Abstract
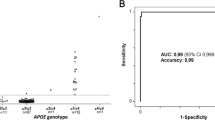
Apolipoprotein E (apo E) polymorphism is associated with increased risk of cardiovascular and Alzheimer diseases, making its genotyping of potentially predictive value. We developed a rapid, reliable and specific method for determining APOE genotypes by fluorescent resonance energy transfer (FRET) over a high number of samples in a single run using a LightTyper™ device and dedicated probes. The method, validated with 75 blood samples, was designed to simultaneously detect three common APOE polymorphisms, ε2, ε3 and ε4, and to identify in a single reaction any of the six following genotypes: ε2/ε2, ε3/ε3, ε4/ε4, ε3/ε4, ε4/ε2, ε3/ε2. The assay involved three phases: (1) DNA extraction, (2) amplification, and (3) melting curve analysis using FRET technique. Briefly, genomic DNA of patients was extracted from total blood. Fragment of APOE was amplified by a first PCR run. Fluorescent labeled probes were added in a second PCR run. FRET genotyping showed following distribution: (1) 1.3% for ε2/ε2 and ε4/ε4 homozygotes, (2) 4.0, 6.6 and 14.7% for ε2/ε4, ε2/ε3 and ε3/ε4 heterozygotes, respectively, and (3) 72.0% for ε3/ε3 homozygotes. Moreover, a careful analysis of the FRET melting curves allowed us to determine the presence of a new polymorphism on the third position of the codon 158 (-AAGCGT-), namely, two nucleotides downstream from the known polymorphism. When the FRET analysis was compared to those obtained by RFLP and sequencing, the presence of this new polymorphism was confirmed only by sequencing thus indicating that RFLP analysis is not always reliable for genotyping.




Similar content being viewed by others
References
Abdollahi MR, Guthrie PA, Smith GD, Lawlor DA, Ebrahim S, Day IN (2006) Integrated single-label liquid-phase assay of APOE codons 112 and 158 and a lipoprotein study in British women. Clin Chem 52(7):1420–1423
Aslanidis C, Schmitz G (1999) High-speed apolipoprotein E genotyping and apolipoprotein B3500 mutation detection using real-time fluorescence PCR and melting curves. Clin Chem 45(7):1094–1097
Ballerini S, Bellincampi L, Bernardini S, Casciani S, Motti C, Cortese C, Federici G (2002) Apolipoprotein E genotyping: a comparative study between restriction endonuclease mapping and allelic discrimination with the LightCycler. Clin Chim Acta 317(1–2):71–76
Bennet AM, Di Angelantonio E, Ye Z, Wensley F, Dahlin A, Ahlbom A, Keavney B, Collins R, Wiman B, de Faire U, Danesh J (2007) Association of apolipoprotein E genotypes with lipid levels and coronary risk. JAMA 298(11):1300–1311
Bennett CD, Campbell MN, Cook CJ, Eyre DJ, Nay LM, Nielsen DR, Rasmussen RP, Bernard PS (2003) The LightTyper: high-throughput genotyping using fluorescent melting curve analysis. Biotechniques 34(6):1288–1292, 1294–1295
Francés F, Corella D, Sorlí JV, Guillén M, González JI, Portolés O (2005) Validating a rapid method for detecting common polymorphisms in the APOA5 gene by melting curve analysis using LightTyper. Clin Chem 51(7):1279–1282
Hixson JE, Vernier DT (1990) Restriction isotyping of human apolipoprotein E by gene amplification and cleavage with HhaI. J Lipid Res 31(3):545–548
Lin Z, Suzow JG, Fontaine JM, Naylor EW (2004) A high throughput beta-globin genotyping method by multiplexed melting temperature analysis. Mol Genet Metab 81(3):237–243
Mahley RW, Weisgraber KH, Huang Y (2006) Apolipoprotein E4: a causative factor and therapeutic target in neuropathology, including Alzheimer’s disease. Proc Natl Acad Sci USA 103(15):5644–5651
Mahley RW, Rall SC Jr (2000) Apolipoprotein E: far more than a transport lipid protein. Annu Rev Genomics Hum Genet 1:507–537
Mahley RW, Huang Y (1999) Apolipoprotein E: from atherosclerosis to Alzheimer’s disease and beyond. Curr Opin Lipidol 10(3):207–217
Mahley RW, Huang Y, Rall SC Jr (1999) Pathogenesis of type III hyperlipoproteinemia (dysbetalipoproteinemia). Questions, quandaries, and paradoxes. J Lipid Res 40(11):1933–1949
Mahley RW (1988) Apolipoprotein E: cholesterol transport protein with expanding role in cell biology. Science 240(4852):622–630 Review
Murugesan G, Kottke-Marchant K, Ellis S, Agah R, Tubbs R (2005) LightTyper platform for high-throughput clinical genotyping. Expert Rev Mol Diagn 5(3):457–471
Nauck M, Hoffmann MM, Wieland H, März W (2000) Evaluation of the apo E genotyping kit on the LightCycler. Clin Chem 46(5):722–724
Olaisen B, Teisberg P, Gedde-Dahl T Jr (1982) The locus for apolipoprotein E (apoE) is linked to the complement component C3 (C3) locus on chromosome 19 in man. Hum Genet 62(3):233–236
Acknowledgments
We would like to thank S. Visvikis-Siest, M. Pfister and Roche Diagnostics (Meylan, France) for technical support.
Conflict of interest statement
The authors declare that they have no conflict of interest related to the publication of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
An erratum to this article can be found at http://dx.doi.org/10.1007/s10238-009-0045-1
Rights and permissions
About this article
Cite this article
Rihn, B.H., Berahmoune, S., Jouma, M. et al. APOE genotyping: comparison of three methods. Clin Exp Med 9, 61–65 (2009). https://doi.org/10.1007/s10238-008-0012-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10238-008-0012-2