Abstract
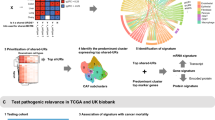
The reprogramming of cellular metabolism is a hallmark of tumorigenesis. However, the prognostic value of metabolism-related genes in colon cancer remains unclear. This study aimed to identify a metabolic gene signature to categorize colon cancer patients into high- and low-risk groups and predict prognosis. Samples from the Gene Expression Omnibus database were used as the training cohort, while samples from The Cancer Genome Atlas database were used as the validation cohort. A metabolic gene signature was established to investigate a robust risk stratification for colon cancer. Subsequently, a prognostic nomogram was established combining the metabolism-related risk score and clinicopathological characteristics of patients. A total of 351 differentially expressed metabolism-related genes were identified in colon cancer. After univariate analysis and least absolute shrinkage and selection operator-penalized regression analysis, an eight-gene metabolic signature (MTR, NANS, HADH, IMPA2, AGPAT1, GGT5, CYP2J2, and ASL) was identified to classify patients into high- and low-risk groups. High-risk patients had significantly shorter overall survival than low-risk patients in both the training and validation cohorts. A high-risk score was positively correlated with proximal colon cancer (P = 0.012), BRAF mutation (P = 0.049), and advanced stage (P = 0.027). We established a prognostic nomogram based on metabolism-related gene risk score and clinicopathologic factors. The areas under the curve and calibration curves indicated that the established nomogram showed a good accuracy of prediction. We have established a novel metabolic gene signature that could predict overall survival in colon cancer patients and serve as a biomarker for colon cancer.







Similar content being viewed by others
Availability of data and material
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. https://doi.org/10.3322/caac.21492.
Edwards BK, Ward E, Kohler BA, et al. Annual report to the nation on the status of cancer, 1975-2006, featuring colorectal cancer trends and impact of interventions (risk factors, screening, and treatment) to reduce future rates. Cancer. 2010;116(3):544–73. https://doi.org/10.1002/cncr.24760.
Ullmann P, Nurmik M, Begaj R, Haan S, Letellier E. Hypoxia- and MicroRNA-Induced Metabolic Reprogramming of Tumor-Initiating Cells. Cells. 2019. https://doi.org/10.3390/cells8060528.
O’Connell JB, Maggard MA, Ko CY. Colon cancer survival rates with the new American Joint Committee on Cancer sixth edition staging. J Natl Cancer Inst. 2004;96(19):1420–5. https://doi.org/10.1093/jnci/djh275.
Warburg O. On the origin of cancer cells. Science. 1956;123(3191):309–14. https://doi.org/10.1126/science.123.3191.309.
Hirschhaeuser F, Sattler UG, Mueller-Klieser W. Lactate: a metabolic key player in cancer. Cancer Res. 2011;71(22):6921–5. https://doi.org/10.1158/0008-5472.Can-11-1457.
Gao P, Tchernyshyov I, Chang TC, et al. c-Myc suppression of miR-23a/b enhances mitochondrial glutaminase expression and glutamine metabolism. Nature. 2009;458(7239):762–5. https://doi.org/10.1038/nature07823.
Ward PS, Thompson CB. Metabolic reprogramming: a cancer hallmark even warburg did not anticipate. Cancer Cell. 2012;21(3):297–308. https://doi.org/10.1016/j.ccr.2012.02.014.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646–74. https://doi.org/10.1016/j.cell.2011.02.013.
Bustamante E, Morris HP, Pedersen PL. Energy metabolism of tumor cells. Requirement for a form of hexokinase with a propensity for mitochondrial binding. J Biol Chem. 1981;256(16):8699–704.
Li S, Xie L, Du M, et al. Association study of genetic variants in estrogen metabolic pathway genes and colorectal cancer risk and survival. Arch Toxicol. 2018;92(6):1991–9. https://doi.org/10.1007/s00204-018-2195-y.
Ose J, Botma A, Balavarca Y, et al. Pathway analysis of genetic variants in folate-mediated one-carbon metabolism-related genes and survival in a prospectively followed cohort of colorectal cancer patients. Cancer Med. 2018. https://doi.org/10.1002/cam4.1407.
Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16(5):284–7. https://doi.org/10.1089/omi.2011.0118.
Wolbers M, Koller MT, Witteman JC, Steyerberg EW. Prognostic models with competing risks: methods and application to coronary risk prediction. Epidemiology. 2009;20(4):555–61. https://doi.org/10.1097/EDE.0b013e3181a39056.
Dekker E, Tanis PJ, Vleugels JLA, Kasi PM, Wallace MB. Colorectal cancer. Lancet (London, England). 2019;394(10207):1467–80. https://doi.org/10.1016/S0140-6736(19)32319-0.
Wolf AMD, Fontham ETH, Church TR, et al. Colorectal cancer screening for average-risk adults: 2018 guideline update from the American Cancer Society. CA Cancer J Clin. 2018;68(4):250–81. https://doi.org/10.3322/caac.21457.
Kasi PM, Shahjehan F, Cochuyt JJ, Li Z, Colibaseanu DT, Merchea A. Rising proportion of young individuals with rectal and colon cancer. Clin Colorectal Cancer. 2019;18(1):e87–95. https://doi.org/10.1016/j.clcc.2018.10.002.
Lasabová Z, Kalman M, Holubeková V, et al. Mutation analysis of POLE gene in patients with early-onset colorectal cancer revealed a rare silent variant within the endonuclease domain with potential effect on splicing. Clin Exp Med. 2019;19(3):393–400. https://doi.org/10.1007/s10238-019-00558-7.
Vatandoust S, Price TJ, Karapetis CS. Colorectal cancer: metastases to a single organ. World J Gastroenterol. 2015;21(41):11767–76. https://doi.org/10.3748/wjg.v21.i41.11767.
Singh B, Mitragotri S. Harnessing cells to deliver nanoparticle drugs to treat cancer. Biotechnol Adv. 2019. https://doi.org/10.1016/j.biotechadv.2019.01.006.
Zhang Y, Yuan J, Zhang HY, et al. Natural resistance to apoptosis correlates with resistance to chemotherapy in colorectal cancer cells. Clin Exp Med. 2012. https://doi.org/10.1007/s10238-011-0146-5.
Fouad YA, Aanei C. Revisiting the hallmarks of cancer. Am J Cancer Res. 2017;7(5):1016–36.
Zhang C, Aldrees M, Arif M, Li X, Mardinoglu A, Aziz MA. Elucidating the reprograming of colorectal cancer metabolism using genome-scale metabolic modeling. Front Oncol. 2019;9:681. https://doi.org/10.3389/fonc.2019.00681.
Liu GM, Xie WX, Zhang CY, Xu JW. Identification of a four-gene metabolic signature predicting overall survival for hepatocellular carcinoma. J Cell Physiol. 2019. https://doi.org/10.1002/jcp.29081.
Ma B, Jiang H, Wen D, et al. Transcriptome analyses identify a metabolic gene signature indicative of dedifferentiation of papillary thyroid cancer. J Clin Endocrinol Metab. 2019;104(9):3713–25. https://doi.org/10.1210/jc.2018-02686.
Motamedian E, Taheri E, Bagheri F. Proliferation inhibition of cisplatin-resistant ovarian cancer cells using drugs screened by integrating a metabolic model and transcriptomic data. Cell Prolif. 2017. https://doi.org/10.1111/cpr.12370.
Kuroda K, Fukuda T, Isogai H, Okumura K, Krstic-Demonacos M, Isogai E. Antimicrobial peptide FF/CAP18 induces apoptotic cell death in HCT116 colon cancer cells via changes in the metabolic profile. Int J Oncol. 2015;46(4):1516–26. https://doi.org/10.3892/ijo.2015.2887.
Bahreyni A, Samani SS, Rahmani F, et al. Role of adenosine signaling in the pathogenesis of breast cancer. J Cell Physiol. 2018;233(3):1836–43. https://doi.org/10.1002/jcp.25944.
Goswami MT, Chen G, Chakravarthi BV, et al. Role and regulation of coordinately expressed de novo purine biosynthetic enzymes PPAT and PAICS in lung cancer. Oncotarget. 2015;6(27):23445–61. https://doi.org/10.18632/oncotarget.4352.
Barfeld SJ, Fazli L, Persson M, et al. Myc-dependent purine biosynthesis affects nucleolar stress and therapy response in prostate cancer. Oncotarget. 2015;6(14):12587–602. https://doi.org/10.18632/oncotarget.3494.
Wang X, Yang K, Xie Q, et al. Purine synthesis promotes maintenance of brain tumor initiating cells in glioma. Nat Neurosci. 2017;20(5):661–73. https://doi.org/10.1038/nn.4537.
Yin J, Ren W, Huang X, Deng J, Li T, Yin Y. Potential Mechanisms Connecting Purine Metabolism and Cancer Therapy. Front Immunol. 2018;9:1697. https://doi.org/10.3389/fimmu.2018.01697.
Eroglu A, Canbolat O, Demirci S, Kocaoglu H, Eryavuz Y, Akgül H. Activities of adenosine deaminase and 5′-nucleotidase in cancerous and noncancerous human colorectal tissues. Med Oncol. 2000;17(4):319–24. https://doi.org/10.1007/bf02782198.
Vannoni D, Di Pietro MC, Rosi F, et al. Metabolism of adenosine in human colorectal tumour. Nucleosides Nucleotides Nucleic Acids. 2004;23(8–9):1455–7. https://doi.org/10.1081/ncn-200027676.
Asante I, Chui D, Pei H, et al. Alterations in folate-dependent one-carbon metabolism as colon cell transition from normal to cancerous. J Nutr Biochem. 2019;69:1–9. https://doi.org/10.1016/j.jnutbio.2019.02.008.
Lopes-Ramos CM, Kuijjer ML, Ogino S, et al. Gene regulatory network analysis identifies sex-linked differences in colon cancer drug metabolism. Cancer Res. 2018;78(19):5538–47. https://doi.org/10.1158/0008-5472.Can-18-0454.
Voloshanenko O, Schwartz U, Kranz D, et al. β-catenin-independent regulation of Wnt target genes by RoR2 and ATF2/ATF4 in colon cancer cells. Sci Rep. 2018;8(1):3178. https://doi.org/10.1038/s41598-018-20641-5.
Fernández LP, Ramos-Ruiz R, Herranz J, et al. The transcriptional and mutational landscapes of lipid metabolism-related genes in colon cancer. Oncotarget. 2018;9(5):5919–30. https://doi.org/10.18632/oncotarget.23592.
Zara-Lopes T, Galbiatti-Dias ALS, Castanhole-Nunes MMU, et al. Polymorphisms in, and genes involved in folate metabolism and thyroid cancer: a case-control study. Arch Med Sci. 2019;15(2):522–30. https://doi.org/10.5114/aoms.2018.73091.
Shen C, Song YH, Xie Y, et al. Downregulation of HADH promotes gastric cancer progression via Akt signaling pathway. Oncotarget. 2017;8(44):76279–89. https://doi.org/10.18632/oncotarget.19348.
Funding
None.
Author information
Authors and Affiliations
Contributions
JR and JF designed the experiment. JR, JF, WS, and CTW undertook the data acquisition. JR, JF, WS, and YHG were involved in the interpretation of data. JR and JF analyzed and visualized the data. All authors drafted and revised the manuscript. The final manuscript was read and approved by all authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest in this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ren, J., Feng, J., Song, W. et al. Development and validation of a metabolic gene signature for predicting overall survival in patients with colon cancer. Clin Exp Med 20, 535–544 (2020). https://doi.org/10.1007/s10238-020-00652-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10238-020-00652-1