Abstract
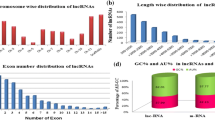
Non-coding RNAs with lengths greater than 200 bp are known as long non-coding RNAs (lncRNAs), and these RNAs play important role in gene regulation and plant development. However, to date, little is known regarding the role played by lncRNAs during flowering in hickory (Carya cathayensis). Here, we performed whole transcriptome RNA-sequencing of samples from hickory female and male floral buds, in which three samples (H0311PF, H0318PF, and H0402PF) represent pre-flowering, flowering, and post-flowering, respectively, while eight male samples collected from May 8th to June 13th as this time course are the key stage for male floral bud differentiation. We identified 2163 lncRNAs in hickory during flowering, containing 213 intronic, 1488 intergenic, and 462 antisense lncRNAs. We noticed that 510 and 648 lncRNAs were differentially expressed corresponding to female and male floral buds, respectively. And some of the lncRNAs were in a tightly tissue-specific or stage-specific manner. To further understand the roles of the lncRNAs, we predicted the function of the lncRNAs in cis- and trans-acting modes. The results showed that 924 lncRNAs were cis-correlated with 1536 protein-coding genes, while 1207 lncRNAs co-expressed (trans-acting) with 7432 protein-coding genes (R > 0.95 or R < − 0.95). These lncRNAs were all enriched in flower development-associated biological processes, i.e., circadian rhythm, vernalization response, response to gibberellin, inflorescence development, floral organ development, etc. To further understand the relationships between lncRNAs and floral-core genes, we build a co-expressing lncRNA-mRNA flowering network. We classified these floral genes into different pathway (photoperiod, vernalization, gibberellin, autonomous, and sucrose pathway) according to their particular functions. We found a set of lncRNAs that preferentially expressed in these pathways. The network showed that some lncRNAs (i.e., XLOC_038669, XLOC_017938) functioned in a particular flowering time pathway, while others (i.e., XLOC_011251, XLOC_04110) were involved in multiple pathway. Furthermore, some lncRNAs (i.e., XLOC_038669, XLOC_009597, and XLOC_049539) played roles in single or multiple pathways via interaction with each other. This study provides a genome-wide survey of hickory flower-related lncRNAs and will contribute to further understanding of the molecular mechanism underpinning flowering in hickory.










Similar content being viewed by others
References
Achard P, Gong F, Cheminant S, Alioua M, Hedden P, Genschik P (2008) The cold-inducible CBF1 factor-dependent signaling pathway modulates the accumulation of the growth-repressing DELLA proteins via its effect on gibberellin metabolism. Plant Cell 20:2117–2129
Amasino RM, Michaels SD (2010) The timing of flowering. Plant Physiol 154:516–520
An YQ, McDowell JM, Huang S, McKinney EC, Chambliss S, Meagher RB (1996) Strong, constitutive expression of the Arabidopsis ACT2/ACT8 actin subclass in vegetative tissues. Plant J 10:107–121
Ariel F, Jegu T, Latrasse D, Romero-Barrios N, Christ A, Benhamed M, Crespi M (2014) Noncoding transcription by alternative RNA polymerases dynamically regulates an auxin-driven chromatin loop. Mol Cell 55:383–396
Arrial RT, Togawa RC, Brigido MM (2009) Screening non-coding RNAs in transcriptomes from neglected species using PORTRAIT: case study of the pathogenic fungus Paracoccidioides brasiliensis. BMC Bioinformatics 10:239
Arrom L, Munné-Bosch S (2012) Sucrose accelerates flower opening and delays senescence through a hormonal effect in cut lily flowers. Plant Sci 188-189:41–47
Bardou F, Ariel F, Simpson CG, Reomero-Barrios N, Laporte P, Balzergue S, Brown JW, Crespi M (2014) Long noncoding RNA modulates alternative splicing regulators in Arabidopsis. Dev Cell 30:166–176
Becker A, Theissen G (2003) The major clades of MADS-box genes and their role in the development and evolution of flowering plants. Mol Phylogenet Evol 29:464–489
Berry S, Dean C (2015) Environmental perception and epigenetic memory: mechanistic insight through FLC. Plant J 83:133–148
Caldelari D, Wang G, Framer EE, Dong X (2011) Arabidopsis lox3 lox4 double mutants are male sterile and defective in global proliferative arrest. Plant Mol Biol 75:25–33
Chekanova JA, Gregory BD, Reverdatto SV, Chen H, Kumar R, Hooker T, Yazaki J, Li P, Skiba N, Peng Q, Alonso J, Brukhin V, Grossniklaus U, Ecker JR, Belostotsky DA (2007) Genome-wide high-resolution mapping of exosome substrates reveals hidden features in the Arabidopsis transcriptome. Cell 131:1340–1353
Chen X, Zhang Z, Liu D, Zhang K, Li A, Mao L (2010) SQUAMOSA promoter-binding protein-like transcription factors: star players for plant growth and development. J Int Plant Biol 52:946–951
Chung KS, Yoo SY, Yoo SJ, Lee JS, Ahn JH (2010) BROTHER OF FT AND TFL1 (BFT), a member of the FT/TFL1 family, shows distinct pattern of expression during the vegetative growth of Arabidopsis. Plant Signal Behav 5:1102–1104
De LF, Crevillen P, Jones AM, Greb T, Dean C (2008) A PHD-polycomb repressive complex 2 triggers the epigenetic silencing of FLC during vernalization. Proc Natl Acad Sci U S A 105:16831–16836
DeYoung BJ, Bickle KL, Schrage KJ, Muskett P, Patel K, Clark SE (2006) The CLAVATA1-related BAM1, BAM2 and BAM3 receptor kinase-like proteins are required for meristem function in Arabidopsis. Plant J 45:1–16
Di C, Yuan J, Wu Y, Li J, Lin H, Hu L, Zhang T, Qi Y, Gerstein MB, Guo Y, Lu ZJ (2014) Characterization of stress-responsive lncRNAs in Arabidopsis thaliana by integrating expression, epigenetic and structural features. Plant J 80:848–861
Eiβmann M, Gutschner T, Hämmerle M, Günther S, Caudron-Herger M, Groβ M, Schirmacher P, Rippe K, Braun T, Zörnig M, Diederichs S (2012) Loss of the abundant nuclear non-coding RNAMALAT1is compatible with life and development. RNA Biol 9:1076–1087
Fatica A, Bozzoni I (2014) Long non-coding RNAs: new players in cell differentiation and development. Nat Rev Genet 15:7–21
Franco-Zorrilla JM, Valli A, Todesco N, Mateos I, Puga MI, Rubio-Somoza I, Leyva A, Weigel D, Garcia JA, Paz-Ares J (2007) Target mimicry provides a new mechanism for regulation of microRNA activity. Nat Genet 39:1033–1037
Golicz AA, Bhalla PL, Singh MB (2018) lncRNAs in plant and animal sexual reproduction. Trends Plant Sci 23:195–205
Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D, Huarte M, Zuk O, Carey BW, Cassady JP, Cabili MN, Jaenisch R, Mikkelsen TS, Jacks T, Hacohen N, Bernstein BE, Kellis M, Regev A, Rinn JL, Lander ES (2009) Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458:223–227
Hatayama T, Takeno K (2003) The metabolic pathway of salicylic acid rather than of chlorogenic acid is involved in the stress-induced flowering of Pharbitis nil. J Plant Physiol 160:461–467
Henriques R, Wang H, Liu J, Boix M, Huang LF, Chua NH (2017) The antiphasic regulatory module comprising CDF5 and its antisense RNA FLORE links the circadian clock to photoperiodic flowering. New Phytol 216:854–867
Hord CL, Chen C, Deyoung BJ, Clark SE, Ma H (2006) The BAM1/BAM2 receptor-like kinases are important regulators of Arabidopsis early anther development. Plant Cell 18:1667–1680
Huang YJ, Xia GH, Wang ZJ, Zheng BS, Liang JY, Huang JQ (2007) Studies on anatomy of development of female flower in Carya cathayensis Sarg. Acta Agric Univ Jiangxiensis 29:723e726
Huang d W, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists, using DAVID bioinformatics resources. Nat Protoc 4:44–57
Huang YJ, Liu LL, Huang JQ, Wang ZJ, Chen FF, Zhang QX, Zheng BS, Chen M (2013) Use of transcriptome sequencing to understand the pistillate flowering in hickory (Carya cathayensis Sarg.). BMC Genomics 14:691
Hwang K, Susila H, Nasim Z, Jung JY, Ahn JH (2019) Arabidopsis ABF3 and ABF4 transcription factors act with the NF-YC complex to regulate SOC1 expression and mediate drought-accelerated flowering. Mol Plant 12:489–505
Jandura A, Krause HM (2017) The new RNA world: growing evidence for long noncoding RNA functionality. Trends Genet 33:665–676
Kang YJ, Yang DC, Kong L, Hou M, Meng YQ, Wei L, Gao G (2017) CPC2: a fast and accurate coding potential calculator based on sequence intrinsic features. Nucleic Acids Res 45:W12–W16
Kapranov P, Cheng J, Dike S, Nix DA, Duttagupta R, Willingham AT, Stadler PF, Hertel J, Hackermüller J, Hofacker IL, Bell I, Cheung E, Drenkow J, Dumais E, Patel S, Helt G, Ganesh M, Ghosh S, Piccolboni A, Sementchenko V, Tammana H, Gingeras TR (2007) RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science 316:1484–1488
Katayama S, Tomaru Y, Kasukawa T, Waki K, Nakanishi M, Nakamura M, Nishida H, Yap CC, Suzuki M, Kawai J, Suzuki H, Carninci P, Hayashizaki Y, Wells C, Frith M, Ravasi T, Pang KC, Hallinan J, Mattick J, Hume DA, Lipovich L, Batalov S, Engström PG, Mizuno Y, Faghihi MA, Sandelin A, Chalk AM, Mottagui-Tabar S, Liang Z, Lenhard B, Wahlestedt C, RIKEN Genome Exploration Research Group, Genome Science Group (Genome Network Project Core Group), FANTOM Consortium (2005) Antisense transcription in the mammalian transcriptome. Science 309:1564–1566
Khalil AM, Guttman M, Huarte M, Garber M, Raj A, Rivea Morales D, Thomas K, Presser A, Bernstein BE, van Oudenaarden A, Regev A, Lander ES, Rinn JL (2009) Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc Natl Acad Sci U S A 106:11667–11672
Kim DH, Sung S (2017) Vernalization-triggered intragenic chromatin loop formation by long noncoding RNAs. Dev Cell 40:302–312
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12:357–360
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R, 1000 Genome Project Data Processing Subgroup (2009) The sequence alignment/map (SAM) format and SAMtools. Bioinformatics 25:2078–2079
Li S, Yamada M, Han X, Ohler U, Benfey PN (2016) High-resolution expression map of the Arabidopsis root reveals alternative splicing and lincRNA regulation. Dev Cell 39:508–522
Li L, Eichten SR, Shimizu R, Petsch K, Yeh CT, Wu W, Chettoor AM, Givan SA, Cole RA, Fowler JE, Evans MMS, Scanlon MJ, Yu J, Schnable PS, Timmermans MCP, Springer NM, Muehlbauer GJ (2018) Genome-wide discovery and characterization of maize long non-coding RNAs. Genome Biol 15:R40
Lin SI, Wang JG, Poon SY, Su CL, Wang SS, Chiou TJ (2005) Differential regulation of FLOWERING LOCUS C expression by vernalization in cabbage and Arabidopsis. Plant Physiol 137:1037–1048
Liu C, Bai B, Skogerbo G, Gai L, Deng W, Zhang Y, Bu D, Zhao Y, Chen R (2005) NONCODE: an integrated knowledge database of non-coding RNAs. Nucleic Acids Res 33:D112–D115
Liu C, Chen H, Er HL, Soo HM, Kumar PP, Han JH, Liou YC, Yu H (2008) Direct interaction of AGL24 and SOC1 integrates flowering signals in Arabidopsis. Development 135:1481–1491
Liu B, Zuo Z, Liu H, Lin C (2011) Arabidopsis cryptochrome 1 interacts with SPA1 to suppress COP1 activity in response to blue light. Genes Dev 25:1029–1034
Liu L, Liu C, Hou X, Xi W, Shen L, Tao Z, Wang Y, Yu H (2012) FTIP1 is an essential regulator required for florigen transport. PLoS Biol 10:e1001313
Liu J, Feng L, Gu X, Deng X, Qiu Q, Li Q, Zhang Y, Wang M, Deng Y, Wang E, He Y, Bäurle I, Li J, Cao X, He Z (2019) An H3K27me3 demethylase-HSFA2 regulatory loop orchestrates transgenerational thermomemory in Arabidopsis. Cell Res 29:379–390
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-delta delta C(T)) method. Methods 25:402–408
Macknight R, Bancroft I, Page T, Lister C, Schmidt R, Love K, Westphal L, Murphy G, Sherson S, Cobbett C, Dean C (1997) FCA, a gene controlling flowering time in Arabidopsis, encodes a protein containing RNA-binding domains. Cell 89:737–745
Matsumoto N, Okada K (2001) A homeobox gene, PRESSED FLOWER, regulates lateral axis-dependent development of Arabidopsis flowers. Genes Dev 15:3355–3364
Matzke MA, Mosher RA (2014) RNA-directed DNA methylation: an epigenetic pathway of increasing complexity. Nat Rev Genet 15:394–408
Morris KV, Mattick JS (2014) The rise of regulatory RNA. Nat Rev Genet 15:423–437
Mouradov A, Cremer F, Coupland G (2002) Control of flowering time: interacting pathways as a basis for diversity. Plant Cell 14:S111–S130
Mu J, Tan H, Hong S, Liang Y, Zuo J (2013) Arabidopsis transcription factor genes NF-YA1, 5, 6, and 9 play redundant roles in male gametogenesis, embryogenesis, and seed development. Mol Plant 6:188–201
Nambirajan G, Karunanidhi K, Ganesan A, Rajendran R, Kandasamy R, Elangovan A, Thilagar S (2018) Evaluation of antidiabetic activity of bud and flower of Avaram Senna (Cassia auriculate L.) in high fat diet and streptozotocin induced diabetic rats. Biomed Pharmacother 108:1495–1506
Ohgishi M, Saji K, Okada K, Sakai T (2004) Functional analysis of each blue light receptor, Cry1, Cry2, Phot1, and Phot2, by using combinatorial multiple mutants in Arabidopsis. Proc Natl Acad Sci U S A 101:2223–2228
Ørom UA, Derrien T, Beringer M, Gumireddy K, Gardini A, Bussotti G, Lai F, Zytnicki M, Notredame C, Huang Q, Guigo R, Shiekhattar R (2010) Long noncoding RNAs with enhancer-like function in human cells. Cell 143:46–58
Parcy F (2005) Flowering: a time for integration. Int J Dev Biol 49:585–593
Paytuví GA, Hermoso PA, de Anzar Martínez LI, Sanseverino W, Aiese CR (2016) GREENC: a wiki-based database of plant lncRNAs. Nucleic Acids Res 44:D1161–D1166
Pelaz S, Ditta GS, Baumann E, Wisman E, Yanofsky MF (2000) B and C floral organ identity functions require SEPALLATA MADS-box genes. Nature 405:200–203
Peng FY, Hu Z, Yang RC (2015) Genome-wide comparative analysis of flowering-related genes in Arabidopsis, wheat, and barley. Int J Plant Genomics 2015:874361
Pertea M, Pertea GM, Antonescu CM, Chang TC, Mendell JT, Salzberg SL (2015) Stringtie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol 33:290–295
Rieu I, Ruiz-Rivero O, Fernandez-Garcia N, Griffiths J, Powers SJ, Gong F, Linhartova T, Eriksson S, Nilsson O, Thomas SG, Phillips AL, Hedden P (2008) The gibberellin biosynthetic genes AtGA20ox1 and AtGA20ox2 act, partially redundantly, to promote growth and development throughout the Arabidopsis life cycle. Plant J 53:488–504
Rinn JL, Chang HY (2012) Genome regulation by long noncoding RNAs. Annu Rev Biochem 81:145–166
Roldán M, Gómez-Mena C, Ruiz-Garcia L, Salinas J, Martínez-Zapater JM (1999) Sucrose availability on the aerial part of the plant promotes morphogenesis and flowering of Arabidopsis in the dark. Plant J 20:581–590
Sauvageau M, Goff LA, Lodato S et al (2013) Multiple knockdown mouse models reveal lincRNAs are required for life and brain development. Elife 2:e01749
Seo PJ, Ryu J, Kang SK, Park CM (2011) Modulation of sugar metabolism by an INDETERMINATE DOMAIN transcription factor contributes to photoperiodic flowering in Arabidopsis. Plant J 65:418–429
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504
Shen C, Xu Y, Huang J, Wang Z, Qiu J, Huang Y (2014) Molecular characterization and expression analysis of the critical floral genes in hickory (Carya cathayensis Sarg.). Plant Physiol Biochem 83:142–150
Shin JH, Chekanova JA (2014) Arabidopsis RRP6L1 and RRP6L2 function in FLOWERING LOCUS C silencing via regulation of antisense RNA synthesis. PLoS Genet 10:e1004612
Shinozaki M, Hirai N, Kojima Y, Koshimizu K, Takimoto A (1994) Correlation between level of phenylpropanoids in cotyledons and flowering in Pharbitis seedling under high-fluence illumination. Plant Cell Physiol 35:807–810
Silverstone AL, Chang C, Krol E, Sun TP (1997) Developmental regulation of the gibberellin biosynthetic gene GA1, in Arabidopsis thaliana. Plant J 12:9–19
Simpson GG, Dijkwel PP, Quesada V, Henderson I, Dean C (2003) FY is an RNA 3′ end-processing factor that interacts with FCA to control the floral transition. Cell 113:777–787
Swiezewski S, Liu F, Magusin A, Dean C (2009) Cold-induced silencing by long antisense transcripts of an Arabidopsis Polycomb target. Nature 462:799–802
Takeno K (2016) Stress-induced flowering: the third category of flowering response. J Exp Bot 67:4925–4934
Takeuchi H, Higashiyama T (2016) Tip-localized receptors control pollen tube growth and LURE sensing in Arabidopsis. Nature 531:245–248
Wahlestedt C (2006) Natural antisense and noncoding RNA transcripts as potential drug targets. Drug Discov Today 11:503–508
Wang ZJ, Huang JQ, Huang YJ, Li Z, Zheng BS (2012) Discovery and profiling of novel and conserved microRNAs during flower development in Carya cathayensis via deep sequencing. Planta 236:613–621
Wang Y, Fan X, Lin F, He G, Terzaghi W, Zhu D, Deng XW (2014) Arabidopsis noncoding RNA mediates control of photomorphogenesis by red light. Proc Natl Acad Sci U S A 111:10359–10364
Wang Z, Huang J, Sun Z, Zheng B (2015) Identification of microRNAs differentially expressed involved in male flower development. Funct Integr Genomics 15:225–232
Wang CY, Liu SR, Zhang XY, Ma YJ, Hu CG, Zhang JZ (2017a) Genome-wide screening and characterization of long non-coding RNAs involved in flowering development of trifoliate orange (Poncirus trifoliata L. Raf). Sci Rep 7:43226
Wang D, Qu Z, Yang L, Zhang Q, Liu ZH, Do T, Adelson DL, Wang ZY, Searle I, Zhu JK (2017b) Transposable elements (TEs) contribute to stress-related long intergenic noncoding RNAs in plants. Plant J 90:133–146
Wang Z, Zhu T, Ma W, Wang N, Qu G, Zhang S, Wang J (2018) Genome-wide analysis of long non-coding RNAs in Catalpa bungei and their potential function in floral transition using high-throughput sequencing. BMC Genet 19:86
Wen CK, Chang C (2002) Arabidopsis RGL1 encodes a negative regulator of gibberellin responses. Plant Cell 14:87–100
Winkelshirley B (2001) Flavonoid biosynthesis. A colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiol 126:485–493
Wu X, Shi T, Iqbal S, Zhang Y, Liu L, Gao Z (2019) Genome-wide discovery and characterization of flower development related long non-coding RNAs in Prunus mume. BMC Plant Biol 19:64
Yamaguchi A, Abe M (2012) Regulation of reproductive development by non-coding RNA in Arabidopsis: to flower or not to flower. J Plant Res 125:693–704
Yang Y, Wu Y, Pirrello J, Regad F, Bouzayen M, Deng W, Li Z (2010) Silencing SI-EBF1 and SI-EBF2 expression causes constitutive ethylene response phenotype, accelerated plant senescence, and fruit ripening in tomato. J Exp Bot 61:697–708
Yao T, Park BS, Mao HZ, Seo JS, Ohama N, Li Y, Yu N, Mustafa NFB, Huang CH, Chua NH (2019) Regulation of flowering time by SPL10/MED25 module in Arabidopsis. New Phytol 224:493–504
Yuan J, Zhang Y, Dong J, Sun Y, Lim BL, Liu D, Lu ZJ (2016) Systematic characterization of novel lncRNAs responding to phosphate starvation in Arabidopsis thaliana. BMC Genomics 17:655
Zhan S, Dong Y, Zhao W, Guo J, Zhong T, Wang L, Li L, Zhang H (2016) Genome-wide identification and characterization of long non-coding RNAs in developmental skeletal muscle of fetal goat. BMC Genomics 17:666
Zhang YC, Liao JY, Li ZY, Yu Y, Zhang JP, Li QF, Qu LH, Shu WS, Chen YQ (2014) Genome-wide screening and functional analysis identify a large number of long noncoding RNAs involved in the sexual reproduction of rice. Genome Biol 15:512
Zhang Z, Zheng Y, Ham BK, Zhang S, Fei Z, Lucas WJ (2019) Plant lncRNAs are enriched in and move systemically through the phloem in response to phosphate deficiency. J Integr Plant Biol 61:492–508
Zhao X, Li J, Lian B, Gu H, Li Y, Qi Y (2018) Global identification of Arabidopsis lncRNAs reveals the regulation of MAF4 by a natural antisense RNA. Nat Commun 9:5056
Data availability statement
The raw RNA-seq datasets analyzed during the current study were deposited in National Center for Biotechnology Information (NCBI) Short Read Archive (SRA) accession SRP134709 (https://www.ncbi.nlm.nih.gov/sra/SRP134709).
Funding
The research was financially supported by the National Natural Science Foundation of China (31570666, 31971672, 31670682, and 31600547), the Natural Science Foundation of Zhejiang Province (LY18C150002).
Author information
Authors and Affiliations
Contributions
Youjun Huang and Zhengjia Wang designed the experiment. Qixiang Zhang participated in discussion part. Tongqiang Fan, Youjun Huang, Yuanyuan Hu, Qixiang Zhang, and Zhengjia Wang participated in the revision of manuscript. Tongqiang Fan carried out statistical analysis and manuscript draft. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics statement
No specific permissions were required for these locations/activities because all samples were collected from Caray cathayensis flower buds of Donghu Campus of Zhejiang A&F University (30°N, 119°W), Lin’an, China. We collected flower buds from hickory for research, and also confirmed that the field studies did not involve any endangered or protected species.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Table S1
Summary of the RNA-seq data and reads mapped to the Carya cathayensis reference genome (XLSX 13 kb)
Table S2
List of the lncRNAs identified in Carya cathayensis pistillate and staminate bud libraries (XLSX 100 kb)
Table S3
List of differentially expressed lncRNAs from three female floral bud libraries (XLSX 101 kb)
Table S4
List of differentially expressed lncRNAs from eight male floral bud libraries (XLSX 214 kb)
Table S5
Detected protein-coding genes 10 kb upstream and downstream of all the identified lncRNAs (XLSX 484 kb)
Table S6
Functional enrichment analysis of protein-coding genes targeted by cis-acting lncRNAs (XLSX 102 kb)
Table S7
Functional enrichment analysis of protein-coding genes targeted by cis-acting lncRNAs that were involved in some key processes (XLSX 20 kb)
Table S8
Co-expression analysis of protein-coding genes and lncRNAs with PCC (Pearson’s correlation coefficient) > 0.95 or <− 0.95 (XLSX 1390 kb)
Table S9
Functional enrichment analysis of protein-coding genes that co-expressed with lncRNAs (XLSX 190 kb)
Table S10
Co-expression analysis of lncRNAs and mRNA involved in floral development (XLSX 23 kb)
Table S11
(XLSX 35 kb)
Table S12
(XLSX 4576 kb)
Table S13
(XLSX 11 kb)
Rights and permissions
About this article
Cite this article
Fan, T., Zhang, Q., Hu, Y. et al. Genome-wide identification of lncRNAs during hickory (Carya cathayensis) flowering. Funct Integr Genomics 20, 591–607 (2020). https://doi.org/10.1007/s10142-020-00737-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-020-00737-w