Abstract
Purpose
Here, we investigated expression modules reflecting the reciprocal expression of the cancer microenvironment and immune response-related genes associated with poor prognosis in primary central nervous system lymphoma (PCNSL).
Methods
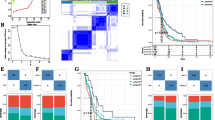
Weighted gene coexpression network analysis revealed representative modules, including neurogenesis, immune response, anti-virus, microenvironment, gene expression and translation, extracellular matrix, morphogenesis, and cell adhesion in the transcriptome data of 31 PCNSL samples.
Results
Gene expression networks were also reflected by protein–protein interaction networks. In particular, some of the hub genes were highly expressed in patients with PCNSL with prognoses as follows: AQP4, SLC1A3, GFAP, CXCL9, CXCL10, GBP2, IFI6, OAS2, IFIT3, DCN, LRP1, and LUM with good prognosis; and STAT1, IFITM3, GZMB, ISG15, LY6E, TGFB1, PLAUR, MMP4, FTH1, PLAU, CSF3R, FGR, POSTN, CCR7, TAS1R3, small ribosomal subunit genes, and collagen type 1/3/4/6 genes with poor prognosis. Furthermore, prognosis prediction formulae were constructed using the Cox proportional-hazards regression model, which demonstrated that the IP-10 receptor gene CXCR3 and type I interferon-induced protein gene IFI44L could predict patient survival in PCNSL.
Conclusion
These results indicate that the differential expression and balance of immune response and microenvironment genes may be required for PCNSL tumor growth or prognosis prediction, which would help understanding the mechanism of tumorigenesis and potential therapeutic targets in PCNSL.






Similar content being viewed by others
Availability of data and materials
Data and materials for the study are included in the manuscript and supplementary information.
Abbreviations
- ABC:
-
Activated B cell
- AUC:
-
Area under the curve
- BBB:
-
Blood–brain barrier
- CI:
-
Confidential interval
- CNS:
-
Central nervous system
- COL:
-
Collagen
- DEGs:
-
Differentially expressed genes
- DLBCL:
-
Diffuse large B cell lymphoma
- EBV:
-
Epstein–Barr virus
- ECM:
-
Extracellular matrix
- FDR:
-
False discovery rate
- FL:
-
Follicular lymphoma
- FPKM:
-
Fragments per kilobase of exon per million reads mapped
- GO:
-
Gene ontology
- GSEA:
-
Gene set enrichment analysis
- HD-MTX:
-
High-dose methotrexate
- HR:
-
Hazard ratio
- IELSG:
-
International Extranodal Lymphoma Study Group
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- KPS:
-
Karnofsky Performance Status
- LDH:
-
Lactate dehydrogenase
- MAD:
-
Mean absolute deviation
- MCODE:
-
Molecular Complex Detection
- MSKCC:
-
Memorial Sloan Kettering Cancer Center
- NAFLD:
-
Non-alcoholic fatty liver disease
- NGS:
-
Next generation sequencing
- NHL:
-
Non-Hodgkin’s lymphoma
- non-GCB:
-
Non-germinal center B cell-like
- OS:
-
Overall survival
- PCNSL:
-
Primary central nervous system lymphoma
- PKA:
-
Protein kinase A
- PPI:
-
Protein–protein interaction
- PRPS:
-
Ribosomal subunits
- ROC:
-
Receiver operating characteristic
- SNV/indel:
-
Single nucleotide variant/insertion/deletion
- STRING:
-
Search Tool of the Retrieval of Interacting Genes Database
- WCNA:
-
Weighted correlation network analysis
- WGCNA:
-
Weighted gene coexpression network analysis
- WHO:
-
World Health Organization
References
Louis DN, Perry A, Reifenberger G et al (2016) The 2016 world health organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131:803–820
Fox CP, Phillips EH, Smith J et al (2019) Guidelines for the diagnosis and management of primary central nervous system diffuse large B-cell lymphoma. Br J Haematol 184:348–363
Hans CP, Weisenburger DD, Greiner TC et al (2004) Confirmation of the molecular classification of diffuse large B-cell lymphoma by immunohistochemistry using a tissue microarray. Blood 103:275–282
Courts C, Montesinos-Rongen M, Brunn A et al (2008) Recurrent inactivation of the PRDM1 gene in primary central nervous system lymphoma. J Neuropathol Exp Neurol 67:720–727
Raoux D, Duband S, Forest F et al (2010) Primary central nervous system lymphoma: immunohistochemical profile and prognostic significance. Neuropathology 30:232–240
Chukwueke UN, Nayak L (2019) Central nervous system lymphoma. Hematol Oncol Clin North Am 33:597–611
Montesinos-Rongen M, Siebert R, Deckert M (2009) Primary lymphoma of the central nervous system: just DLBCL or not? Blood 113:7–10
Grommes C, DeAngelis LM (2017) Primary CNS lymphoma. J Clin Oncol 35:2410–2418
Prodduturi P, Bierman PJ (2012) Current and emerging pharmacotherapies for primary CNS lymphoma. Clin Med Insights Oncol 6:219–231
Mead GM, Bleehen NM, Gregor A et al (2000) A medical research council randomized trial in patients with primary cerebral non-Hodgkin lymphoma: cerebral radiotherapy with and without cyclophosphamide, doxorubicin, vincristine, and prednisone chemotherapy. Cancer 89:1359–1370
Gavrilovic IT, Hormigo A, Yahalom J et al (2006) Long-term follow-up of high-dose methotrexate-based therapy with and without whole brain irradiation for newly diagnosed primary CNS lymphoma. J Clin Oncol 24:4570–4574
Thiel E, Korfel A, Martus P et al (2010) High-dose methotrexate with or without whole brain radiotherapy for primary CNS lymphoma (G-PCNSL-SG-1): a phase 3, randomised, non-inferiority trial. Lancet Oncol 11:1036–1047
Langner-Lemercier S, Houillier C, Soussain C et al (2016) Primary CNS lymphoma at first relapse/progression: characteristics, management, and outcome of 256 patients from the French LOC network. Neuro Oncol 18:1297–1303
Nayak L, Hedvat C, Rosenblum MK et al (2011) Late relapse in primary central nervous system lymphoma: clonal persistence. Neuro Oncol 13:525–529
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinform 9:559
Zhang B, Horvath S (2005) A general framework for weighted gene co-expression network analysis. Stat Appl Genet Mol Biol. https://doi.org/10.2202/1544-6115.1128
Presson AP, Yoon NK, Bagryanova L et al (2011) Protein expression based multimarker analysis of breast cancer samples. BMC Cancer 11:230
Langfelder P, Mischel PS, Horvath S (2013) When is hub gene selection better than standard meta-analysis? PLoS ONE 8:e61505
Luo Y, Coskun V, Liang A et al (2015) Single-cell transcriptome analyses reveal signals to activate dormant neural stem cells. Cell 161:1175–1186
Bruno A, Boisselier B, Labreche K et al (2014) Mutational analysis of primary central nervous system lymphoma. Oncotarget 5:5065–5075
Braggio E, Van Wier S, Ojha J et al (2015) Genome-wide analysis uncovers novel recurrent alterations in primary central nervous system lymphomas. Clin Cancer Res 21:3986–3994
Vater I, Montesinos-Rongen M, Schlesner M et al (2015) The mutational pattern of primary lymphoma of the central nervous system determined by whole-exome sequencing. Leukemia 29:677–685
Zhou Y, Liu W, Xu Z et al (2018) Analysis of genomic alteration in primary central nervous system lymphoma and the expression of some related genes. Neoplasia 20:1059–1069
Takashima Y, Sasaki Y, Hayano A et al (2018) Target amplicon exome-sequencing identifies promising diagnosis and prognostic markers involved in RTK-RAS and PI3K-AKT signaling as central oncopathways in primary central nervous system lymphoma. Oncotarget 9:27471–27486
Kaulen LD, Erson-Omay EZ, Henegariu O et al (2021) Exome sequencing identifies SLIT2 variants in primary CNS lymphoma. Br J Haematol 193:375–379
Gandhi MK, Hoang T, Law SC et al (2021) EBV-associated primary CNS lymphoma occurring after immunosuppression is a distinct immunobiological entity. Blood 137:1468–1477
Takashima Y, Kawaguchi A, Sato R et al (2019) Differential expression of individual transcript variants of PD-1 and PD-L2 genes on Th-1/Th-2 status is guaranteed for prognosis prediction in PCNSL. Sci Rep 9:10004
Chen Z, Shen Z, Zhang Z et al (2021) RNA-associated co-expression network identifies novel biomarkers for digestive system cancer. Front Genet 12:659788
Subramanian A, Tamayo P, Mootha VK et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A 102:15545–15550
Yu G, He QY (2016) ReactomePA: an R/Bioconductor package for reactome pathway analysis and visualization. Mol BioSyst 12:477–479
Takashima Y, Hamano M, Fukai J et al (2020) GSEA-assisted gene signatures valid for combinations of prognostic markers in PCNSL. Sci Rep 10:8435
Yu G, Wang LG, Yan GR et al (2015) DOSE: an R/Bioconductor package for disease ontology semantic and enrichment analysis. Bioinformatics 31:608–609
Szklarczyk D, Gable AL, Lyon D et al (2019) STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 47:D607–D613
Bader GD, Hogue CW (2003) An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinform 4:2
Tashiro K, Tada H, Heilker R et al (1993) Signal sequence trap: a cloning strategy for secreted proteins and type I membrane proteins. Science 261:600–603
Venetz D, Ponzoni M, Schiraldi M et al (2010) Perivascular expression of CXCL9 and CXCL12 in primary central nervous system lymphoma: T-cell infiltration and positioning of malignant B cells. Int J Cancer 127:2300–2312
Gao P, Grigoryev DN, Rafaels NM et al (2010) CD14, a key candidate gene associated with a specific immune response to cockroach. Clin Exp Allergy 40:1353–1364
Luo S, Wu R, Li Q et al (2022) Epigenetic regulation of IFI44L expression in monocytes affects the functions of monocyte-derived dendritic cells in systemic lupus erythematosus. J Immunol Res 2022:4053038
Lei J, Yin X, Shang H et al (2019) IP-10 is highly involved in HIV infection. Cytokine 115:97–103
Ozturk T, Kollhoff A, Anderson AM et al (2019) Linked CSF reduction of phosphorylated tau and IL-8 in HIV associated neurocognitive disorder. Sci Rep 9:8733
Fang M, Xu N, Shao X et al (2012) Inhibitory effects of human immunodeficiency virus gp120 and Tat on CpG-A-induced inflammatory cytokines in plasmacytoid dendritic cells. Acta Biochim Biophys Sin (Shanghai) 44:797–804
Fletcher JS, Wu J, Jessen WJ et al (2019) Cxcr3-expressing leukocytes are necessary for neurofibroma formation in mice. JCI Insight 4:e98601
Nitcheu J, Bonduelle O, Combadiere C et al (2003) Perforin-dependent brain-infiltrating cytotoxic CD8+ T lymphocytes mediate experimental cerebral malaria pathogenesis. J Immunol 170:2221–2228
Campanella GS, Tager AM, El Khoury JK et al (2008) Chemokine receptor CXCR3 and its ligands CXCL9 and CXCL10 are required for the development of murine cerebral malaria. Proc Natl Acad Sci U S A 105:4814–4819
Suidan GL, Mcdole JR, Chen Y et al (2008) Induction of blood brain barrier tight junction protein alterations by CD8 T cells. PLoS ONE 3:e3037
Martín-Fontecha A, Thomsen LL, Brett S et al (2004) Induced recruitment of NK cells to lymph nodes provides IFN-gamma for T(H)1 priming. Nat Immunol 5:1260–1265
Hansen DS, Bernard NJ, Nie CQ et al (2007) NK cells stimulate recruitment of CXCR3+ T cells to the brain during Plasmodium berghei-mediated cerebral malaria. J Immunol 178:5779–5788
Jinquan T, Jing C, Jacobi HH et al (2000) CXCR3 expression and activation of eosinophils: role of IFN-gamma-inducible protein-10 and monokine induced by IFN-gamma. J Immunol 165:1548–1556
Shen Q, Zhang R, Bhat NR (2006) MAP kinase regulation of IP10/CXCL10 chemokine gene expression in microglial cells. Brain Res 1086:9–16
Bonacchi A, Romagnani P, Romanelli RG et al (2001) Signal transduction by the chemokine receptor CXCR3: activation of Ras/ERK, Src, and phosphatidylinositol 3-kinase/Akt controls cell migration and proliferation in human vascular pericytes. J Biol Chem 276:9945–9954
Acknowledgements
This study was supported in part by the MEXT KAKENHI Grant Number 21H03045 to RY. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Funding
This study was supported in part by the MEXT KAKENHI Grant Number 21H03045 to RY. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
YT and RY designed the experiments. JF, YI, KK and HH diagnosed and treated patients and collected samples. YT, MH, KY, AH and RY performed the experiments. YT, MH, KY, AH and RY analyzed data. YT and RY wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethical approval
All experiments comply with the Ethics Committee of Kyoto Prefectural University of Medicine (RBMR-G-146).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
About this article
Cite this article
Takashima, Y., Hamano, M., Yoshii, K. et al. Reciprocal expression of the immune response genes CXCR3 and IFI44L as module hubs are associated with patient survivals in primary central nervous system lymphoma. Int J Clin Oncol 28, 468–481 (2023). https://doi.org/10.1007/s10147-022-02285-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10147-022-02285-8