization of the cellulolytic complex (cellulosome) produced by Clostridium cellulolyticum. Appl Environ Microbiol 63:903–909
Gietz RD, Woods RA (2002) Transformation of yeast by lithium acetate/single-stranded carrier DNA/polyethylene glycol method. Methods Enzymol 350:87–96
Gundllapalli Moses SB, Cordero Otero RR, Pretorius IS (2005) Domain engineering of Saccharomyces cerevisiae exoglucanases. Biotechnol Lett 27:355–362
Kataeva IA, Blum DL, Li XL, Ljungdahl LG (2001) Do domain interactions of glycosyl hydrolases from Clostridium thermocellum contribute to protein thermostability? Protein Eng 14:167–172
Lehtio J, Sugiyama J, Gustavsson M, Fransson L, Linder M, Teeri TT (2003) The binding specificity and affinity determinants of family 1 and family 3 cellulose binding modules. Proc Natl Acad Sci USA 100:484–489
Lemos MA, Teixeira JA, Domingues MRM, Mota M, Gama FM (2003) The enhancement of the cellulolytic activity of cellobiohydrolase I and endoglucanase by the addition of cellulose binding domains derived from Trichoderma reesei. Enzyme Microb Technol 32:35–40
Lönn A, Gardonyi M, van Zyl W, Hahn Hagerdahl B, Cordero Otero R (2002) Cold adaptation of xylose isomerase from Thermus thermophilus through random PCR mutagenesis. Gene cloning and protein characterization. Eur J Biochem 269:157–163
Lymar ES, Li B, Renganathan V (1995) Purification and characterization of a cellulose-binding β-glucosidase from cellulose-degrading cultures of Phanerochaete chrysosporium. Appl Environ Microbiol 61:2976–2980
Machida M, Ohtsuki I, Fukui S, Yamashita I (1988) Nucleotide sequences of Saccharomycopsis fibuligera genes for extracellular β-glucosidases as expressed in Saccharomyces cerevisiae. Appl Environ Microbiol 54:3147–3155
Mangala SL, Kittur FS, Nishimoto M, Sakka K, Ohmiya K, Kitaoka M, Hayashi K (2003) Fusion of family VI cellulose binding domains to Bacillus halodurans xylanase increases its catalytic activity and substrate-binding capacity to insoluble xylan. J Mol Catal B Enzym 21:221–230
Nidetzky B, Steiner W, Hayn M, Claeyssens M (1994) Cellulose hydrolysis by the cellulolytic enzymes from Trichoderma reesei: a new model for synergistic interaction. Biochem J 298:705–710
Reinikainen T, Ruohonen L, Nevanen T, Laaksonen L, Kraulis P, Jones TA, Knowles JK, Teeri TT (1992) Investigation of the function of mutated cellulose-binding domains of Trichoderma reesei cellobiohydrolase I. Proteins 14:475–482
Stahlberg J, Johansson G, Pettersson G (1991) A new model for enzymatic hydrolysis of cellulose based on the two-domain structure of cellobiohydrolase I. Biotechnology 9:286–290
Suzuki K, Yabe T, Maruyama Y, Abe K, Nakajima T (2001) Characterization of recombinant yeast exo-beta-1,3-glucanase (Exg1p) expressed in Escherichia coli cells. Biosci Biotechnol Biochem 65:1310–1314
Teeri TT, Reinikainen T, Ruohonen L, Jones TA, Knowles JKC (1992) Domain function in Trichoderma reesei cellobiohydrolase. J Biotechnol 24:169–176
Tomme P, Warren RAJ, Miller RC Jr, Kilburn DG, Gilkers NR (1995) Cellulose-binding domains: classification and properties. In: Saddler JN, Pennaer MH (eds) Enzymatic degradation of insoluble carbohydrates. ACS Symposium Series No. 618. American Chemical Society, Washington, DC, pp 142–163
Tormo J, Lamed R, Chirino AJ, Morag E, Bayer EA, Shoham Y, Steitz TA (1996) Crystal structure of a bacterial family-III cellulose-binding domain: a general mechanism for attachment to cellulose. EMBO J 15:5739–5751
Towbin H, Staehelin T, Gordon J (1979) Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc Natl Acad Sci USA 76:4350–4354
Van Rensburg P, van Zyl WH, Pretorius IS (1996) Co-expression of a Phanerochaete chrysosporium cellobiohydrolase gene and a Butyrivibrio fibrisolvens endo-β-1,4-glucanase gene in Saccharomyces cerevisiae. Curr Genet 30:246–250
Acknowledgments
This project was financially supported by South Africa’’ grapegrowers and winemakers through their investment body Winetech, and by South Africa’s National Research Foundation. The authors would like to thank Dr. P. Van Rensburg, IWBT, for kindly providing the plasmid pEFB19; Mr. J. Minaar of the analytical facility (Institute for Wine Biotechnology, Stellenbosch University) for the HPLC analyses; and Ms. M. Waldron of the Electron Microscope Facility at the University of Cape Town, South Africa, for SEM analyses.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gundllapalli, S.B., Pretorius, I.S. & Cordero Otero, R.R. Effect of the cellulose-binding domain on the catalytic activity of a β-glucosidase from Saccharomycopsis fibuligera . J Ind Microbiol Biotechnol 34, 413–421 (2007). https://doi.org/10.1007/s10295-007-0213-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-007-0213-9