Abstract
In the present study, de novo transcriptome analysis of Selaginella bryopteris in frond and root was performed to understand the regulation of flavonoid (FL) biosynthesis. High-quality data of 5.84 and 5.86 Gb was generated for frond and root, respectively, that assembled into 94,713 and 81,567 transcripts. A total of 87,471 and 73,395 unigenes were obtained from frond and root, respectively. A total of 41,267 and 31,048 CDS of frond and root, respectively, were annotated by BLASTX, which showed maximum hits against S. moellendorffii. Out of 11,285 differentially expressed genes, a total of 5639 genes were found to be down-regulated and 5628 genes up-regulated in frond as compared to those in root. In silico analysis of expression of genes in frond as compared to that in root was done for those related to phenylpropanoid (PP)/FL biosynthesis along with transcription factors (TFs) after DESeq and MapMan-based information. Results showed that genes of PP/FL biosynthesis pathway namely SbCHS, SbCHI, SbF3H, SbF3’H, SbDFR, SbUF3GT, SbCCOAMT, and SbCATOMT and TFs (SbMYB1, SbMYB2, SbMYB3, SbBHLH1, and SbWD40-5) were up-regulated in frond in comparison to those in root. Further, this in silico expression data was validated by RT-PCR analysis which showed predominant expression of most of these genes in frond and indicated their importance in the biosynthesis of flavonoids in S. bryopteris. A total of 9074 simple sequence repeats (SSRs) were also identified for frond and 3811 SSRs for root; these can be used for experimental validation.







Similar content being viewed by others
References
Agarwal AV, Gupta P, Singh D, Dhar YV, Chandra D, Trivedi PK (2017) Comprehensive assessment of the genes involved in withanolide biosynthesis from Withania somnifera: chemotype-specific and elicitor-responsive expression. Funct Integr Genomics 17(4):477–490
Aguilar MI, Romero MG, Chávez MI, King-Díaz B, Lotina-Hennsen B (2008) Biflavonoids isolated from Selaginella lepidophylla inhibit photosynthesis in spinach chloroplasts. J Agric Food Chem 56:6994–7000
Appert C, Logemann E, Hahlbrock K, Schmid J, Amrhein N (1994) Structural and catalytic properties of the four phenylala- nine ammonia-lyase isoenzymes from parsley (ktroselinum crispum Nym.) Eur J Biochem 225:491–499
Bate NJ, Orr J, Ni W, Meromi A, Nadler-Hassar T, Doerner PW, Dixon RA, Lamb CJ, Elkind Y (1994) Quantitative relationship between phenylalanine ammonia-lyase levels and phenylpropanoid accumulation in transgenic tobacco identifies a rate-limiting step in natural product synthesis. Proc Natl Acad Sci 91:7608–7612
Bogs J, Jaffé FW, Takos AM, Walker AR, Robinson SP (2007) The grapevine transcription factor VvMYBPA1 regulates proanthocyanidin synthesis during fruit development. Plant Physiol 143(3):1347–1361
Brodowska KM (2017) Natural flavonoids: classification, potential role, and application of flavonoid analogues. Eur. J Biol Res 7(2):108–123
Cao Y, Tan N, Chen J, Zeng G, Ma Y, Wu Y, Yan H, Yang J, Lu L, Wang Q (2010) Bioactive flavones and biflavones from Selaginella moellendorffii Hieron. Fitoterapia 81:253–258
Cordeiro GM, Casu R, McIntyre CL, Manners JM, Henry RJ (2001) Microsatellite markers from sugarcane (Saccharum spp.) ESTs cross transferable to erianthus and sorghum. Plant Science 160(6):1115-1123
Deeba F, Pandey V, Pathre U, Kanojiya S (2009) Proteome analysis of detached fronds from a resurrection plant selaginella bryopteris - response to dehydration and rehydration. Journal of Proteomics & Bioinformatics 02(02):108–116
Dixon RA, Paiva NL (1995) Stress-induced phenylpropanoid metabolism. Plant Cell 7:1085–1097
Ehlting J, Buettner D, Wang Q, Douglas CJ, Somssich IE, Kombrink E (1999) Three 4-coumarate: coenzyme a ligases in Arabidopsis thaliana represent two evolutionarily divergent classes in angiosperms. Plant J 19:9–20
Feldbrügge M, Sprenger M, Hahlbrock K, Weisshaar B (1997) PcMYB1, a novel plant protein containing a DNA-binding domain with one MYB repeat, interacts in vivo with a light-regulatory promoter unit. Plant J 11(5):1079–1093
Ganeshaiah KN, Vasudeva R, Uma Shaanker R (2009) In search of Sanjeevani. Curr Sci 97(4):484–489
Gupta PK, Balyan HS, Sharma PC, Ramesh B (1996) Microsatellites in plants: a new class of molecular markers. Curr Sci 70:45–54
Jiao Y, Jia HM, Li XW, Chai ML, Jia HJ, Chen Z, Wang GY, Chai CY, van de Weg E, Gao ZS (2012) Development of simple sequence repeat (SSR) markers from a genome survey of Chinese bayberry (Myrica rubra). BMC Genomics 13:201
Jorrin J, Dixon RA (1990) Stress responses in alfalfa (Medicago sativa L.). II.Purification, characterization, and induction of phenylalanine ammonia-lyaseisofonns from elicitor-treated cell suspension cultures. Plant Physiol 92:447–455
Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, Itoh M, Katayama T, Kawashima S, Okuda S, Tokimatsu T, Yamanishi Y (2008) KEGG for linking genomes to life and the environment. Nucleic Acids Res. 36(Database issue):D480–4
Kim HJ, Jung J, Kim MS, Lee JM, Choi D, Yeam I (2015) Molecular marker development and genetic diversity exploration by RNA-seq in Platycodon grandiflorum. Genome 58(10):441–451
Kumar S, Kalra S, Singh B, Kumar A, Kaur J, Singh K (2016) RNA-Seq mediated root transcriptome analysis of Chlorophytum borivilianum for identification of genes involved in saponin biosynthesis. Funct Integr Genomics 16(1):37–55
Li S (2014) Transcriptional control of flavonoid biosynthesis: fine-tuning of the MYB-bHLH-WD40 (MBW) complex. Plant Signal Behav 9(1):e27522
Li W, Godzik A (2006) Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22:1658–1659
Liu J, Osbourn A, Ma P (2015) MYB transcription factors as regulators of phenylpropanoid metabolism in plants. Mol Plant 8(5):689–708
Liu JF, Xu KP, Li FS, Shen J, Hu CP, Zou H, Yang F, Liu GR, Xiang HL, Zhou YJ, Li YJ, Tan GS (2010) A new flavonoid from Selaginella tamariscina (Beauv.) Spring. Chem Pharmaceu Bull 58:549–551
Lohse M, Nagel A, Herter T, May P, Schroda M, Zrenner R et al (2014) Mercator: a fast and simple web server for genome scale functional annotation of plant sequence data. Plant Cell Environ 37:1250–1258
Marchler-Bauer A, Lu S, Anderson JB, Chitsaz F, Derbyshire MK, DeWeese-Scott C, Fong JH, Geer LY, Geer RC, Gonzales NR, Gwadz M, Hurwitz DI, Jackson JD, Ke Z, Lanczycki CJ, Lu F, Marchler GH, Mullokandov M, Omelchenko MV, Robertson CL, Song JS, Thanki N, Yamashita RA, Zhang D, Zhang N, Zheng C, Bryant SH (2010) CDD: a Conserved Domain Database for the functional annotation of proteins. Nucleic Acids Res 39(Database):D225-D229
Martinelli F, Tonutti P (2012) Flavonoid metabolism and gene expression in developing olive (Olea europaea L.) fruit. Plant Biosys. https://doi.org/10.1080/11263504.2012.68132
Pandey A, Misra P, Trivedi PK (2015) Constitutive expression of Arabidopsis MYB transcription factor, AtMYB11, in tobacco modulates flavonoid biosynthesis in favor of flavonol accumulation. Plant Cell Rep 34(9):1515–1528
Payyavula RS, Singh RK, Navarre DA (2013) Transcription factors, sucrose, and sucrose metabolic genes interact to regulate potato phenylpropanoid metabolism. J Exp Bot 64(16):5115–5131
Qi LJ, Long P, Jiang C, Yuan Y, Huang LQ (2015) Development of microsatellites and genetic diversity analysis of Scutellaria baicalensis Georgi using genomic-SSR markers. Yao Xue Xue Bao 50(4):500–505
Sah P (2008) Does the magical Himalayan Herb “Sanjeevani Booti” really exist in Nature. J American Sci 4(3):65–67
Sah NK, Singh SN, Sahdev S, Banerji S, Jha V, Khan Z, Hasnain SE (2005) Indian herb 'Sanjeevani' (Selaginella bryopteris) can promote growth and protect against heat shock and apoptotic activities of ultra violet and oxidative stress. J Biosci 30(4): 499–505
Singh K, Kumar S, Rani A, Gulati A, Ahuja PS (2009) Phenylalanine ammonia-lyase (PAL) and cinnamate 4-hydroxylase (C4H) and catechins (flavan-3-ols) accumulation in tea. Funct Integr Genomics 9:125–134
Singh K, Rani A, Kumar S, Sood P, Mahajan M, Yadav SK, Bikram S, Ahuja PS (2008) An early gene of the flavonoid pathway, flavanone 3-hydroxylase, exhibits a positive relationship with the concentration of catechins in tea (Camellia sinensis). Tree Physiol 28:1349–1356
Singh RS, Kumar S (2012) A protocol to remove colored metabolites and other inhibitors (RECOIN) from plant tissues to facilitate RNA isolation suitable for downstream applications. Biotechnol Prog 28(5):1303–1307
Singh K, Raizada J, Bhardwaj P, Ghawana S, Rani A, Singh H, Kaul K, Kumar S (2004) 26S rRNA based internal control gene primer pair for reverse transcription-polymerase chain reaction-based quantitative expression studies in diverse plant species. Anal Biochem 335:330–333
Singh R, Kumar R, Mahato AK, Paliwal R, Singh AK, Kumar S, Marla SS, Kumar A, Singh NK (2016) De novo transcriptome sequencing facilitates genomic resource generation in Tinospora cordifolia. Funct Integr Genomics 16(5):581–591
Singh RS, Gara RK, Bhardwaj PK, Kaachra A, Malik S, Kumar R, Sharma M, Ahuja PS, Kumar S (2010) Expression of 3-hydroxy-3-methylglutaryl-CoA reductase, p-hydroxybenzoate-m-geranyltransferase and genes of phenylpropanoid pathway exhibits positive correlation with shikonins content in arnebia [Arnebia euchroma (Royle) Johnston]. BMC Mol Biol 11:88
Sobhani NA, Naghavi MR, Farahmand H, Abbasi A (2017) Transcriptome and metabolome analysis of Ferula gummosa Boiss. to reveal major biosynthetic pathways of galbanum compounds. Funct Integr Genomics 17(6):725–737
Thimm O, Bläesing O, Gibon Y, Nagel A, Meyer S, Krüger P et al (2004) MAPMAN: a user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J 37:914–939
URL http://www.genscript.com/cgi-bin/tools/sequencing_primer_design
Varshney RK, Graner A, Sorrells ME (2005) Genic microsatellites in plants: features and applications. Trends Bitechnol 23:48–55
Verpoorte R, van der Heijden R, Memelink J (2000) Engineering the plant cell factory for secondary metabolite production. Transgenic Res 9(4):323–343
Vieira ML, Santini L, Diniz AL, Munhoz Cde F (2016) Microsatellite markers: what they mean and why they are so useful. Genet Mol Biol 39(3):312–328
Weng JK, Noel JP (2013) Chemodiversity in Selaginella: a reference system for parallel and convergent metabolic evolution in terrestrial plants. Front Plant Sci 4:1–17
Wei X, Wang L, Zhang Y, Qi X, Wang X, Ding X, Zhang J, Zhang X (2014) Development of simple sequence repeat (SSR) markers of sesame (Sesamum indicum) from a genome survey. Molecules 19(4):5150–5162
Xu W, Dubos C, Lepiniec L (2015) Transcriptional control of flavonoid biosynthesis by MYB–bHLH–WDR complexes. Trends Plant Sci 20:176–185
Yang SF, Chu SC, Liu SJ, Chen YC, Chang YZ, Hsieh YS (2007) Antimetastatic activities of Selaginella tamariscina (Beauv.) on lung cancer cells in vitro and in vivo. J Ethnopharmacol 110(3):483–489
Ye J, Fang L, Zheng HK, Zhang Y, Chen J, Zhang ZJ, Wang J, Li S, Li R, Bolund L, Wang J (2006) WEGO: a web tool for plotting GO annotations. Nucleic Acids Res 34:W293–W297
Yin D, Li J, Lei X, Liu Y, Yang Z, Chen K (2014) Antiviral activity of total flavonoid extracts from Selaginella moellendorffii Hieron against Coxsackie virus B3 in vitro and in vivo. Evid Based Complement Alternat Med 2014:1–7
Zhai R, Wang Z, Zhang S, Meng G, Song L, Wang Z, Li P, Ma F, Xu L (2016) Two MYB transcription factors regulate flavonoid biosynthesis in pear fruit (Pyrus bretschneideri Rehd.) J Exp Bot 67(5):1275–1284
Zhang J, Song TT, Meng XN, Han ZY, Yao YC (2015) Early phenylpropanoid biosynthetic pathway genes are responsible for flavonoid accumulation in the leaves of three crabapple (Malus spp.) cultivars. J Hort Sci Biotech 90(5): 489–502
Zhao L, Gao L, Wang H, Xiaotian C, Yunsheng W, Hua Y, Chaoling W, Xiaochun W, Tao X (2013) The R2R3-MYB, bHLH, WD40, and related transcription factors in flavonoid biosynthesis. Funct Integr Genomics 13:75–98
Zhao P, Chen KL, Zhang GL, Deng GR, Li J (2017) Pharmacological basis for use of Selaginella moellendorffii in gouty arthritis: antihyperuricemic, anti-inflammatory, and xanthine oxidase inhibition. Evid Based Complement Alternat Med 2017:2103254
Zhou H, Cai BH, Lü ZQ, Gao ZH, Qiao YS (2016) Development, characterization, and annotation of potential simple sequence repeats by transcriptome sequencing in pears (Pyrus pyrifolia Nakai). Genet Mol Res 15(3)
Zheng X-K, Wang W-W, Zhang L, C-f S, Wu Y-Y, Ke Y-Y, Hou Q-W, Liu Z-Y, Gao A-S, Feng W-S (2013) Antihyperlipidaemic and antioxidant effect of the total flavonoids in Selaginella tamariscina (Beauv.) Spring in diabetic mice. J Pharm Pharmacol 65:757–766
Zou Z, Xu K, Xu P, Li X, Cheng F, Li J, Yu X, Cao D, Li D, Zeng W, Zhang G, Tan G (2017) Seladoeflavones A-F, six novel flavonoids from Selaginella doederleinii. Fitoterapia 116:66–71
Zoua H, Xua KP, Lia FS, Zoua ZX, Liua R, Liua RH, Lia J, Tana LH, Tana GS (2014) Unciflavones A–F, six novel flavonoids from Selaginella uncinata. Fitoterapia 99:328–333
Acknowledgements
Authors thank and acknowledge the SERB-DST, New Delhi, Board of Research in Nuclear Sciences-Department of Atomic Energy, Mumbai and Bihar Agricultural University, Sabour for all the supports and facilities. M/s Xcelris Labs Ltd., Ahmedabad, India, is acknowledged for their sequencing services. The manuscript represents Bihar Agricultural University Communication No. 259/2017.
Funding
Grants received from the SERB-DST (SB/YS/LS-209/2013) and the Board of Research in Nuclear Sciences-Department of Atomic Energy (2013/35/39/BRNS/1362) for the study on Selaginella bryopteris.
Author information
Authors and Affiliations
Contributions
Conceived the idea and designed the experiment: RSS; performed the experiments and gene expression analysis: RSS, UJ, VJ, and AK; result/data interpretation: RSS and RK; prepared the manuscript: RSS; edited the manuscript: RSS, RK, AKP, TK, and PKS.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Electronic supplementary material
Supplementary Fig. 1
Unigene length distribution in frond and root transcriptome. (PPT 214 kb)
Supplementary Fig. 2
Transcript length distribution in frond and root transcriptome. (PPT 219 kb)
Supplementary Fig. 3
CDS length distribution in frond and root transcriptome. (PPT 180 kb)
Supplementary Fig. 4
MapMan visualization of the metabolic pathways. (A) Total frond CDS mapped, (B) Top 500 frond CDS mapped. (PPT 288 kb)
Supplementary Fig. 5
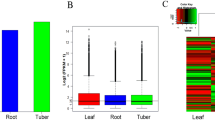
Distribution of gene expression for root and frond is shown with Volcano plot. DEseq was performed to show the differentially expressed genes. Green, red, and gray correspond to genes with p-value <0.05 and logFC <0, p-value <0.05 and logFC >0, p-value >0.05 (Not DEG) respectively (Panel a); Digital gene expression between root Sample and frond sample. Each point represents a gene. The x- and y-axis are the log2 of the normalized gene expression level of control and experimental sample. Red and green points indicate significant change at the absolute value of log2 (FPKM ratio in two groups) and p-value<0.05 Red points indicate up-regulated genes and green points indicate down-regulated genes in the two groups which its expression level is represented by the y-axis (Panel b). (PPT 673 kb)
Supplementary Table 1
Primer sequences and PCR conditions used in RT-PCR based expression analysis (DOC 38 kb)
Supplementary Table 2
Frond transcriptome Annotation (XLS 13456 kb)
Supplementary Table 3
Root transcriptome Annotation (XLS 10336 kb)
Supplementary Table 4
Frond transcriptome GO annotation (XLS 3414 kb)
Supplementary Table 5
Root transcriptome GO annotation (XLS 2479 kb)
Supplementary Table 6
Frond genes KEGG pathway annotation (XLS 741 kb)
Supplementary Table 7
Root genes KEGG pathway annotation (XLS 686 kb)
Supplementary Table 8
Top 500 frond CDS mapped for secondary metabolites by MapMan (XLS 45 kb)
Supplementary Table 9
All frond CDS mapped for secondary metabolism by MapMan (XLS 125 kb)
Supplementary Table 10
Differentially expressed genes in fronds and root (XLS 21379 kb)
Supplementary Table 11
SSR in frond transcriptome of S. bryopteris (XLS 3223 kb)
Supplementary Table 12
SSR in root transcriptome of S. bryopteris (XLS 1496 kb)
Rights and permissions
About this article
Cite this article
Singh, R.S., Kesari, R., Kumar, U. et al. Candidate genes of flavonoid biosynthesis in Selaginella bryopteris (L.) Baker identified by RNA-Seq. Funct Integr Genomics 18, 505–517 (2018). https://doi.org/10.1007/s10142-018-0603-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-018-0603-2