Abstract
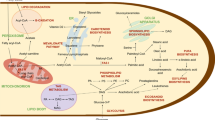
Beta oxidation is the principal metabolic pathway for fatty acid degradation. The pathway is virtually universally present throughout eukaryotes yet displays different forms in enzyme architecture, substrate specificity, and subcellular location. In this review, we examine beta oxidation across the fungal kingdom by conducting a large-scale in silico screen and localization prediction for all relevant enzymes in >50 species. The survey reveals that fungi exhibit an astounding diversity of beta oxidation pathways and shows that the combined presence of distinct mitochondrial and peroxisomal pathways is the prevailing and likely ancestral type of beta oxidation in fungi. In addition, the available information indicates that the mitochondrial pathway was lost in the common ancestor of Saccharomycetes. Finally, we infer the existence of a hybrid peroxisomal pathway in several Sordariomycetes, including Neurospora crassa. In these cases, a typically mitochondrion-located enzyme compensates for the lack of a peroxisomal one.


Similar content being viewed by others
References
Cornell MJ, Alam I, Soanes DM, Wong HM, Hedeler C, Paton NW, Rattray M, Hubbard SJ, Talbot NJ, Oliver SG (2007) Comparative genome analysis across a kingdom of eukaryotic organisms: specialization and diversification in the fungi. Genome Res 17:1809–1822
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Filppula SA, Yagi AI, Kilpelainen SH, Novikov D, FitzPatrick DR, Vihinen M, Valle D, Hiltunen JK (1998) Delta3,5-delta2,4-dienoyl-CoA isomerase from rat liver. Molecular characterization. J Biol Chem 273:349–355
Hiltunen JK, Qin Y (2000) Beta-oxidation—strategies for the metabolism of a wide variety of acyl-CoA esters. Biochim Biophys Acta 1484:117–128
Hiltunen JK, Wenzel B, Beyer A, Erdmann R, Fossa A, Kunau WH (1992) Peroxisomal multifunctional beta-oxidation protein of Saccharomyces cerevisiae. Molecular analysis of the fox2 gene and gene product. J Biol Chem 267:6646–6653
Kawachi H, Atomi H, Ueda M, Tanaka A (1996) Peroxisomal and mitochondrial carnitine acetyltransferases of the n-alkane-assimilating yeast Candida tropicalis. Analysis of gene structure and translation products. Eur J Biochem 238:845–852
Kionka C, Kunau WH (1985) Inducible beta-oxidation pathway in Neurospora crassa. J Bacteriol 161:153–157
Kurihara T, Ueda M, Okada H, Kamasawa N, Naito N, Osumi M, Tanaka A (1992) Beta-oxidation of butyrate, the short-chain-length fatty acid, occurs in peroxisomes in the yeast Candida tropicalis. J Biochem 111:783–787
Lartillot N, Philippe H (2004) A Bayesian mixture model for across-site heterogeneities in the amino-acid replacement process. Mol Biol Evol 21:1095–1109
Maggio-Hall LA, Keller NP (2004) Mitochondrial beta-oxidation in Aspergillus nidulans. Mol Microbiol 54:1173–1185
Masterson C, Wood C (2000) Mitochondrial beta-oxidation of fatty acids in higher plants. Physiol Plant 109:217–224
Poirier Y, Antonenkov VD, Glumoff T, Hiltunen JK (2006) Peroxisomal beta-oxidation—a metabolic pathway with multiple functions. Biochim Biophys Acta 1763:1413–1426
Thieringer R, Kunau WH (1991) The beta-oxidation system in catalase-free microbodies of the filamentous fungus Neurospora crassa. Purification of a multifunctional protein possessing 2-enoyl-CoA hydratase, L-3-hydroxyacyl-CoA dehydrogenase, and 3-hydroxyacyl-CoA epimerase activities. J Biol Chem 266:13110–13117
Uchida Y, Izai K, Orii T, Hashimoto T (1992) Novel fatty acid beta-oxidation enzymes in rat liver mitochondria. II. Purification and properties of enoyl-coenzyme A (CoA) hydratase/3-hydroxyacyl-CoA dehydrogenase/3-ketoacyl-CoA thiolase trifunctional protein. J Biol Chem 267:1034–1041
Wanders RJ, Waterham HR (2006) Biochemistry of mammalian peroxisomes revisited. Annu Rev Biochem 75:295–332
Wang ZY, Soanes DM, Kershaw MJ, Talbot NJ (2007) Functional analysis of lipid metabolism in Magnaporthe grisea reveals a requirement for peroxisomal fatty acid beta-oxidation during appressorium-mediated plant infection. Mol Plant Microbe Interact 20:475–491
Acknowledgements
This work was supported by the Genome-Canada and Canadian Institutes of Health Research (CIHR, Institute of Genetics, MOP-79309). We would like to thank B. Franz Lang, Henner Brinkmann, Pierre Rioux, and Nicolas Lartillot (Université de Montréal) for their help with phylogenetic analyses. We also thank Emmet O’Brien (Université de Montréal) for improving the manuscript. YQS is a Canadian Institute for Health Research (CIHR) Strategic Training Fellow in Bioinformatics.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary materials
Below is the link to the electronic supplementary materials.
Supplementary Table 1
Fungal proteins with significant sequence similarity to both mitochondrial and peroxisomal keto-acyl-CoA thiolase (DOC 152 KB)
Supplementary Table 2
Sequence IDs for mitochondrial beta oxidation enzymes (DOC 127 KB)
Supplementary Table 3
Sequence IDs for peroxisomal beta oxidation enzymes (DOC 105 KB)
Supplementary Table 4
Keto-acyl-CoA thiolase homologs predicted as targeted to both mitochondria and peroxisomes (DOC 68.0 KB)
Supplementary Table 5
Protein sequences used for constructing the phylogenetic tree (DOC 142 KB)
Rights and permissions
About this article
Cite this article
Shen, YQ., Burger, G. Plasticity of a key metabolic pathway in fungi. Funct Integr Genomics 9, 145–151 (2009). https://doi.org/10.1007/s10142-008-0095-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-008-0095-6